| Rank | Motif | Name | P-value | log P-pvalue | q-value (Benjamini) | # Target Sequences with Motif | % of Targets Sequences with Motif | # Background Sequences with Motif | % of Background Sequences with Motif | Motif File |
PDF |
| 1 |  | BORIS(Zf)/K562-CTCFL-ChIP-Seq(GSE32465)/Homer | 1e-255 | -5.879e+02 | 0.0000 | 29160.0 | 46.12% | 188490.6 | 39.00% | motif file (matrix) |
pdf |
| 2 |  | CTCF(Zf)/CD4+-CTCF-ChIP-Seq(Barski_et_al.)/Homer | 1e-163 | -3.762e+02 | 0.0000 | 25529.0 | 40.37% | 168248.1 | 34.81% | motif file (matrix) |
pdf |
| 3 |  | CTCF-SatelliteElement(Zf?)/CD4+-CTCF-ChIP-Seq(Barski_et_al.)/Homer | 1e-80 | -1.861e+02 | 0.0000 | 3900.0 | 6.17% | 21279.5 | 4.40% | motif file (matrix) |
pdf |
| 4 |  | c-Myc(bHLH)/mES-cMyc-ChIP-Seq(GSE11431)/Homer | 1e-23 | -5.445e+01 | 0.0000 | 8763.0 | 13.86% | 60035.7 | 12.42% | motif file (matrix) |
pdf |
| 5 |  | SeqBias: GA-repeat | 1e-17 | -4.066e+01 | 0.0000 | 55720.0 | 88.12% | 420021.7 | 86.90% | motif file (matrix) |
pdf |
| 6 |  | PIF5ox(bHLH)/Arabidopsis-PIF5ox-ChIP-Seq(GSE35062)/Homer | 1e-17 | -4.037e+01 | 0.0000 | 14674.0 | 23.21% | 104821.7 | 21.69% | motif file (matrix) |
pdf |
| 7 |  | n-Myc(bHLH)/mES-nMyc-ChIP-Seq(GSE11431)/Homer | 1e-15 | -3.607e+01 | 0.0000 | 11324.0 | 17.91% | 80300.4 | 16.61% | motif file (matrix) |
pdf |
| 8 |  | SeqBias: CG bias | 1e-15 | -3.503e+01 | 0.0000 | 60502.0 | 95.68% | 458977.1 | 94.96% | motif file (matrix) |
pdf |
| 9 |  | PIF4(bHLH)/Seedling-PIF4-ChIP-Seq(GSE35315)/Homer | 1e-14 | -3.414e+01 | 0.0000 | 16894.0 | 26.72% | 122074.0 | 25.26% | motif file (matrix) |
pdf |
| 10 |  | Max(bHLH)/K562-Max-ChIP-Seq(GSE31477)/Homer | 1e-14 | -3.397e+01 | 0.0000 | 10441.0 | 16.51% | 73946.8 | 15.30% | motif file (matrix) |
pdf |
| 11 |  | ARE(NR)/LNCAP-AR-ChIP-Seq(GSE27824)/Homer | 1e-13 | -3.018e+01 | 0.0000 | 4665.0 | 7.38% | 31837.9 | 6.59% | motif file (matrix) |
pdf |
| 12 |  | BMAL1(bHLH)/Liver-Bmal1-ChIP-Seq(GSE39860)/Homer | 1e-11 | -2.723e+01 | 0.0000 | 22604.0 | 35.75% | 165980.1 | 34.34% | motif file (matrix) |
pdf |
| 13 |  | ZNF143|STAF(Zf)/CUTLL-ZNF143-ChIP-Seq(GSE29600)/Homer | 1e-11 | -2.640e+01 | 0.0000 | 6447.0 | 10.20% | 45135.5 | 9.34% | motif file (matrix) |
pdf |
| 14 |  | GATA3(Zf),DR8/iTreg-Gata3-ChIP-Seq(GSE20898)/Homer | 1e-10 | -2.335e+01 | 0.0000 | 1365.0 | 2.16% | 8632.8 | 1.79% | motif file (matrix) |
pdf |
| 15 |  | Rfx5(HTH)/GM12878-Rfx5-ChIP-Seq(GSE31477)/Homer | 1e-9 | -2.194e+01 | 0.0000 | 5507.0 | 8.71% | 38608.1 | 7.99% | motif file (matrix) |
pdf |
| 16 |  | Trl(Zf)/S2-GAGAfactor-ChIP-Seq(GSE40646)/Homer | 1e-8 | -1.943e+01 | 0.0000 | 36353.0 | 57.49% | 272013.7 | 56.28% | motif file (matrix) |
pdf |
| 17 |  | SeqBias: CA-repeat | 1e-8 | -1.893e+01 | 0.0000 | 51552.0 | 81.53% | 389473.6 | 80.58% | motif file (matrix) |
pdf |
| 18 |  | ZNF189(Zf)/HEK293-ZNF189.GFP-ChIP-Seq(GSE58341)/Homer | 1e-4 | -1.129e+01 | 0.0003 | 11821.0 | 18.69% | 87025.5 | 18.01% | motif file (matrix) |
pdf |
| 19 |  | RUNX(Runt)/HPC7-Runx1-ChIP-Seq(GSE22178)/Homer | 1e-4 | -1.065e+01 | 0.0005 | 11081.0 | 17.52% | 81566.4 | 16.88% | motif file (matrix) |
pdf |
| 20 |  | NeuroD1(bHLH)/Islet-NeuroD1-ChIP-Seq(GSE30298)/Homer | 1e-4 | -9.930e+00 | 0.0009 | 12785.0 | 20.22% | 94548.8 | 19.56% | motif file (matrix) |
pdf |
| 21 |  | RUNX2(Runt)/PCa-RUNX2-ChIP-Seq(GSE33889)/Homer | 1e-4 | -9.592e+00 | 0.0013 | 12301.0 | 19.45% | 90961.8 | 18.82% | motif file (matrix) |
pdf |
| 22 |  | Ptf1a(bHLH)/Panc1-Ptf1a-ChIP-Seq(GSE47459)/Homer | 1e-3 | -8.520e+00 | 0.0035 | 33267.0 | 52.61% | 250667.5 | 51.86% | motif file (matrix) |
pdf |
| 23 |  | PABPC1(?)/MEL-PABC1-CLIP-Seq(GSE69755)/Homer | 1e-3 | -8.175e+00 | 0.0047 | 20423.0 | 32.30% | 152821.9 | 31.62% | motif file (matrix) |
pdf |
| 24 |  | GFY-Staf(?,Zf)/Promoter/Homer | 1e-3 | -7.917e+00 | 0.0059 | 1473.0 | 2.33% | 10244.9 | 2.12% | motif file (matrix) |
pdf |
| 25 |  | RBFox2(?)/Heart-RBFox2-CLIP-Seq(GSE57926)/Homer | 1e-3 | -7.257e+00 | 0.0109 | 22843.0 | 36.13% | 171477.8 | 35.48% | motif file (matrix) |
pdf |
| 26 |  | MafK(bZIP)/C2C12-MafK-ChIP-Seq(GSE36030)/Homer | 1e-3 | -7.124e+00 | 0.0120 | 3158.0 | 4.99% | 22756.9 | 4.71% | motif file (matrix) |
pdf |
| 27 |  | Olig2(bHLH)/Neuron-Olig2-ChIP-Seq(GSE30882)/Homer | 1e-3 | -7.073e+00 | 0.0122 | 23897.0 | 37.79% | 179556.7 | 37.15% | motif file (matrix) |
pdf |
| 28 |  | RUNX1(Runt)/Jurkat-RUNX1-ChIP-Seq(GSE29180)/Homer | 1e-3 | -7.059e+00 | 0.0122 | 14468.0 | 22.88% | 107910.1 | 22.33% | motif file (matrix) |
pdf |
| 29 |  | SUT1?/SacCer-Promoters/Homer | 1e-2 | -5.510e+00 | 0.0540 | 52609.0 | 83.20% | 400092.5 | 82.78% | motif file (matrix) |
pdf |
| 30 |  | Nkx2.2(Homeobox)/NPC-Nkx2.2-ChIP-Seq(GSE61673)/Homer | 1e-2 | -5.361e+00 | 0.0606 | 24655.0 | 38.99% | 185869.7 | 38.46% | motif file (matrix) |
pdf |
| 31 |  | SeqBias: C/A-bias | 1e-2 | -5.164e+00 | 0.0714 | 63231.0 | 100.00% | 483284.7 | 99.99% | motif file (matrix) |
pdf |
| 32 | 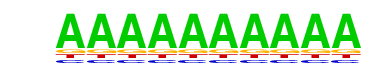 | SeqBias: polyA-repeat | 1e-2 | -4.876e+00 | 0.0922 | 62818.0 | 99.35% | 479748.1 | 99.26% | motif file (matrix) |
pdf |
| 33 | 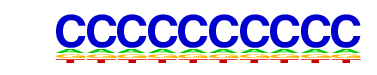 | SeqBias: polyC-repeat | 1e-2 | -4.763e+00 | 0.1002 | 63229.0 | 100.00% | 483257.1 | 99.99% | motif file (matrix) |
pdf |
| 34 |  | Nkx2.1(Homeobox)/LungAC-Nkx2.1-ChIP-Seq(GSE43252)/Homer | 1e-2 | -4.653e+00 | 0.1085 | 33565.0 | 53.08% | 254168.6 | 52.59% | motif file (matrix) |
pdf |



































































