| Rank | Motif | Name | P-value | log P-pvalue | q-value (Benjamini) | # Target Sequences with Motif | % of Targets Sequences with Motif | # Background Sequences with Motif | % of Background Sequences with Motif | Motif File |
PDF |
| 1 |  | ABF1/SacCer-Promoters/Homer | 1e-218 | -5.037e+02 | 0.0000 | 1556.0 | 34.27% | 3136.0 | 15.28% | motif file (matrix) |
pdf |
| 2 |  | REB1/SacCer-Promoters/Homer | 1e-180 | -4.162e+02 | 0.0000 | 977.0 | 21.52% | 1613.6 | 7.86% | motif file (matrix) |
pdf |
| 3 |  | RLR1?/SacCer-Promoters/Homer | 1e-83 | -1.922e+02 | 0.0000 | 2181.0 | 48.03% | 6986.4 | 34.03% | motif file (matrix) |
pdf |
| 4 |  | c-Myc(bHLH)/LNCAP-cMyc-ChIP-Seq(Unpublished)/Homer | 1e-56 | -1.301e+02 | 0.0000 | 602.0 | 13.26% | 1363.6 | 6.64% | motif file (matrix) |
pdf |
| 5 |  | bHLHE40(bHLH)/HepG2-BHLHE40-ChIP-Seq(GSE31477)/Homer | 1e-45 | -1.043e+02 | 0.0000 | 453.0 | 9.98% | 993.5 | 4.84% | motif file (matrix) |
pdf |
| 6 |  | DPL-1(E2F)/cElegans-Adult-ChIP-Seq(modEncode)/Homer | 1e-43 | -9.977e+01 | 0.0000 | 1121.0 | 24.69% | 3405.4 | 16.59% | motif file (matrix) |
pdf |
| 7 |  | n-Myc(bHLH)/mES-nMyc-ChIP-Seq(GSE11431)/Homer | 1e-42 | -9.829e+01 | 0.0000 | 683.0 | 15.04% | 1793.0 | 8.73% | motif file (matrix) |
pdf |
| 8 |  | Usf2(bHLH)/C2C12-Usf2-ChIP-Seq(GSE36030)/Homer | 1e-41 | -9.495e+01 | 0.0000 | 422.0 | 9.29% | 933.0 | 4.55% | motif file (matrix) |
pdf |
| 9 |  | Cbf1(bHLH)/Yeast-Cbf1-ChIP-Seq(GSE29506)/Homer | 1e-39 | -9.126e+01 | 0.0000 | 425.0 | 9.36% | 958.2 | 4.67% | motif file (matrix) |
pdf |
| 10 |  | CLOCK(bHLH)/Liver-Clock-ChIP-Seq(GSE39860)/Homer | 1e-39 | -9.042e+01 | 0.0000 | 584.0 | 12.86% | 1492.7 | 7.27% | motif file (matrix) |
pdf |
| 11 |  | SeqBias: CG-repeat | 1e-39 | -8.982e+01 | 0.0000 | 766.0 | 16.87% | 2142.8 | 10.44% | motif file (matrix) |
pdf |
| 12 |  | USF1(bHLH)/GM12878-Usf1-ChIP-Seq(GSE32465)/Homer | 1e-38 | -8.785e+01 | 0.0000 | 504.0 | 11.10% | 1233.1 | 6.01% | motif file (matrix) |
pdf |
| 13 |  | SFP1/SacCer-Promoters/Homer | 1e-37 | -8.589e+01 | 0.0000 | 671.0 | 14.78% | 1824.5 | 8.89% | motif file (matrix) |
pdf |
| 14 |  | Max(bHLH)/K562-Max-ChIP-Seq(GSE31477)/Homer | 1e-30 | -7.125e+01 | 0.0000 | 611.0 | 13.46% | 1702.7 | 8.29% | motif file (matrix) |
pdf |
| 15 |  | TATA-box/SacCer-Promoters/Homer | 1e-30 | -6.969e+01 | 0.0000 | 2138.0 | 47.08% | 7936.2 | 38.66% | motif file (matrix) |
pdf |
| 16 |  | E-box/Arabidopsis-Promoters/Homer | 1e-27 | -6.397e+01 | 0.0000 | 532.0 | 11.72% | 1466.3 | 7.14% | motif file (matrix) |
pdf |
| 17 |  | PIF4(bHLH)/Seedling-PIF4-ChIP-Seq(GSE35315)/Homer | 1e-27 | -6.361e+01 | 0.0000 | 1025.0 | 22.57% | 3340.7 | 16.27% | motif file (matrix) |
pdf |
| 18 |  | PIF5ox(bHLH)/Arabidopsis-PIF5ox-ChIP-Seq(GSE35062)/Homer | 1e-26 | -6.091e+01 | 0.0000 | 918.0 | 20.22% | 2944.3 | 14.34% | motif file (matrix) |
pdf |
| 19 |  | E-box(bHLH)/Promoter/Homer | 1e-26 | -6.026e+01 | 0.0000 | 139.0 | 3.06% | 218.9 | 1.07% | motif file (matrix) |
pdf |
| 20 |  | IBL1(bHLH)/Seedling-IBL1-ChIP-Seq(GSE51120)/Homer | 1e-22 | -5.270e+01 | 0.0000 | 1368.0 | 30.13% | 4858.9 | 23.67% | motif file (matrix) |
pdf |
| 21 |  | Pho2(bHLH)/Yeast-Pho2-ChIP-Seq(GSE29506)/Homer | 1e-22 | -5.256e+01 | 0.0000 | 421.0 | 9.27% | 1145.6 | 5.58% | motif file (matrix) |
pdf |
| 22 |  | SPCH(bHLH)/Seedling-SPCH-ChIP-Seq(GSE57497)/Homer | 1e-20 | -4.675e+01 | 0.0000 | 1135.0 | 24.99% | 3968.2 | 19.33% | motif file (matrix) |
pdf |
| 23 |  | E2F4(E2F)/K562-E2F4-ChIP-Seq(GSE31477)/Homer | 1e-19 | -4.600e+01 | 0.0000 | 824.0 | 18.15% | 2722.6 | 13.26% | motif file (matrix) |
pdf |
| 24 |  | FHY3(FAR1)/Arabidopsis-FHY3-ChIP-Seq(GSE30711)/Homer | 1e-18 | -4.329e+01 | 0.0000 | 414.0 | 9.12% | 1183.1 | 5.76% | motif file (matrix) |
pdf |
| 25 |  | TOD6?/SacCer-Promoters/Homer | 1e-17 | -3.944e+01 | 0.0000 | 352.0 | 7.75% | 987.3 | 4.81% | motif file (matrix) |
pdf |
| 26 |  | Egr2(Zf)/Thymocytes-Egr2-ChIP-Seq(GSE34254)/Homer | 1e-16 | -3.857e+01 | 0.0000 | 133.0 | 2.93% | 263.9 | 1.29% | motif file (matrix) |
pdf |
| 27 |  | NPAS2(bHLH)/Liver-NPAS2-ChIP-Seq(GSE39860)/Homer | 1e-16 | -3.769e+01 | 0.0000 | 820.0 | 18.06% | 2799.5 | 13.64% | motif file (matrix) |
pdf |
| 28 |  | BMAL1(bHLH)/Liver-Bmal1-ChIP-Seq(GSE39860)/Homer | 1e-15 | -3.580e+01 | 0.0000 | 1477.0 | 32.53% | 5558.8 | 27.08% | motif file (matrix) |
pdf |
| 29 |  | KLF14(Zf)/HEK293-KLF14.GFP-ChIP-Seq(GSE58341)/Homer | 1e-14 | -3.392e+01 | 0.0000 | 563.0 | 12.40% | 1825.6 | 8.89% | motif file (matrix) |
pdf |
| 30 |  | MITF(bHLH)/MastCells-MITF-ChIP-Seq(GSE48085)/Homer | 1e-14 | -3.358e+01 | 0.0000 | 808.0 | 17.79% | 2801.2 | 13.65% | motif file (matrix) |
pdf |
| 31 |  | ZNF467(Zf)/HEK293-ZNF467.GFP-ChIP-Seq(GSE58341)/Homer | 1e-13 | -3.203e+01 | 0.0000 | 332.0 | 7.31% | 969.2 | 4.72% | motif file (matrix) |
pdf |
| 32 |  | Klf4(Zf)/mES-Klf4-ChIP-Seq(GSE11431)/Homer | 1e-12 | -2.979e+01 | 0.0000 | 167.0 | 3.68% | 406.4 | 1.98% | motif file (matrix) |
pdf |
| 33 |  | c-Myc(bHLH)/mES-cMyc-ChIP-Seq(GSE11431)/Homer | 1e-12 | -2.794e+01 | 0.0000 | 355.0 | 7.82% | 1088.2 | 5.30% | motif file (matrix) |
pdf |
| 34 |  | Klf9(Zf)/GBM-Klf9-ChIP-Seq(GSE62211)/Homer | 1e-12 | -2.775e+01 | 0.0000 | 135.0 | 2.97% | 312.4 | 1.52% | motif file (matrix) |
pdf |
| 35 |  | Arnt:Ahr(bHLH)/MCF7-Arnt-ChIP-Seq(Lo_et_al.)/Homer | 1e-10 | -2.476e+01 | 0.0000 | 1057.0 | 23.28% | 3961.7 | 19.30% | motif file (matrix) |
pdf |
| 36 |  | Maz(Zf)/HepG2-Maz-ChIP-Seq(GSE31477)/Homer | 1e-10 | -2.437e+01 | 0.0000 | 289.0 | 6.36% | 873.6 | 4.26% | motif file (matrix) |
pdf |
| 37 |  | Pho4(bHLH)/Yeast-Pho4-ChIP-Seq(GSE29506)/Homer | 1e-10 | -2.377e+01 | 0.0000 | 159.0 | 3.50% | 412.5 | 2.01% | motif file (matrix) |
pdf |
| 38 |  | KLF10(Zf)/HEK293-KLF10.GFP-ChIP-Seq(GSE58341)/Homer | 1e-9 | -2.286e+01 | 0.0000 | 243.0 | 5.35% | 716.6 | 3.49% | motif file (matrix) |
pdf |
| 39 |  | KLF5(Zf)/LoVo-KLF5-ChIP-Seq(GSE49402)/Homer | 1e-9 | -2.114e+01 | 0.0000 | 459.0 | 10.11% | 1561.9 | 7.61% | motif file (matrix) |
pdf |
| 40 |  | Egr1(Zf)/K562-Egr1-ChIP-Seq(GSE32465)/Homer | 1e-9 | -2.099e+01 | 0.0000 | 305.0 | 6.72% | 964.5 | 4.70% | motif file (matrix) |
pdf |
| 41 |  | EKLF(Zf)/Erythrocyte-Klf1-ChIP-Seq(GSE20478)/Homer | 1e-8 | -2.024e+01 | 0.0000 | 116.0 | 2.55% | 287.5 | 1.40% | motif file (matrix) |
pdf |
| 42 |  | HIF-1b(HLH)/T47D-HIF1b-ChIP-Seq(GSE59937)/Homer | 1e-8 | -1.943e+01 | 0.0000 | 1689.0 | 37.19% | 6797.0 | 33.11% | motif file (matrix) |
pdf |
| 43 |  | ZBTB33(Zf)/GM12878-ZBTB33-ChIP-Seq(GSE32465)/Homer | 1e-8 | -1.937e+01 | 0.0000 | 64.0 | 1.41% | 127.0 | 0.62% | motif file (matrix) |
pdf |
| 44 |  | SUT1?/SacCer-Promoters/Homer | 1e-7 | -1.763e+01 | 0.0000 | 3042.0 | 66.99% | 12951.9 | 63.09% | motif file (matrix) |
pdf |
| 45 |  | HIF-1a(bHLH)/MCF7-HIF1a-ChIP-Seq(GSE28352)/Homer | 1e-7 | -1.681e+01 | 0.0000 | 492.0 | 10.83% | 1752.8 | 8.54% | motif file (matrix) |
pdf |
| 46 |  | GAGA-repeat/Arabidopsis-Promoters/Homer | 1e-6 | -1.463e+01 | 0.0000 | 523.0 | 11.52% | 1913.7 | 9.32% | motif file (matrix) |
pdf |
| 47 |  | BATF(bZIP)/Th17-BATF-ChIP-Seq(GSE39756)/Homer | 1e-6 | -1.441e+01 | 0.0000 | 608.0 | 13.39% | 2268.5 | 11.05% | motif file (matrix) |
pdf |
| 48 |  | p53(p53)/Saos-p53-ChIP-Seq(GSE15780)/Homer | 1e-5 | -1.340e+01 | 0.0000 | 83.0 | 1.83% | 214.2 | 1.04% | motif file (matrix) |
pdf |
| 49 |  | p53(p53)/Saos-p53-ChIP-Seq/Homer | 1e-5 | -1.340e+01 | 0.0000 | 83.0 | 1.83% | 214.2 | 1.04% | motif file (matrix) |
pdf |
| 50 |  | PU.1-IRF(ETS:IRF)/Bcell-PU.1-ChIP-Seq(GSE21512)/Homer | 1e-5 | -1.330e+01 | 0.0000 | 1498.0 | 32.99% | 6116.4 | 29.80% | motif file (matrix) |
pdf |
| 51 |  | HIF2a(bHLH)/785_O-HIF2a-ChIP-Seq(GSE34871)/Homer | 1e-5 | -1.242e+01 | 0.0000 | 723.0 | 15.92% | 2791.0 | 13.60% | motif file (matrix) |
pdf |
| 52 |  | FOXP1(Forkhead)/H9-FOXP1-ChIP-Seq(GSE31006)/Homer | 1e-5 | -1.221e+01 | 0.0000 | 802.0 | 17.66% | 3130.1 | 15.25% | motif file (matrix) |
pdf |
| 53 |  | NRF(NRF)/Promoter/Homer | 1e-5 | -1.183e+01 | 0.0001 | 94.0 | 2.07% | 262.8 | 1.28% | motif file (matrix) |
pdf |
| 54 |  | E2F1(E2F)/Hela-E2F1-ChIP-Seq(GSE22478)/Homer | 1e-4 | -9.640e+00 | 0.0005 | 192.0 | 4.23% | 652.8 | 3.18% | motif file (matrix) |
pdf |
| 55 |  | Elk4(ETS)/Hela-Elk4-ChIP-Seq(GSE31477)/Homer | 1e-3 | -8.913e+00 | 0.0009 | 1036.0 | 22.81% | 4227.4 | 20.59% | motif file (matrix) |
pdf |
| 56 |  | CRE(bZIP)/Promoter/Homer | 1e-3 | -8.844e+00 | 0.0010 | 200.0 | 4.40% | 694.9 | 3.39% | motif file (matrix) |
pdf |
| 57 |  | E2F7(E2F)/Hela-E2F7-ChIP-Seq(GSE32673)/Homer | 1e-3 | -8.729e+00 | 0.0011 | 123.0 | 2.71% | 395.6 | 1.93% | motif file (matrix) |
pdf |
| 58 |  | BORIS(Zf)/K562-CTCFL-ChIP-Seq(GSE32465)/Homer | 1e-3 | -8.564e+00 | 0.0013 | 45.0 | 0.99% | 113.1 | 0.55% | motif file (matrix) |
pdf |
| 59 |  | TATA-box/Drosophila-Promoters/Homer | 1e-3 | -7.847e+00 | 0.0026 | 228.0 | 5.02% | 821.4 | 4.00% | motif file (matrix) |
pdf |
| 60 |  | Elk1(ETS)/Hela-Elk1-ChIP-Seq(GSE31477)/Homer | 1e-3 | -7.809e+00 | 0.0026 | 851.0 | 18.74% | 3458.7 | 16.85% | motif file (matrix) |
pdf |
| 61 |  | SeqBias: TA-repeat | 1e-3 | -7.451e+00 | 0.0037 | 4008.0 | 88.26% | 17785.2 | 86.64% | motif file (matrix) |
pdf |
| 62 |  | Zic(Zf)/Cerebellum-ZIC1.2-ChIP-Seq(GSE60731)/Homer | 1e-3 | -7.296e+00 | 0.0042 | 420.0 | 9.25% | 1627.4 | 7.93% | motif file (matrix) |
pdf |
| 63 |  | CTCF(Zf)/CD4+-CTCF-ChIP-Seq(Barski_et_al.)/Homer | 1e-3 | -7.244e+00 | 0.0044 | 46.0 | 1.01% | 124.3 | 0.61% | motif file (matrix) |
pdf |
| 64 |  | PAX5(Paired,Homeobox)/GM12878-PAX5-ChIP-Seq(GSE32465)/Homer | 1e-3 | -7.140e+00 | 0.0048 | 220.0 | 4.84% | 800.2 | 3.90% | motif file (matrix) |
pdf |
| 65 |  | RXR(NR),DR1/3T3L1-RXR-ChIP-Seq(GSE13511)/Homer | 1e-3 | -7.086e+00 | 0.0050 | 513.0 | 11.30% | 2026.7 | 9.87% | motif file (matrix) |
pdf |
| 66 |  | Tbox:Smad(T-box,MAD)/ESCd5-Smad2_3-ChIP-Seq(GSE29422)/Homer | 1e-2 | -6.762e+00 | 0.0068 | 113.0 | 2.49% | 378.1 | 1.84% | motif file (matrix) |
pdf |
| 67 |  | E2F(E2F)/Hela-CellCycle-Expression/Homer | 1e-2 | -6.732e+00 | 0.0069 | 129.0 | 2.84% | 441.9 | 2.15% | motif file (matrix) |
pdf |
| 68 |  | NRF1(NRF)/MCF7-NRF1-ChIP-Seq(Unpublished)/Homer | 1e-2 | -6.619e+00 | 0.0076 | 42.0 | 0.92% | 114.5 | 0.56% | motif file (matrix) |
pdf |
| 69 |  | Fra1(bZIP)/BT549-Fra1-ChIP-Seq(GSE46166)/Homer | 1e-2 | -6.615e+00 | 0.0076 | 511.0 | 11.25% | 2030.8 | 9.89% | motif file (matrix) |
pdf |
| 70 |  | Dorsal(RHD)/Embryo-dl-ChIP-Seq(GSE65441)/Homer | 1e-2 | -6.092e+00 | 0.0125 | 208.0 | 4.58% | 770.0 | 3.75% | motif file (matrix) |
pdf |
| 71 |  | SpiB(ETS)/OCILY3-SPIB-ChIP-Seq(GSE56857)/Homer | 1e-2 | -6.078e+00 | 0.0125 | 279.0 | 6.14% | 1063.2 | 5.18% | motif file (matrix) |
pdf |
| 72 | 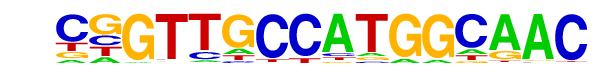 | RFX(HTH)/K562-RFX3-ChIP-Seq(SRA012198)/Homer | 1e-2 | -5.955e+00 | 0.0139 | 77.0 | 1.70% | 249.0 | 1.21% | motif file (matrix) |
pdf |
| 73 |  | AP-1(bZIP)/ThioMac-PU.1-ChIP-Seq(GSE21512)/Homer | 1e-2 | -5.538e+00 | 0.0209 | 614.0 | 13.52% | 2505.5 | 12.21% | motif file (matrix) |
pdf |
| 74 |  | PAX5(Paired,Homeobox),condensed/GM12878-PAX5-ChIP-Seq(GSE32465)/Homer | 1e-2 | -5.506e+00 | 0.0213 | 75.0 | 1.65% | 245.2 | 1.19% | motif file (matrix) |
pdf |
| 75 |  | Unknown3/Arabidopsis-Promoters/Homer | 1e-2 | -5.467e+00 | 0.0218 | 534.0 | 11.76% | 2163.0 | 10.54% | motif file (matrix) |
pdf |
| 76 |  | Pax8(Paired,Homeobox)/Thyroid-Pax8-ChIP-Seq(GSE26938)/Homer | 1e-2 | -5.341e+00 | 0.0244 | 122.0 | 2.69% | 432.5 | 2.11% | motif file (matrix) |
pdf |
| 77 |  | Myf5(bHLH)/GM-Myf5-ChIP-Seq(GSE24852)/Homer | 1e-2 | -5.330e+00 | 0.0244 | 358.0 | 7.88% | 1413.3 | 6.88% | motif file (matrix) |
pdf |
| 78 |  | ETS(ETS)/Promoter/Homer | 1e-2 | -5.212e+00 | 0.0270 | 333.0 | 7.33% | 1310.8 | 6.39% | motif file (matrix) |
pdf |
| 79 |  | Unknown-ESC-element(?)/mES-Nanog-ChIP-Seq(GSE11724)/Homer | 1e-2 | -5.105e+00 | 0.0297 | 181.0 | 3.99% | 676.5 | 3.30% | motif file (matrix) |
pdf |
| 80 |  | PPARE(NR),DR1/3T3L1-Pparg-ChIP-Seq(GSE13511)/Homer | 1e-2 | -4.917e+00 | 0.0354 | 484.0 | 10.66% | 1964.3 | 9.57% | motif file (matrix) |
pdf |
| 81 |  | ZNF692(Zf)/HEK293-ZNF692.GFP-ChIP-Seq(GSE58341)/Homer | 1e-2 | -4.754e+00 | 0.0412 | 51.0 | 1.12% | 161.2 | 0.79% | motif file (matrix) |
pdf |

































































































































































