HiCExplorer QC report
Pairs sequenced
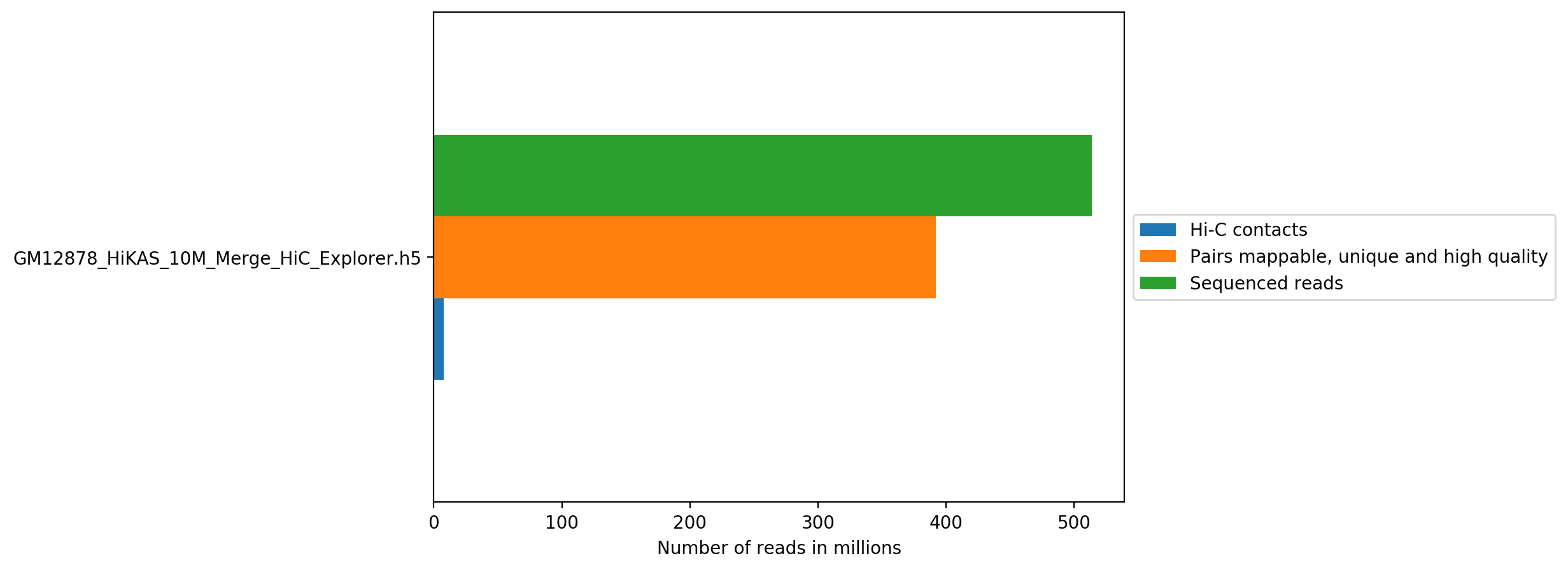
Unmappable and non-unique pairs
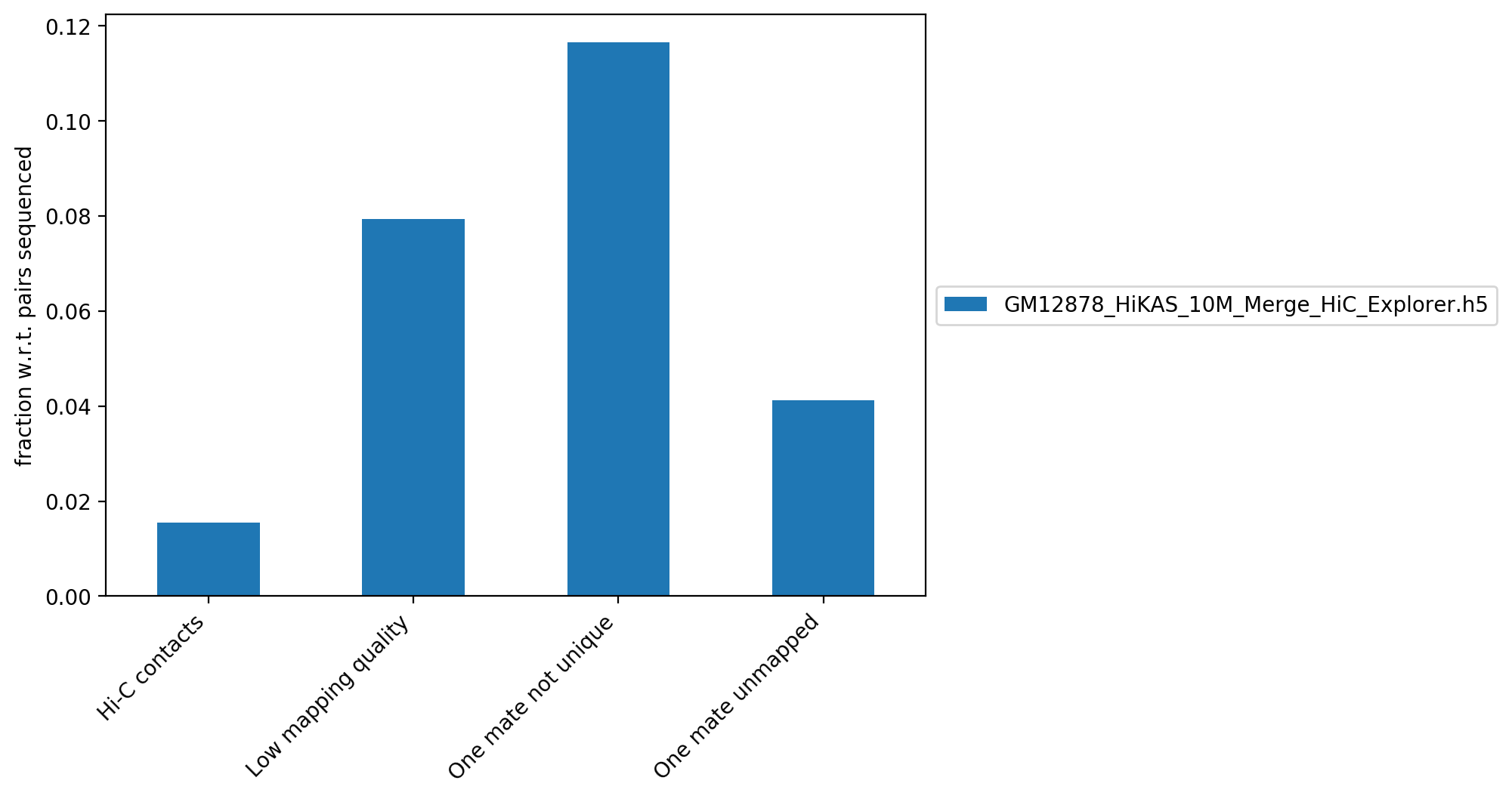
This figure contains the fraction of reads, with respect to the total number of reads, that did not map, that have a low quality score or that didn't map uniquely to the genome.
Unmappable and non-unique table
| Hi-C contacts | Hi-C contacts_% | Low mapping quality | Low mapping quality_% | One mate not unique | One mate not unique_% | One mate unmapped | One mate unmapped_% | |
|---|---|---|---|---|---|---|---|---|
| File | ||||||||
| GM12878_HiKAS_10M_Merge_HiC_Explorer.h5 | 7,963,164 | 1.55% | 40,771,155 | 7.94% | 59,841,028 | 11.65% | 21,158,380 | 4.12% |
Pairs discarded
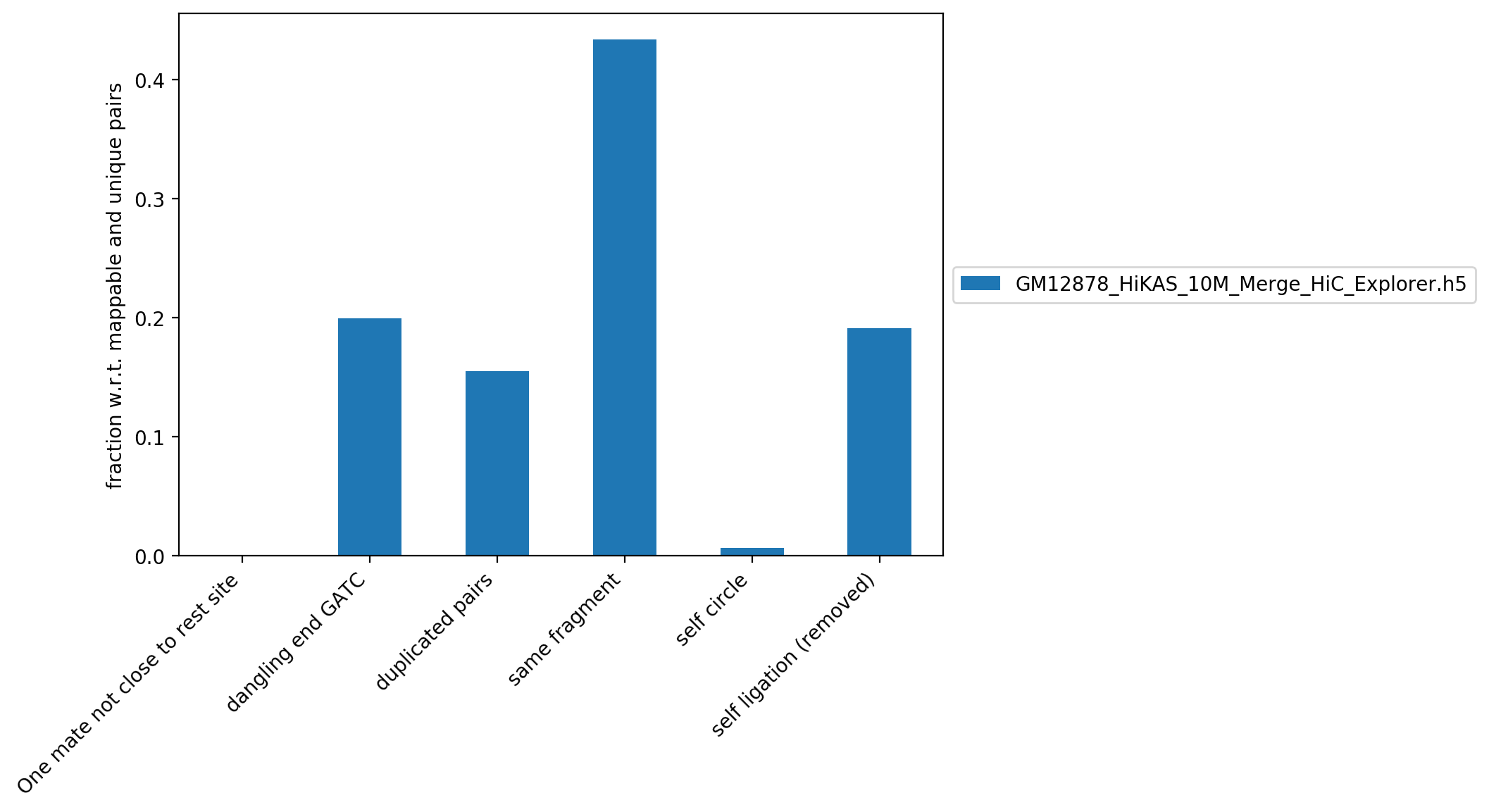
This figure contains the fraction read pairs (with respect to mappable reads) that were discarded when building the Hi-C matrix.
- Dangling ends
- These are reads that start with the restriction site and constitute reads that were digested but no ligated.
- Same fragment
- These are read mates, facing inward, separated by up to 800 bp that do not have a restriction enzyme in between. These read pairs are not valid Hi-C pairs
- Self circle
- self circles are defined as pairs within 25kb with 'outward' read orientation
- Self ligation
- These are read pairs with a restriction site in between that are within 800 bp.
Discarded pairs table
| One mate not close to rest site | One mate not close to rest site % | dangling end GATC % | duplicated pairs | duplicated pairs % | same fragment | same fragment % | self circle | self circle % | self ligation (removed) | self ligation (removed) % | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| File | |||||||||||
| GM12878_HiKAS_10M_Merge_HiC_Explorer.h5 | 0.00% | 0.00% | 19.95% | 60,704,381 | 15.49% | 170,003,494 | 43.38% | 2,532,589 | 0.65% | 75,009,486 | 19.14% |
Contact distance
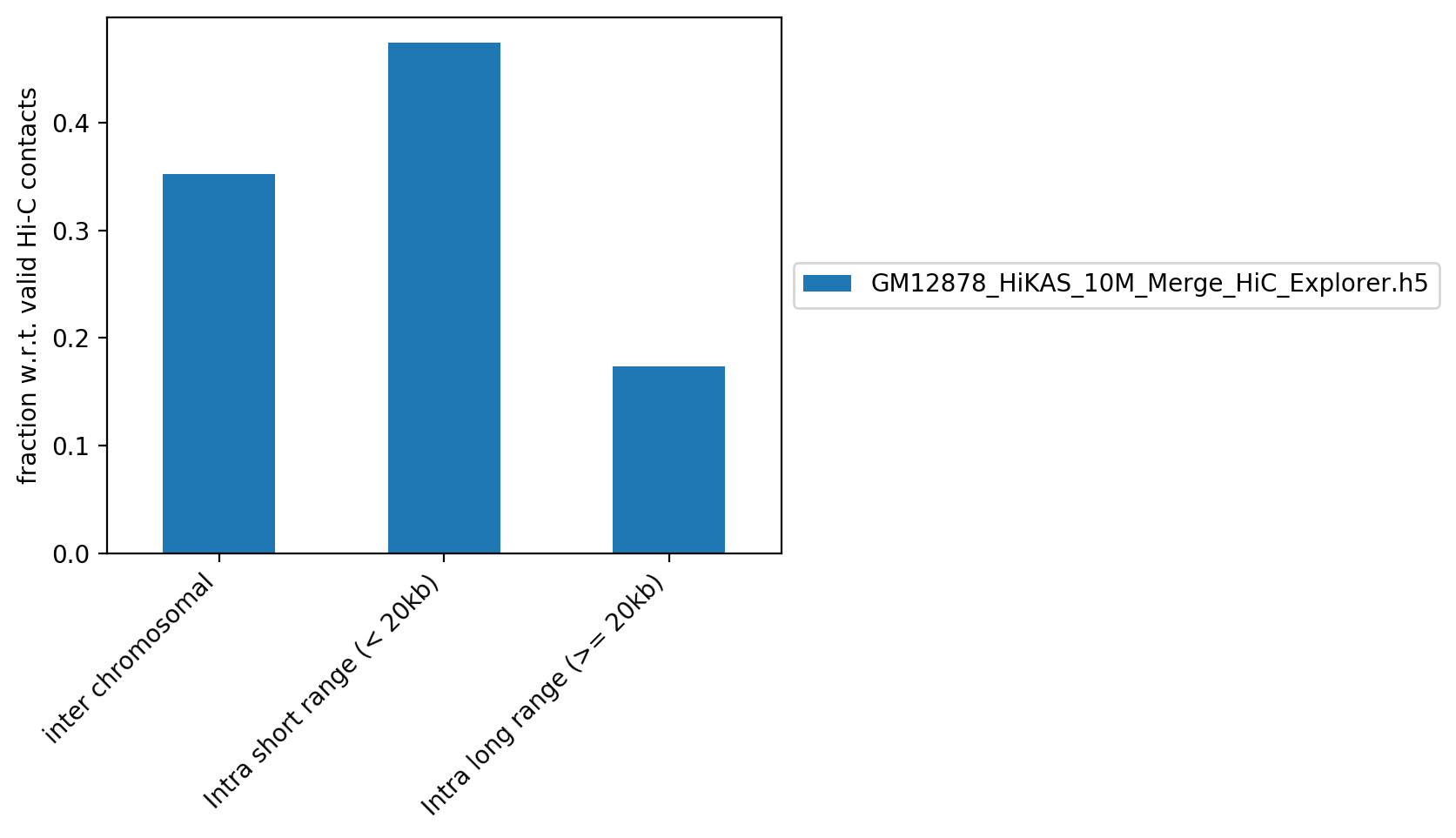
Contact distance table
| inter chromosomal | inter chromosomal % | Intra short range (< 20kb) | Intra short range (< 20kb) % | Intra long range (>= 20kb) | Intra long range (>= 20kb) % | |
|---|---|---|---|---|---|---|
| File | ||||||
| GM12878_HiKAS_10M_Merge_HiC_Explorer.h5 | 2,804,744 | 35.22% | 3,774,177 | 47.40% | 1,384,243 | 17.38% |
Read orientation
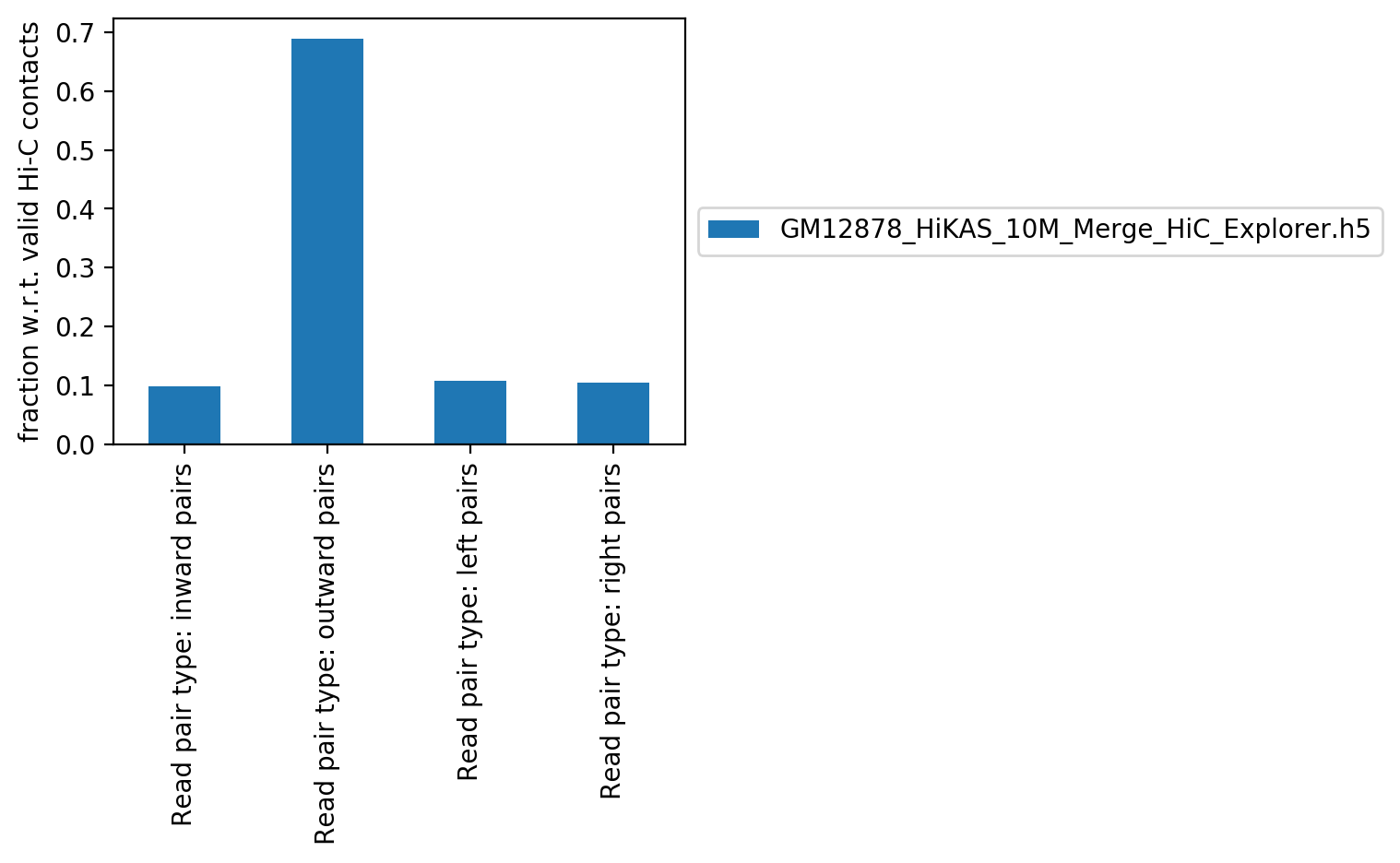
Read orientation table
| Read pair type: inward pairs | Read pair type: inward pairs % | Read pair type: outward pairs | Read pair type: outward pairs % | Read pair type: left pairs | Read pair type: left pairs % | Read pair type: right pairs | Read pair type: right pairs % | |
|---|---|---|---|---|---|---|---|---|
| File | ||||||||
| GM12878_HiKAS_10M_Merge_HiC_Explorer.h5 | 507,713 | 9.84% | 3,553,717 | 68.89% | 554,744 | 10.75% | 542,246 | 10.51% |
Number of reads table
| Sequenced reads | Pairs mappable, unique and high quality | Hi-C contacts | One mate unmapped | One mate not unique | Low mapping quality | dangling end GATC | self ligation (removed) | One mate not close to rest site | same fragment | self circle | duplicated pairs | inter chromosomal | Intra short range (< 20kb) | Intra long range (>= 20kb) | Read pair type: inward pairs | Read pair type: outward pairs | Read pair type: left pairs | Read pair type: right pairs | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File | |||||||||||||||||||
| GM12878_HiKAS_10M_Merge_HiC_Explorer.h5 | 513627084 | 391856521 | 7963164 | 21158380 | 59841028 | 40771155 | 78175996 | 75009486 | 0 | 170003494 | 2532589 | 60704381 | 2804744 | 3774177 | 1384243 | 507713 | 3553717 | 554744 | 542246 |