| p-value: | 1e-4 |
| log p-value: | -9.911e+00 |
| Information Content per bp: | 1.530 |
| Number of Target Sequences with motif | 1.0 |
| Percentage of Target Sequences with motif | 25.00% |
| Number of Background Sequences with motif | 0.0 |
| Percentage of Background Sequences with motif | 0.00% |
| Average Position of motif in Targets | 14.0 +/- 0.0bp |
| Average Position of motif in Background | 0.0 +/- 0.0bp |
| Strand Bias (log2 ratio + to - strand density) | 10.0 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
Nr5a2(NR)/mES-Nr5a2-ChIP-Seq(GSE19019)/Homer
| Match Rank: | 1 |
| Score: | 0.61
| | Offset: | -3
| | Orientation: | reverse strand |
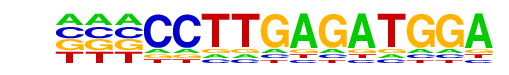
| Alignment: | ---CCTTGAGATGGA
TGACCTTGAN----- |
|


|
|
Nr5a2(NR)/Pancreas-LRH1-ChIP-Seq(GSE34295)/Homer
| Match Rank: | 2 |
| Score: | 0.59
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---CCTTGAGATGGA
TGACCTTGAV----- |
|


|
|
MA0505.1_Nr5a2/Jaspar
| Match Rank: | 3 |
| Score: | 0.59
| | Offset: | -5
| | Orientation: | reverse strand |
| Alignment: | -----CCTTGAGATGGA
GCTGACCTTGAACTN-- |
|


|
|
MA0142.1_Pou5f1::Sox2/Jaspar
| Match Rank: | 4 |
| Score: | 0.59
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CCTTGAGATGGA---
CTTTGTTATGCAAAT |
|


|
|
OCT4-SOX2-TCF-NANOG(POU,Homeobox,HMG)/mES-Oct4-ChIP-Seq(GSE11431)/Homer
| Match Rank: | 5 |
| Score: | 0.56
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CCTTGAGATGGA---
CATTGTTATGCAAAT |
|


|
|
RARg(NR)/ES-RARg-ChIP-Seq(GSE30538)/Homer
| Match Rank: | 6 |
| Score: | 0.55
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---CCTTGAGATGGA
TGACCTTGACCT--- |
|


|
|
PB0098.1_Zfp410_1/Jaspar
| Match Rank: | 7 |
| Score: | 0.55
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --CCTTGAGATGGA---
TATTATGGGATGGATAA |
|


|
|
MA0141.2_Esrrb/Jaspar
| Match Rank: | 8 |
| Score: | 0.54
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---CCTTGAGATGGA
TGACCTTGANNN--- |
|

|
|
HOXA2(Homeobox)/mES-Hoxa2-ChIP-Seq(Donaldson et al.)/Homer
| Match Rank: | 9 |
| Score: | 0.54
| | Offset: | 4
| | Orientation: | reverse strand |
| Alignment: | CCTTGAGATGGA----
----ATGATKGATGRC |
|


|
|
Tcf3(HMG)/mES-Tcf3-ChIP-Seq(GSE11724)/Homer
| Match Rank: | 10 |
| Score: | 0.53
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -CCTTGAGATGGA
CCTTTGATGT--- |
|


|
|





















