# Parameters
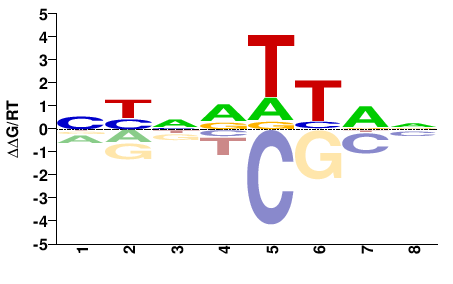
data_name = "Lbx2"
task_idxs = "0-213,216-278"
homer_root = "/srv/scratch/msharmin/mouse_hem/with_tfd/full_mouse50/Naive_scans"
modisco_root = "/srv/scratch/msharmin/mouse_hem/with_tfd/full_mouse50/Naive_modisco2019"
overwrite = False
from matlas.reports import collect_motifDB_to_be_displayed, display_comparative_cell_types
load data from labcluster
Using TensorFlow backend. 2019-08-23 11:16:38,115 [WARNING] git-lfs not installed
homer_matches, modisco_matches = collect_motifDB_to_be_displayed(task_idxs, homer_root, modisco_root,
overwrite=False)
TF-MoDISco is using the TensorFlow backend.
display_comparative_cell_types(data_name, homer_matches, modisco_matches)
Shared Cell types both TF-MoDISco and Homer finds Lbx2: #18
| Cell type | Modisco | Homer |
|---|---|---|
| wES;limb;E11.5;D;En |
Significance: 0.000215574 |
Significance: 0.00858817 |
| wES;Fr_lmb;E11.5;D;En |
Significance: 0.000134401 |
Significance: 0.030745599999999994 |
| wES;Hnd_lmb;E11.5;D;En |
Significance: 0.000132104 |
Significance: 0.00607199 |
| wES;Hnd_BRN;E14.5;D;En |
Significance: 0.0020622 |
Significance: 0.014081 |
| BCK_SKN;A;GEO |
Significance: 0.00135904 |
Significance: 0.00899313 |
| CRBL_GRNL_Precrsr;A;GEO |
Significance: 0.00567005 |
Significance: 0.0135025 |
| RTN;D1;D;En |
Significance: 0.00101796 |
Significance: 0.00400243 |
| wES;RTN;E11.5;D;En |
Significance: 0.00036333 |
Significance: 0.00523812 |
| G_NRN_P;A;GEO |
Significance: 0.00487644 |
Significance: 0.015007 |
| Muller;P12;D;En |
Significance: 0.00212854 |
Significance: 0.00542561 |
| wES;WHL_BRN;E14.5;D;GEO |
Significance: 0.000768605 |
Significance: 0.00403467 |
| wES.Fr_BRN;E14.4;D;En |
Significance: 0.00041474599999999996 |
Significance: 0.00397983 |
| Fr_BRN;P0;D;En |
Significance: 0.0286139 |
Significance: 0.0164003 |
| Md_BRN;D;En |
Significance: 0.00829087 |
Significance: 0.00451696 |
| BL_CN_PHT_RCPTR;A;GEO |
Significance: 0.0242378 |
Significance: 0.011833100000000001 |
| wES;Md_BRN;E14.4;D;En |
Significance: 0.0005397080000000001 |
Significance: 0.0154559 |
| wES;3rd_Rhmbmr;E8.5;A;GEO |
Significance: 0.00313145 |
Significance: 0.0150715 |
| wES;WHL_BRN;E14.5;D;En |
Significance: 0.00042388800000000003 |
Significance: 0.00473719 |
Unique Cell types only TF-MoDISco finds Lbx2: #19
| Cell type | Modisco | Homer |
|---|---|---|
| wES;5th_Rhmbmr;E8.5;A;GEO |
Significance: 0.0254202 | Absent |
| npES;5th_Rhmbmr_delA;E8.5;A;GEO |
Significance: 0.0103002 | Absent |
| wES;posterior;2;E9.5;A;GEO |
Significance: 0.00398197 | Absent |
| wES;FaceP;E14.5;D;En |
Significance: 0.0322059 | Absent |
| wES;posterior;1;E8.5;A;GEO |
Significance: 0.00124343 | Absent |
| npES;posterior_delC;E8.5;A;GEO |
Significance: 0.000197315 | Absent |
| ES;E14;A;GEO |
Significance: 0.038042599999999996 | Absent |
| ES;v6.5;A;GEO |
Significance: 0.0399505 | Absent |
| NPC;F129;A;GEO |
Significance: 0.0482801 | Absent |
| NPC;G4;A;GEO |
Significance: 0.0263468 | Absent |
| NPC;mE6;A;GEO |
Significance: 0.0317732 | Absent |
| iPV_NRN;A;GEO |
Significance: 0.0534631 | Absent |
| wES;MSDRM;E11.5;D;En |
Significance: 0.00266261 | Absent |
| Hnd_BRN;P0;D;En |
Significance: 0.000579601 | Absent |
| npES;ZHBTC4;Brg1fl_fl;untreat;A;GEO |
Significance: 0.0346939 | Absent |
| ES;C57BL_6J;R1;D;En |
Significance: 0.0332122 | Absent |
| npES;3rd_Rhmbmr_delA;E8.5;A;GEO |
Significance: 0.00316383 | Absent |
| npES;3rd_Rhmbmr_delC;E8.5;A;GEO |
Significance: 0.00220586 | Absent |
| wES;5th_Rhmbmr;E9.5;A;GEO |
Significance: 0.028922399999999997 | Absent |
Unique Cell types only Homer finds Lbx2: #8
| Cell type | Modisco | Homer |
|---|---|---|
| BRN;OLFTRY_BLB;A;CL | Absent |
Significance: 0.0498961 |
| wES;FaceP;E11.5;D;En | Absent |
Significance: 0.0356255 |
| wES.CRNL_NRL_CRST;E10.5;A;GEO | Absent |
Significance: 0.0240568 |
| iVIP_NRN;A;GEO | Absent |
Significance: 0.0198388 |
| MTR_NRN_MN1;methyl3;D;En | Absent |
Significance: 0.026036200000000002 |
| MN1;D;En | Absent |
Significance: 0.0371164 |
| Melanotrope;A;GEO | Absent |
Significance: 0.018005700000000003 |
| CN_PHT_RCPTR;A;GEO | Absent |
Significance: 0.012446899999999999 |