| pattern |
num_seqlets |
modisco_cwm_fwd |
modisco_cwm_rev |
match0 |
qval0 |
match0_logo |
match1 |
qval1 |
match1_logo |
match2 |
qval2 |
match2_logo |
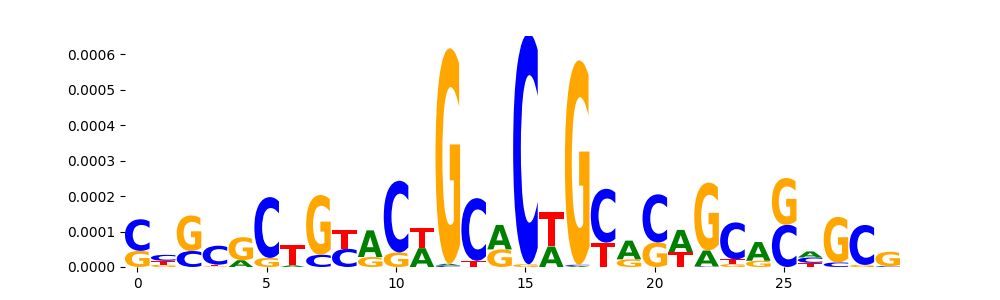
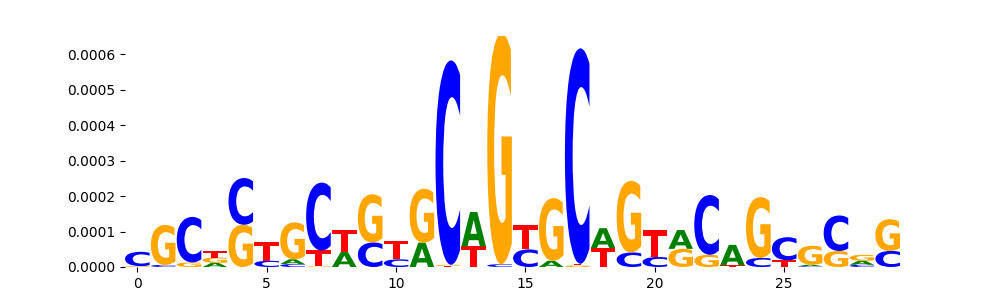
| pos_patterns.pattern_0 |
3587 |
 |
 |
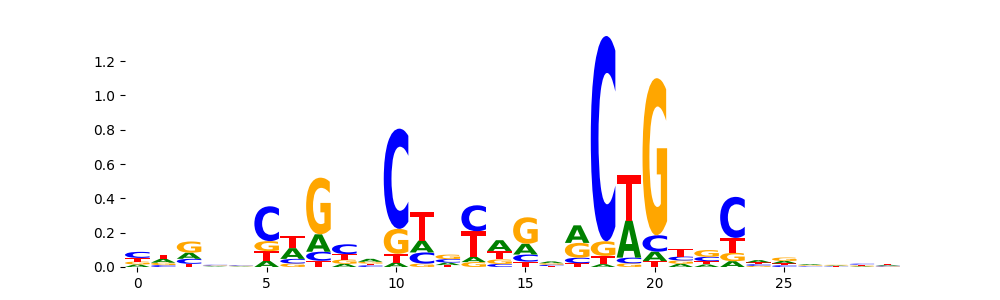
TN5_7 |
1.072620e-03 |
 |
TN5_4 |
4.896430e-02 |
 |
TN5_5 |
4.896430e-02 |
 |
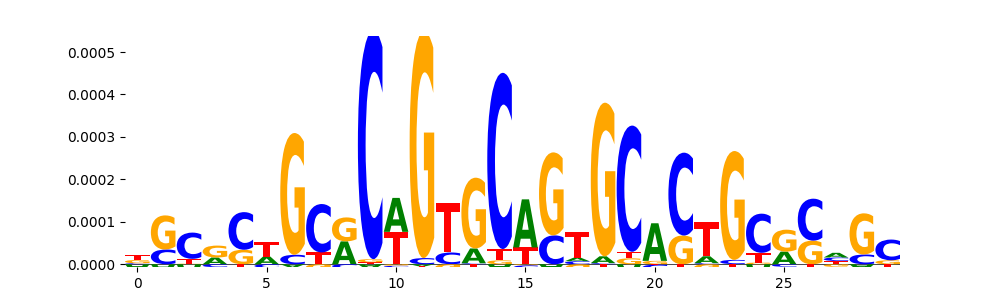
| pos_patterns.pattern_1 |
2787 |
 |
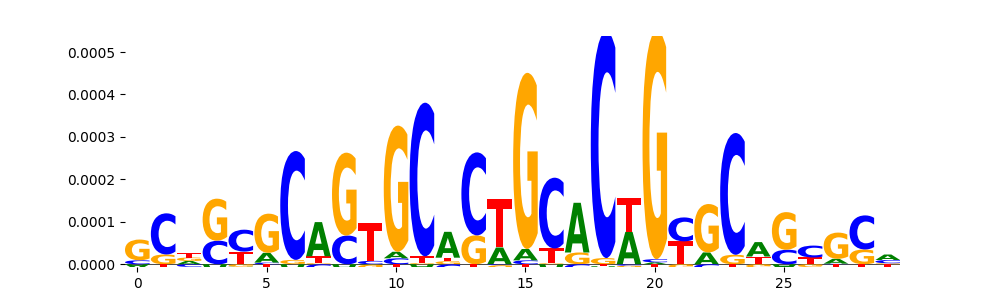 |
TN5_4 |
6.511700e-06 |
 |
TN5_5 |
6.511700e-06 |
 |
TN5_2 |
1.150090e-05 |
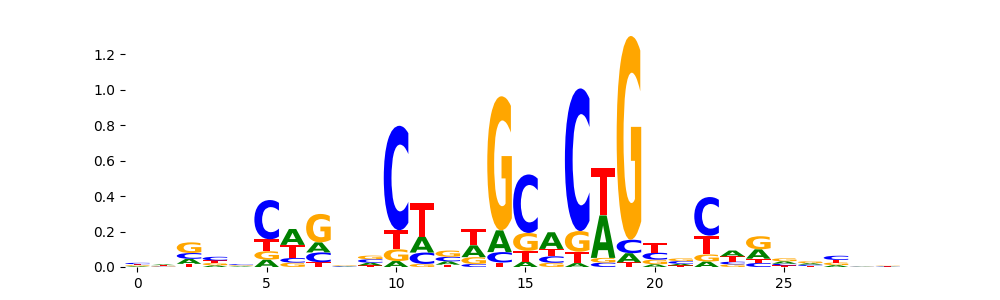 |
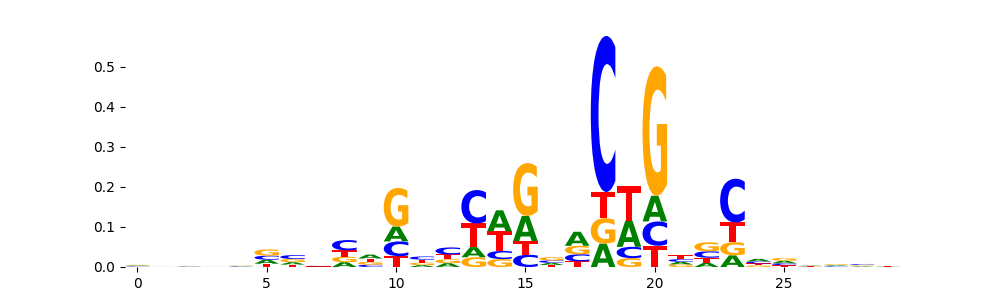
| pos_patterns.pattern_2 |
1705 |
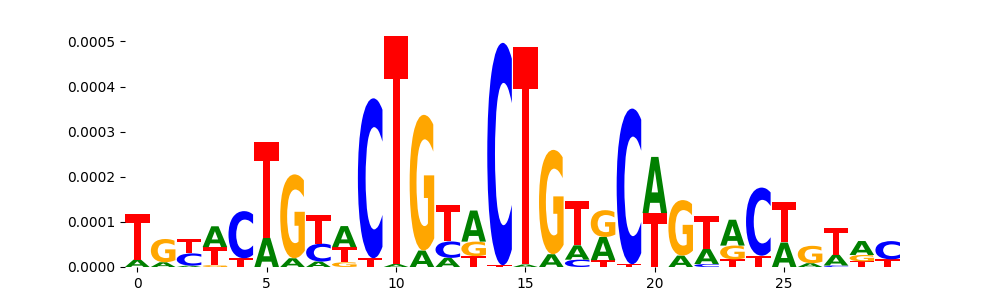 |
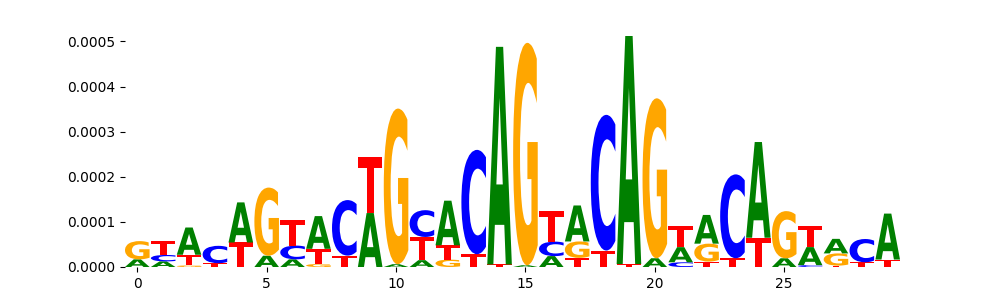 |
TN5_1 |
2.856130e-03 |
 |
TN5_2 |
2.937620e-02 |
 |
TN5_7 |
1.684700e-01 |
 |
| pos_patterns.pattern_3 |
1160 |
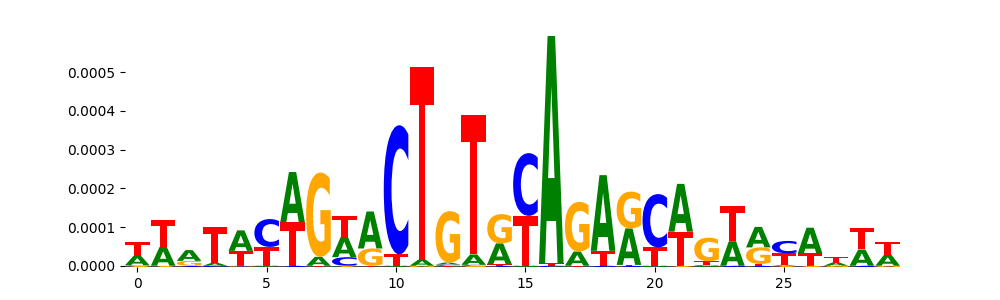 |
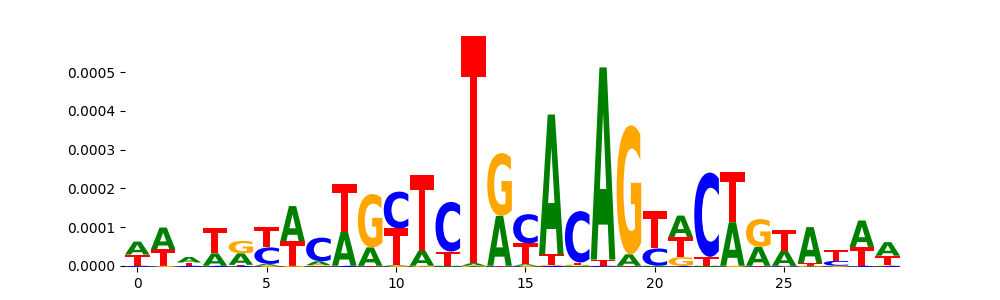 |
TN5_2 |
1.036130e-03 |
 |
TN5_1 |
6.962290e-03 |
 |
TN5_7 |
1.980070e-02 |
 |
| pos_patterns.pattern_4 |
594 |
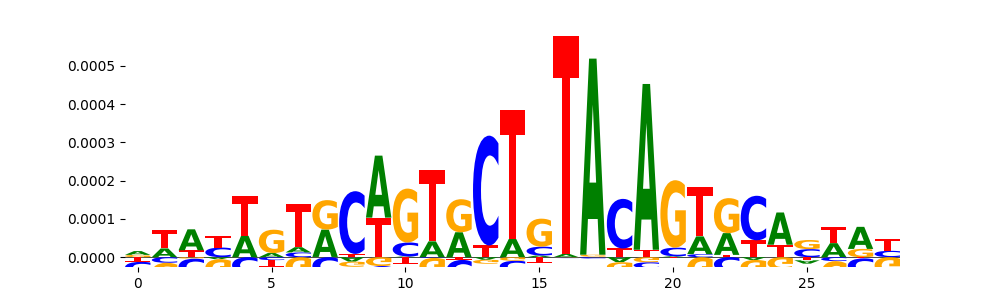 |
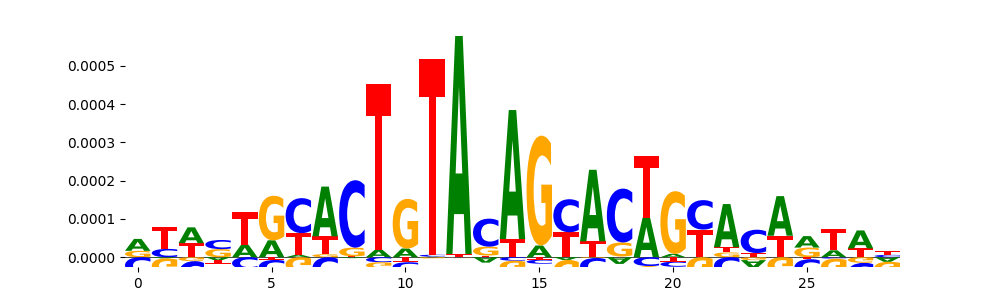 |
TN5_4 |
4.425570e-07 |
 |
TN5_5 |
4.425570e-07 |
 |
TN5_7 |
2.303480e-03 |
 |
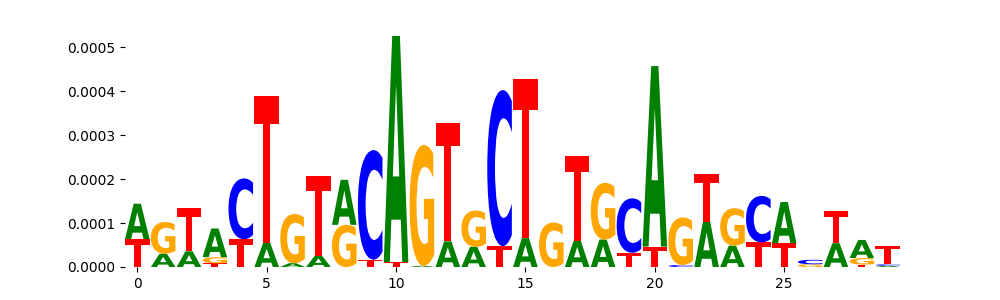
| pos_patterns.pattern_5 |
539 |
 |
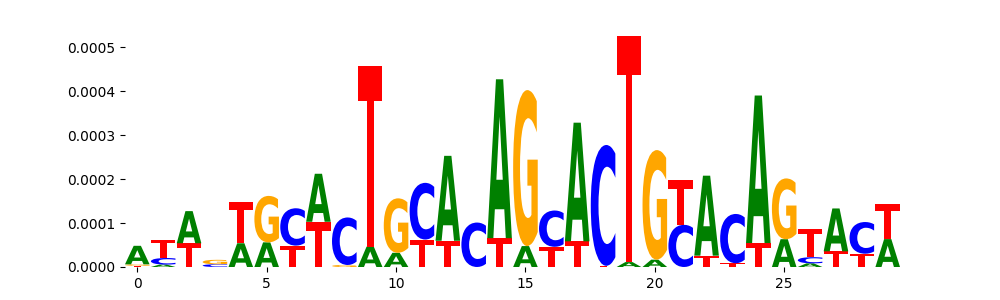 |
TN5_2 |
2.099660e-03 |
 |
TN5_7 |
3.056500e-03 |
 |
TN5_1 |
6.090100e-02 |
 |
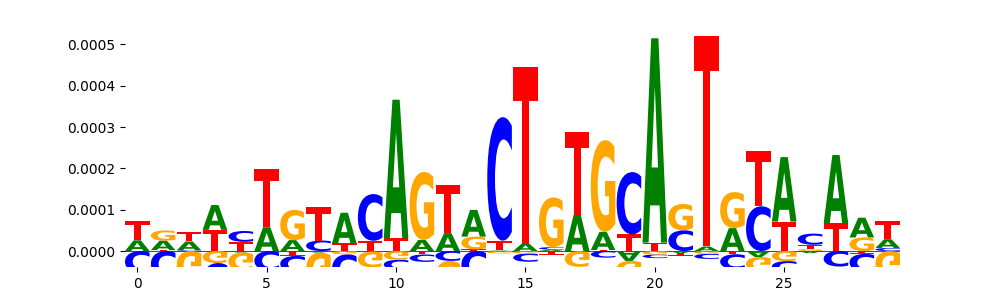
| pos_patterns.pattern_6 |
444 |
 |
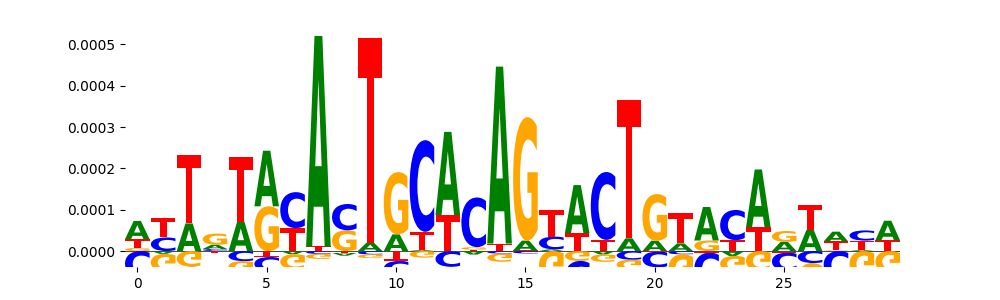 |
TN5_2 |
4.973120e-04 |
 |
TN5_4 |
1.168490e-02 |
 |
TN5_5 |
1.168490e-02 |
 |
| pos_patterns.pattern_7 |
386 |
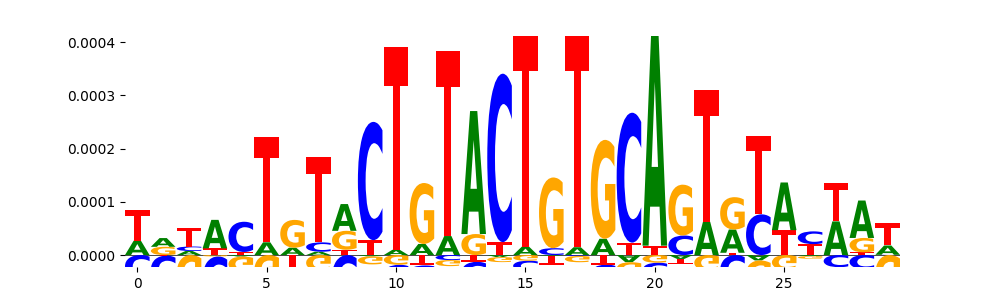 |
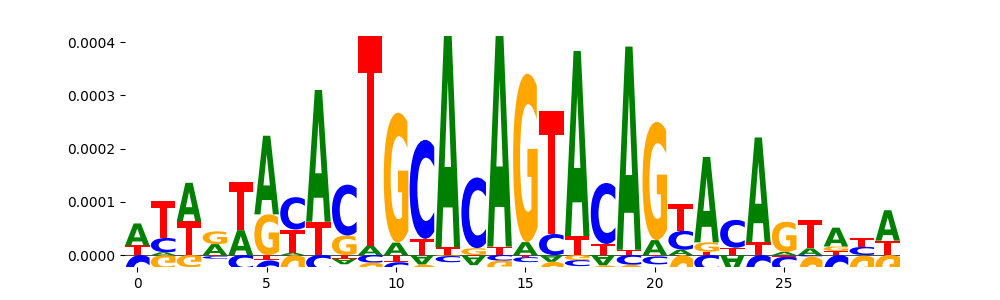 |
TN5_2 |
2.261420e-03 |
 |
TN5_1 |
2.529030e-01 |
 |
TN5_4 |
2.529030e-01 |
 |
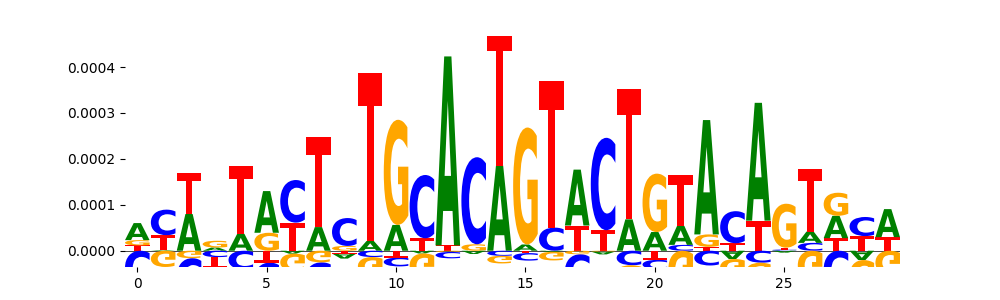
| pos_patterns.pattern_8 |
344 |
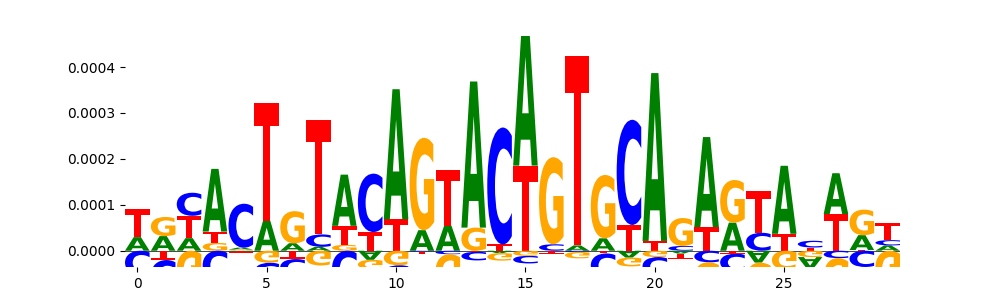 |
 |
TN5_7 |
2.770720e-03 |
 |
TN5_1 |
2.770720e-03 |
 |
TN5_2 |
2.833740e-03 |
 |
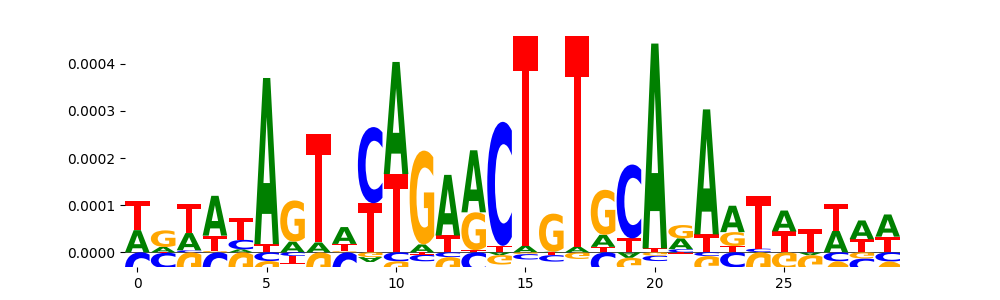
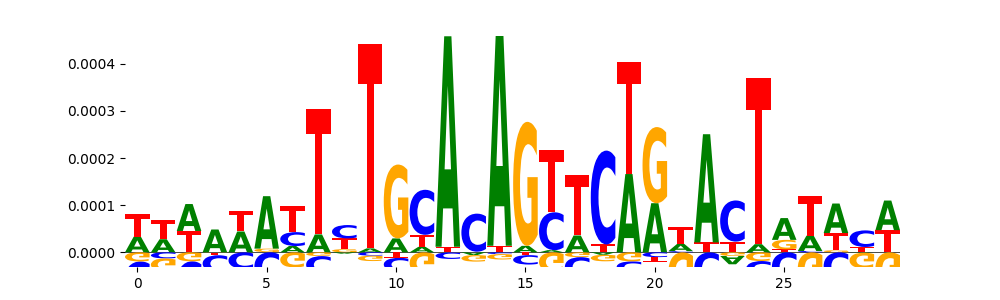
| pos_patterns.pattern_9 |
268 |
 |
 |
TN5_7 |
2.846160e-04 |
 |
TN5_2 |
1.021950e-03 |
 |
TN5_1 |
1.021950e-03 |
 |
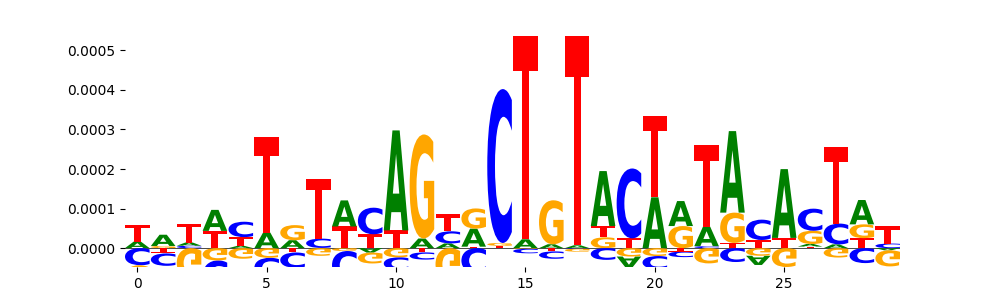
| pos_patterns.pattern_10 |
227 |
 |
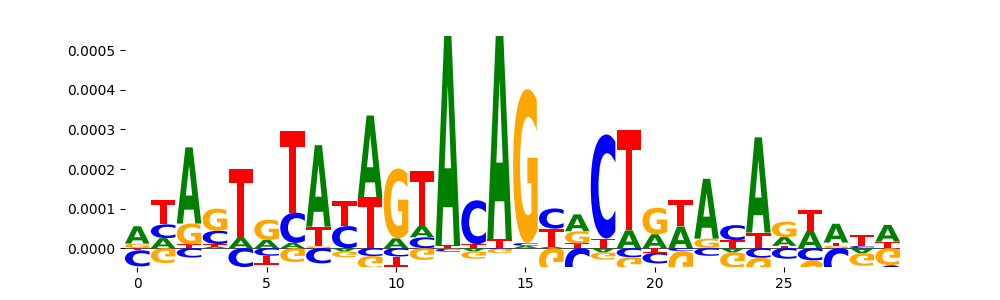 |
TN5_2 |
5.055420e-04 |
 |
TN5_1 |
1.158160e-01 |
 |
TN5_7 |
3.366560e-01 |
 |
| pos_patterns.pattern_11 |
149 |
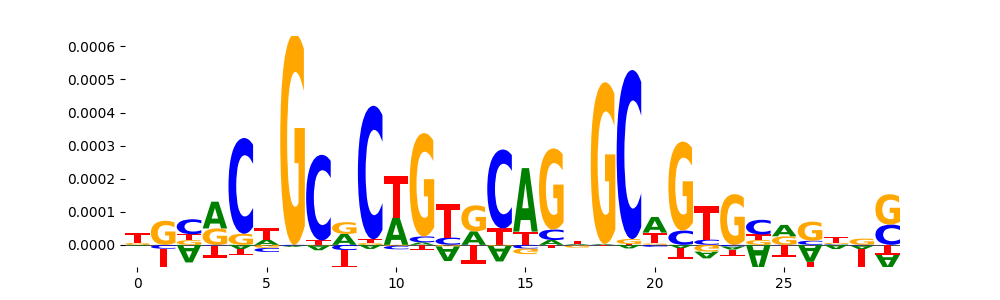 |
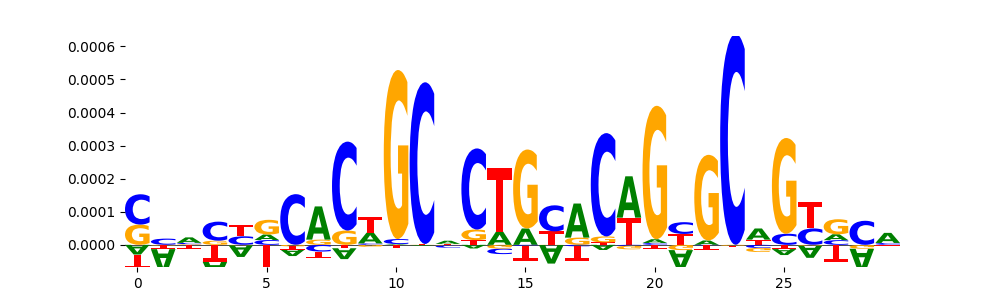 |
TN5_2 |
1.064150e-01 |
 |
TN5_1 |
1.064150e-01 |
 |
TBX1_TBX_4 |
1.064150e-01 |
 |
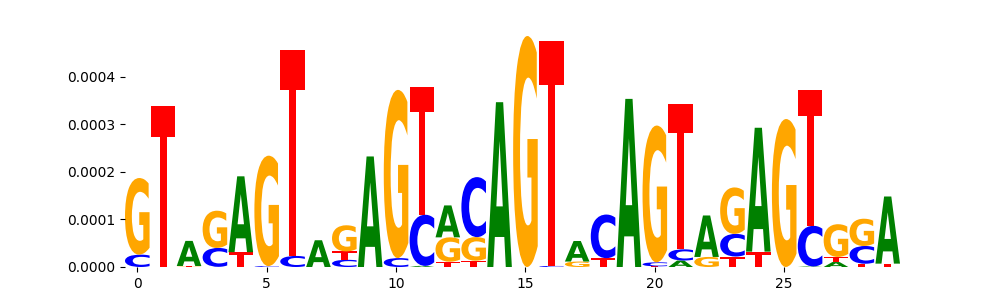
| pos_patterns.pattern_12 |
52 |
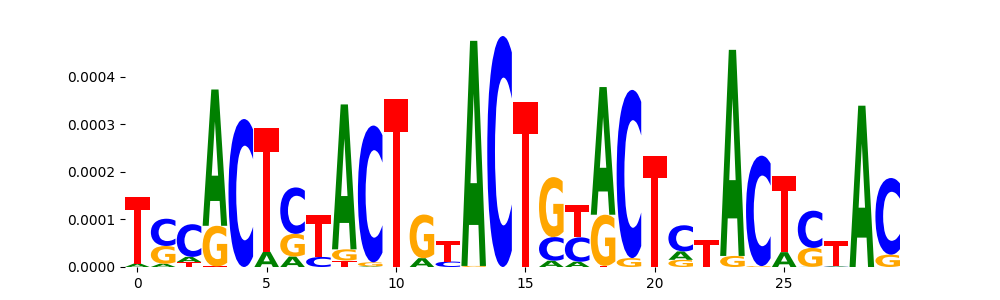 |
 |
ZN502_HUMAN.H11MO.0.C |
8.435460e-02 |
 |
SMCA5_MOUSE.H11MO.0.C |
8.435460e-02 |
 |
ZN394_HUMAN.H11MO.0.C |
1.728940e-01 |
 |
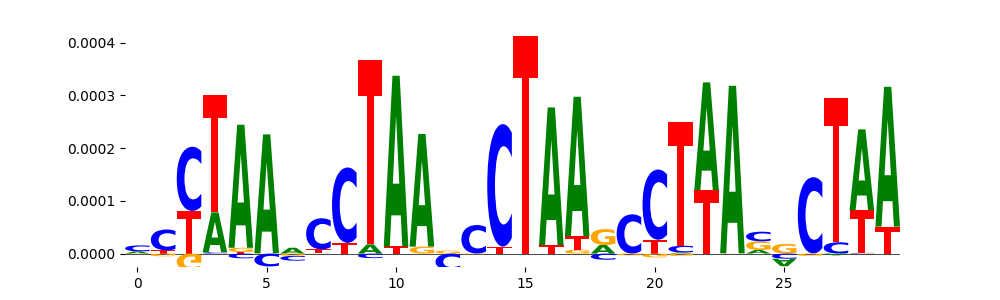
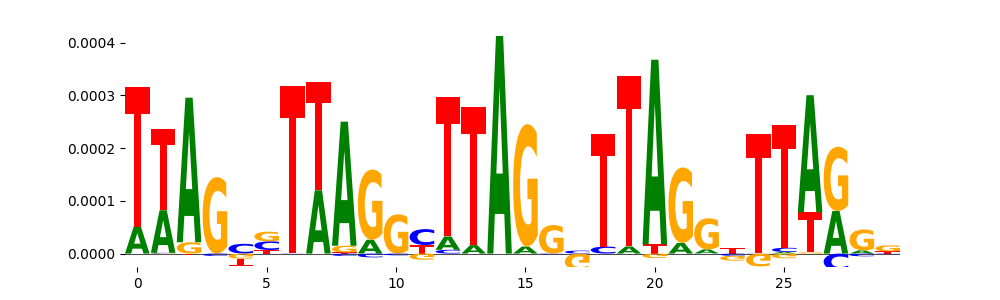
| pos_patterns.pattern_13 |
26 |
 |
 |
RREB1_MA0073.1 |
1.000000e+00 |
 |
ZN281_MOUSE.H11MO.0.A |
1.000000e+00 |
 |
SRBP2_HUMAN.H11MO.0.B |
1.000000e+00 |
 |
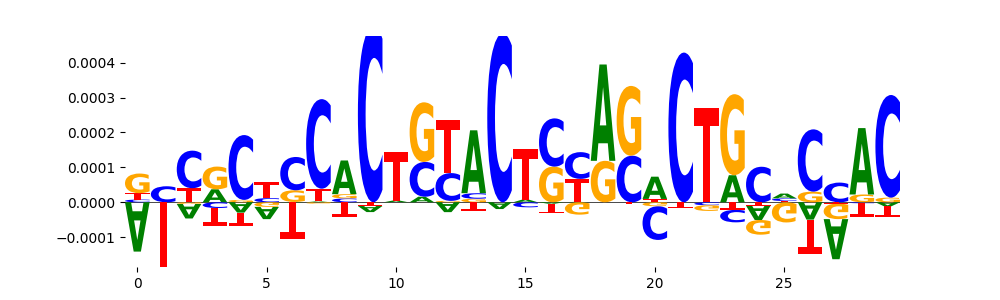
| pos_patterns.pattern_14 |
26 |
 |
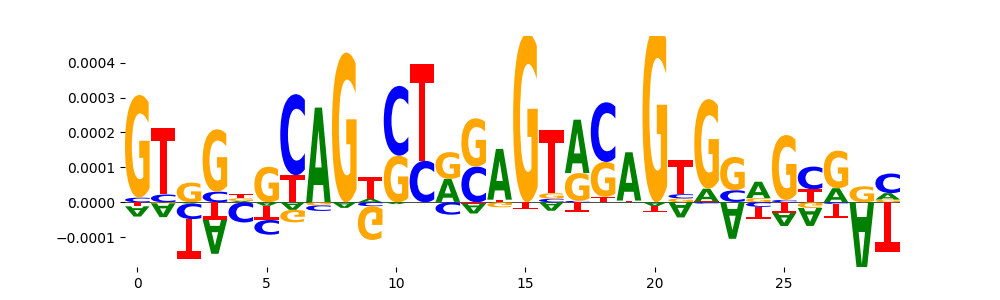 |
TN5_6 |
2.567000e-13 |
 |
TN5_4 |
2.040810e-01 |
 |
TN5_5 |
2.040810e-01 |
 |
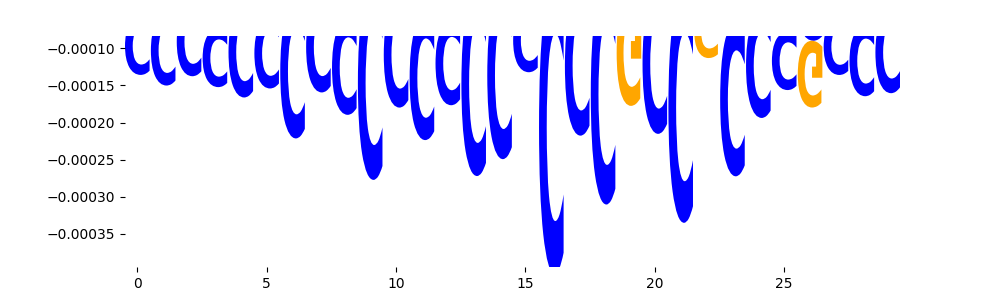
| neg_patterns.pattern_0 |
4777 |
 |
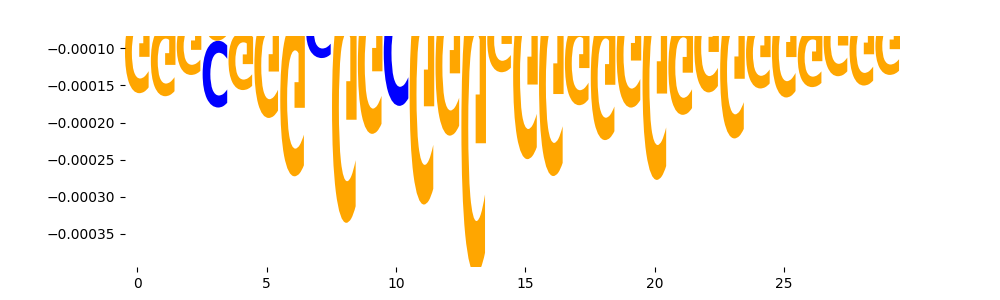 |
SP2_HUMAN.H11MO.0.A |
1.702880e-08 |
 |
SP2_MOUSE.H11MO.0.B |
1.702880e-08 |
 |
SP1_HUMAN.H11MO.0.A |
1.089330e-07 |
 |
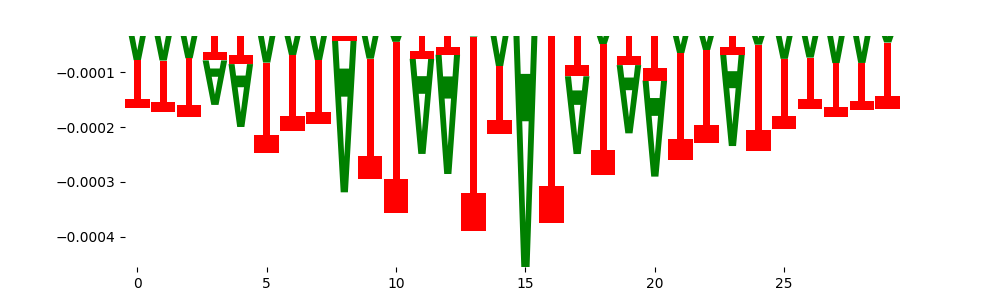
| neg_patterns.pattern_1 |
4188 |
 |
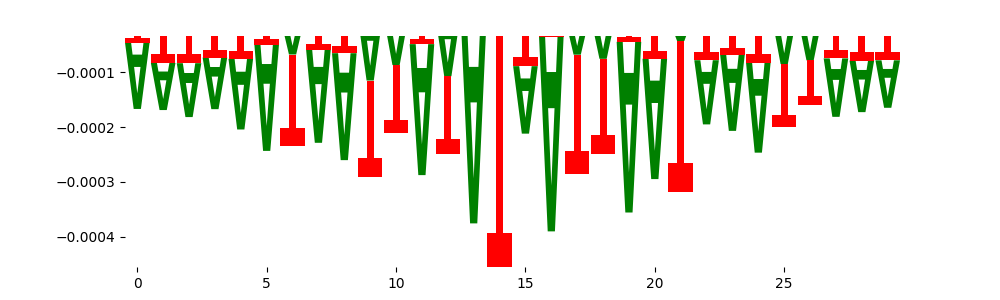 |
DNASE_2 |
1.416810e-01 |
 |
Arid3b_MA0601.1 |
1.416810e-01 |
 |
LHX3_HUMAN.H11MO.0.C |
1.416810e-01 |
 |
| neg_patterns.pattern_2 |
3421 |
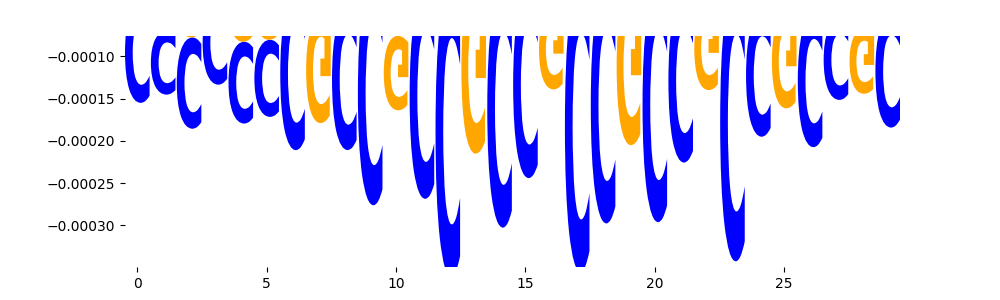 |
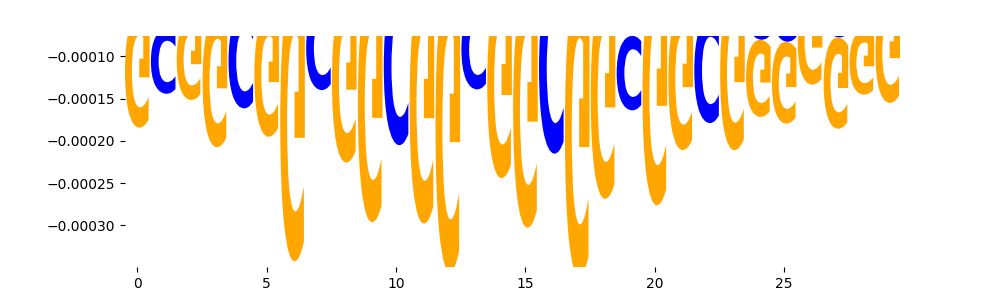 |
SP2_HUMAN.H11MO.0.A |
1.480960e-06 |
 |
SP2_MOUSE.H11MO.0.B |
1.480960e-06 |
 |
SP1_MOUSE.H11MO.0.A |
3.261450e-06 |
 |
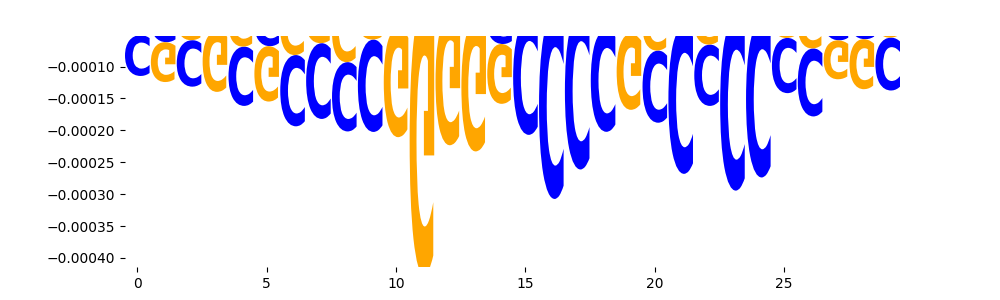
| neg_patterns.pattern_3 |
3333 |
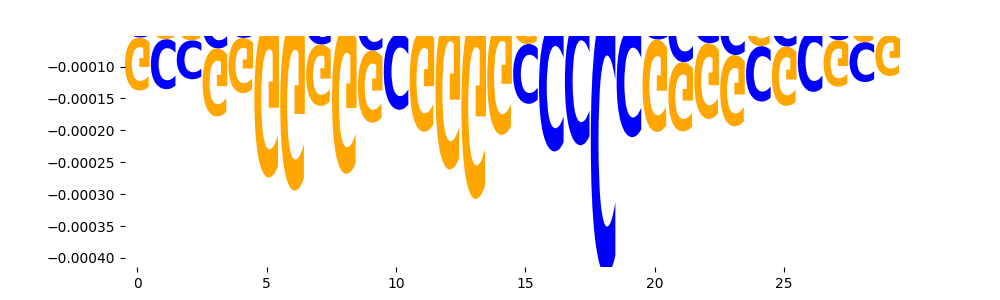 |
 |
SP2_HUMAN.H11MO.0.A |
6.522630e-08 |
 |
SP2_MOUSE.H11MO.0.B |
6.522630e-08 |
 |
SP1_HUMAN.H11MO.0.A |
2.651530e-06 |
 |
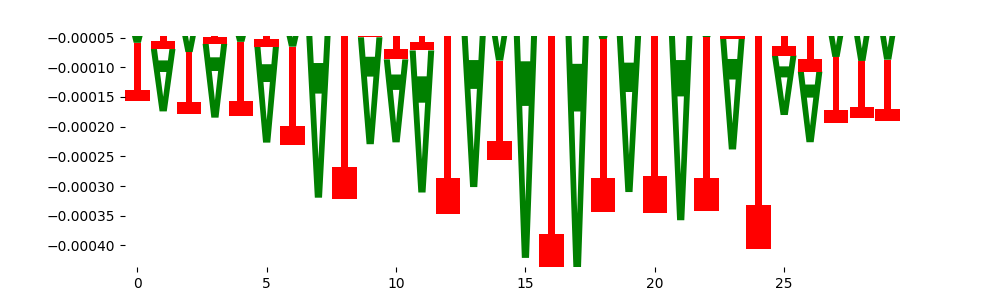
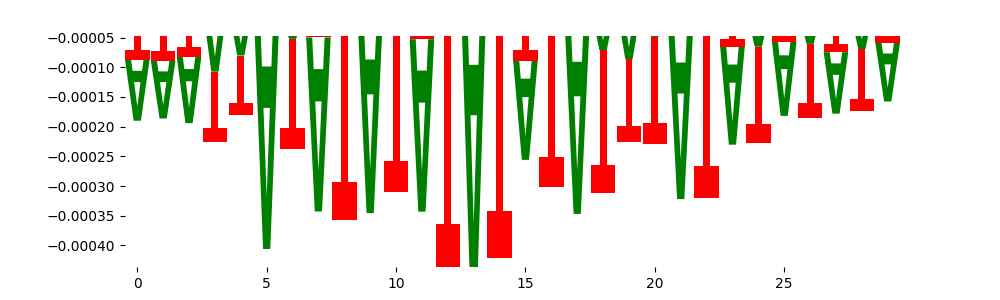
| neg_patterns.pattern_4 |
1646 |
 |
 |
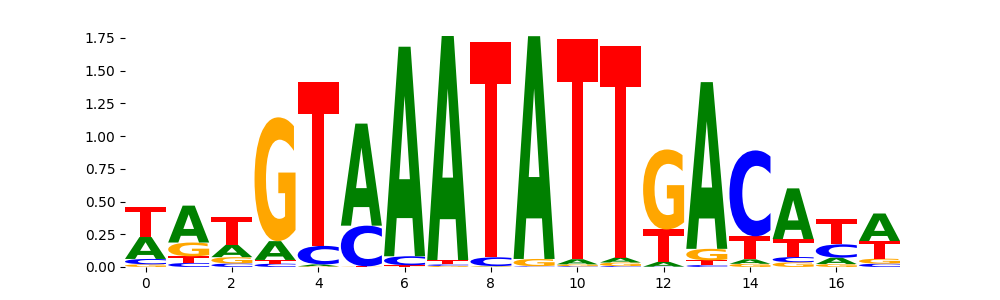
FOXB1_forkhead_2 |
9.242280e-02 |
 |
DNASE_2 |
1.792880e-01 |
 |
FOXD2_forkhead_1 |
1.890470e-01 |
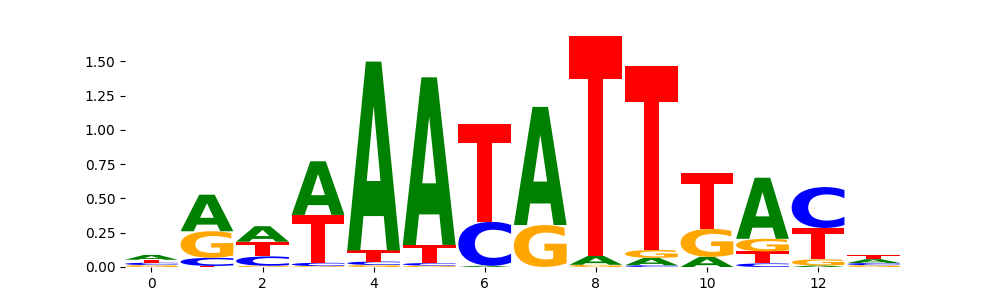 |
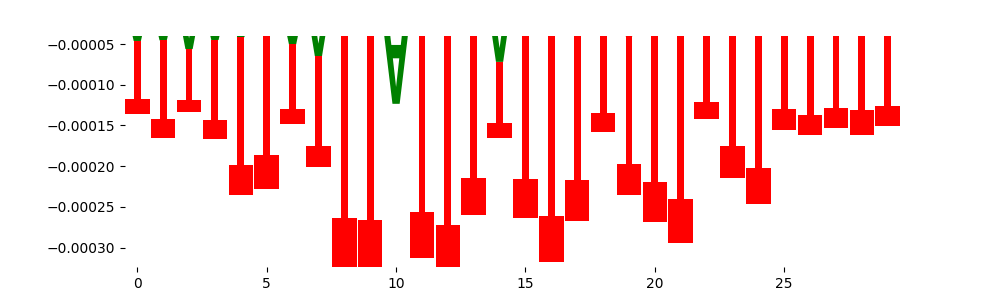
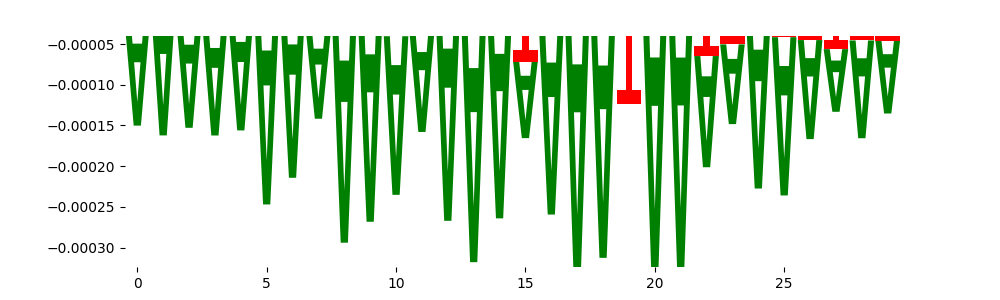
| neg_patterns.pattern_5 |
861 |
 |
 |
PRDM6_HUMAN.H11MO.0.C |
4.149380e-02 |
 |
ZNF384_MA1125.1 |
9.047700e-02 |
 |
STAT1_MOUSE.H11MO.0.A |
9.047700e-02 |
 |
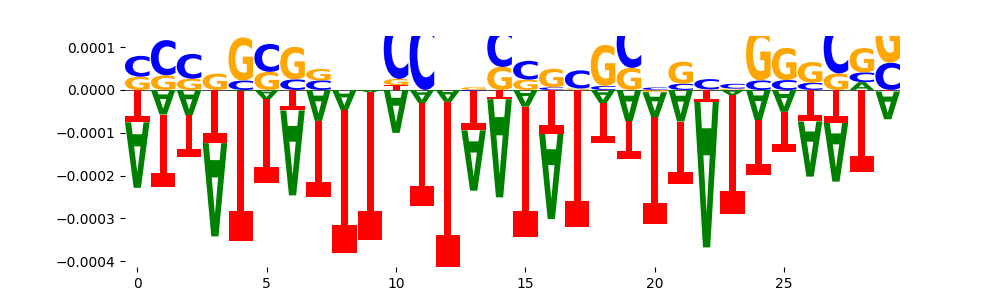
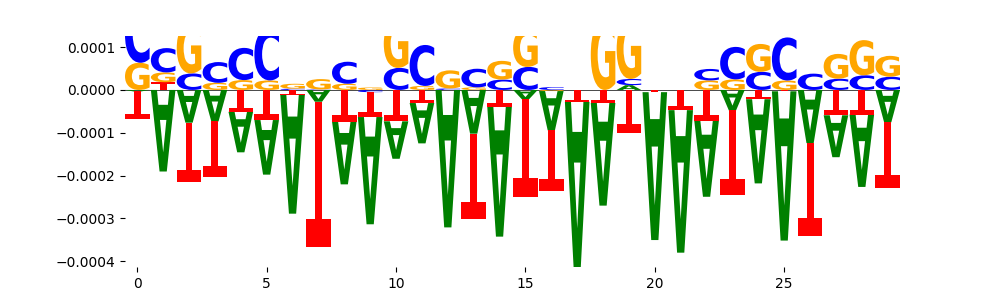
| neg_patterns.pattern_6 |
37 |
 |
 |
Arid3b_MA0601.1 |
2.945860e-01 |
 |
ZNF384_MA1125.1 |
2.945860e-01 |
 |
ONECUT3_CUT_1 |
3.001360e-01 |
 |
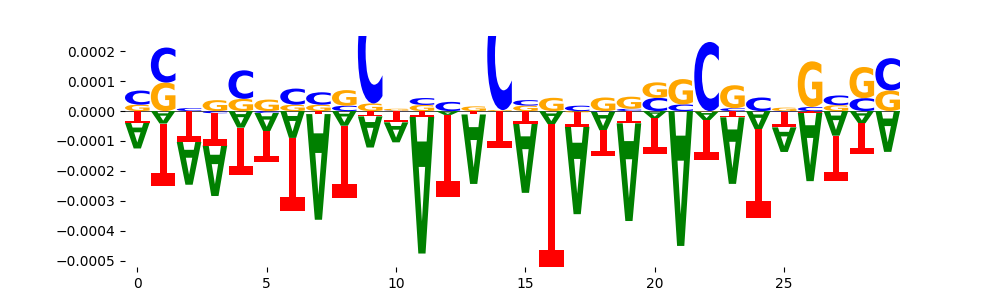
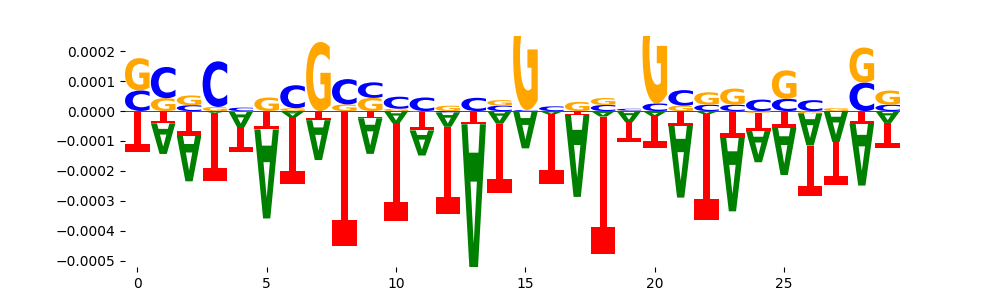
| neg_patterns.pattern_7 |
33 |
 |
 |
DNASE_2 |
9.588780e-01 |
 |
FOXB1_forkhead_2 |
9.588780e-01 |
 |
CDX1_HUMAN.H11MO.0.C |
9.588780e-01 |
 |























































































































