Purpose of the notebook is to generate reports via saved images¶
In [1]:
import os
import sys
sys.path.append("/oak/stanford/groups/akundaje/manyu/C2H2_ZNF_project/utils/modisco_utils")
sys.path.append(os.path.abspath("/oak/stanford/groups/akundaje/manyu/softwares/tfmodisco_tf_models/src/"))
sys.path.append(os.path.abspath("/oak/stanford/groups/akundaje/manyu/softwares/tfmodisco_tf_models/notebooks/reports/"))
from tfmodisco.run_tfmodisco import import_shap_scores, import_tfmodisco_results
from motif.read_motifs import pfm_info_content, trim_motif_by_ic
import motif.moods as moods
import plot.viz_sequence as viz_sequence
from util import figure_to_vdom_image, import_peak_table
import h5py
import pandas as pd
import numpy as np
import modisco
import sklearn.decomposition
import umap
import matplotlib.pyplot as plt
import matplotlib.font_manager as font_manager
import vdom.helpers as vdomh
from IPython.display import display
import tqdm
import io
import base64
import urllib
import deepdish as dd
import shutil
from vdom.helpers import h1, p, img, div, b
Main class to generate reports in HTML form using the saved images and h5 files¶
In [2]:
from shutil import copyfile
def get_read_handle(h5_file):
'''Copy to temp with a hashed name and give the read handle'''
name = h5_file.split('/')[-1]
#copy_file = '/users/manyu/temp/{}_{}'.format(str(hash(h5_file)),name)
copy_file = '/srv/www/kundaje/manyu/tmp/{}_{}'.format(str(hash(h5_file)),name)
if os.path.exists(copy_file):
os.remove(copy_file)
copyfile(h5_file,copy_file)
f = h5py.File(copy_file,'r')
return(f)
def close_h5file_read_handle(h5_file,read_handle):
'''Removes the copied h5 file in temp and the read handle to avoid bloating'''
read_handle.close()
name = h5_file.split('/')[-1]
#copy_file = '/users/manyu/temp/{}_{}'.format(str(hash(h5_file)),name)
copy_file = '/srv/www/kundaje/manyu/tmp/{}_{}'.format(str(hash(h5_file)),name)
if os.path.exists(copy_file):
os.remove(copy_file)
class ReportGenerator():
def __init__(self,tf,dataset):
self.dataset = dataset
self.tf = tf
# Path to saved modisco results on /oak. Images and h5 files
self.reports_base = '/oak/stanford/groups/akundaje/manyu/C2H2_ZNF_project/train_profile_models_2020/modisco_reports'
self.results_dir = '{}/{}/{}'.format(self.reports_base,dataset,tf)
## Paths to the saved modisco report results to copy/copied on Mitra: Images and h5 files
self.modisco_reports_results_mitra = '/srv/www/kundaje/manyu/C2H2_ZNF_project/modisco_report_saved_information'
self.mitra_results_dataset_dir = '{}/{}'.format(self.modisco_reports_results_mitra,self.dataset)
if not os.path.exists(self.mitra_results_dataset_dir):
os.makedirs(self.mitra_results_dataset_dir)
self.save_results_mitra_dir = '{}/{}'.format(self.mitra_results_dataset_dir,self.tf)
self.main_patterns_dir = '{}/main_patterns'.format(self.save_results_mitra_dir)
self.main_patterns = '{}/all_motifs.h5'.format(self.main_patterns_dir)
self.subpatterns_dir = '{}/sub_patterns'.format(self.save_results_mitra_dir)
self.subpatterns = '{}/all_motif_subclusters.h5'.format(self.subpatterns_dir)
## Paths to the modisco reports html on lab cluster
self.modisco_reports_base = '/srv/www/kundaje/manyu/C2H2_ZNF_project/modisco_reports'
self.modisco_reports_dataset_base = '{}/{}'.format(self.modisco_reports_base,self.dataset)
if not os.path.exists(self.modisco_reports_dataset_base):
os.makedirs(self.modisco_reports_dataset_base)
self.modisco_report_filename = '{}/{}.html'.format(self.modisco_reports_dataset_base,self.tf)
## Path to webpages and weblinks to images(needs weblinks to load; not local links)
self.mitra_web_base = 'http://mitra.stanford.edu/kundaje/manyu/C2H2_ZNF_project/'
self.mitra_web_results_base = '{}/modisco_report_saved_information/{}/{}'.format(self.mitra_web_base,self.dataset,self.tf)
self.mitra_web_results_main_patterns = '{}/main_patterns'.format(self.mitra_web_results_base)
self.mitra_web_results_sub_patterns = '{}/sub_patterns'.format(self.mitra_web_results_base)
self.mitra_web_results_profiles = '{}/profiles_dir'.format(self.mitra_web_results_base)
self.mitra_web_results_b1h_aln = '{}/b1h_aligments'.format(self.mitra_web_results_base)
##Do we need this here?
#self.mitra_web_reports_base = '{}/modisco_reports'.format(self.mitra_web_base)
def copy_to_mitra(self):
assert(os.path.exists(self.results_dir))
if os.path.exists(self.save_results_mitra_dir):
shutil.rmtree(self.save_results_mitra_dir)
shutil.copytree(self.results_dir,self.save_results_mitra_dir)
print('Finished copying files to Mitra')
permissions_cmd = 'chmod -R 777 {}'.format(self.save_results_mitra_dir)
os.system(permissions_cmd)
def show_subcluster_table(self):
assert(os.path.exists(self.main_patterns))
## Opening a read handle for h5. Remmeber to close it also
try:
patterns_read_handle = get_read_handle(self.main_patterns)
except Exception as e:
print(e)
try:
sub_patterns_read_handle = get_read_handle(self.subpatterns)
except Exception as e:
print(e)
num_patterns = len(patterns_read_handle.keys())
for pattern_i in range(num_patterns):
display(vdomh.h3("Pattern {}/{}".format(str(pattern_i + 1), num_patterns)))
# Sort this out:
colgroup = vdomh.colgroup(
vdomh.col(style={"width": "5%"}),
vdomh.col(style={"width": "5%"}),
vdomh.col(style={"width": "50%"}),
vdomh.col(style={"width": "40%"})
)
header = vdomh.thead(
vdomh.tr(
vdomh.th("Subpattern", style={"text-align": "center"}),
vdomh.th("Seqlets", style={"text-align": "center"}),
vdomh.th("Embeddings", style={"text-align": "center"}),
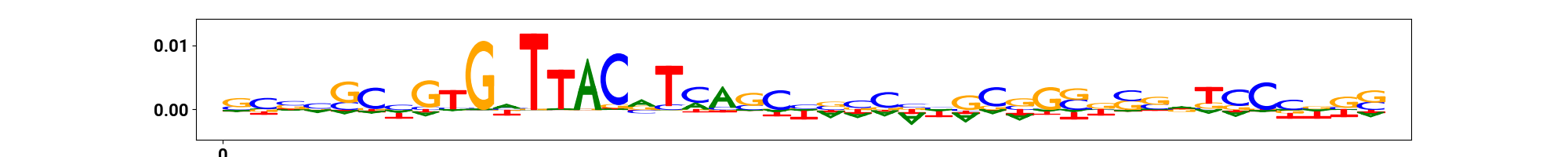
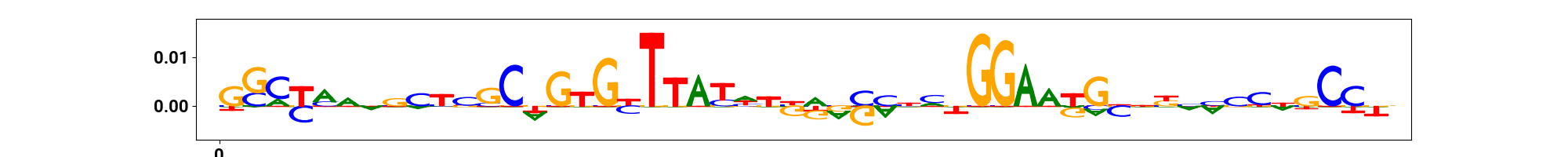
vdomh.th("CWM", style={"text-align": "center"}),
vdomh.th("PFM", style={"text-align": "center"})
)
)
## Fill in the paths and seqlets of the main pattern
total_seqlets = patterns_read_handle[str(pattern_i)]['n_seqlets'].value
emb_agg_path = '{}/{}_subcluster_agg.png'.format(self.mitra_web_results_sub_patterns,str(pattern_i))
pwm_path = '{}/{}_pfm_full.png'.format(self.mitra_web_results_main_patterns,str(pattern_i))
cwm_path = '{}/{}_cwm_full.png'.format(self.mitra_web_results_main_patterns,str(pattern_i))
table_rows = [vdomh.tr(
vdomh.td("Agg."),
vdomh.td(str(total_seqlets)),
vdomh.td(img(src=emb_agg_path)),
vdomh.td(img(src=cwm_path)),
vdomh.td(img(src=pwm_path))
)]
total_sub_pats = len(sub_patterns_read_handle[str(pattern_i)].keys())
for subpat_i in range(total_sub_pats):
emb_path = '{}/{}_subcluster_{}.png'.format(self.mitra_web_results_sub_patterns,str(pattern_i),str(subpat_i))
total_seqlets = sub_patterns_read_handle[str(pattern_i)]['subcluster_{}'.format(str(subpat_i))]['n_seqlets'].value
pwm_path = '{}/{}_subcluster_{}_pfm_trimmed.png'.format(self.mitra_web_results_sub_patterns,str(pattern_i),str(subpat_i))
cwm_path = '{}/{}_subcluster_{}_cwm_trimmed.png'.format(self.mitra_web_results_sub_patterns,str(pattern_i),str(subpat_i))
table_rows.append(vdomh.tr(
vdomh.td('{}'.format(str(subpat_i))),
vdomh.td(str(total_seqlets)),
vdomh.td(img(src=emb_path)),
vdomh.td(img(src=cwm_path)),
vdomh.td(img(src=pwm_path))
))
table = vdomh.table(header,vdomh.tbody(*table_rows))
display(table)
#Remove unnecessary temp files
close_h5file_read_handle(self.main_patterns,patterns_read_handle)
close_h5file_read_handle(self.subpatterns,sub_patterns_read_handle)
def show_footprints_and_peak_dist(self):
try:
patterns_read_handle = get_read_handle(self.main_patterns)
except Exception as e:
print(e)
num_patterns = len(patterns_read_handle.keys())
for pattern_i in range(num_patterns):
display(vdomh.h3("Pattern {}/{} Footprint".format(str(pattern_i + 1), num_patterns)))
footprint_plot = '{}/profiles_pattern_{}.png'.format(self.mitra_web_results_profiles,str(pattern_i))
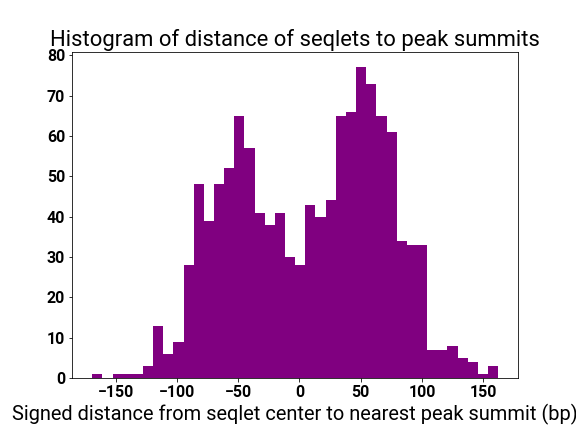
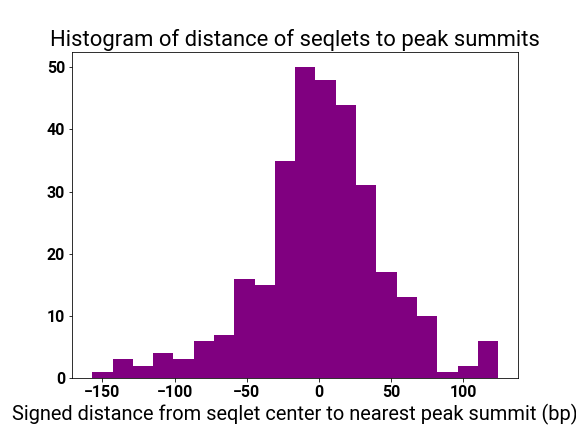
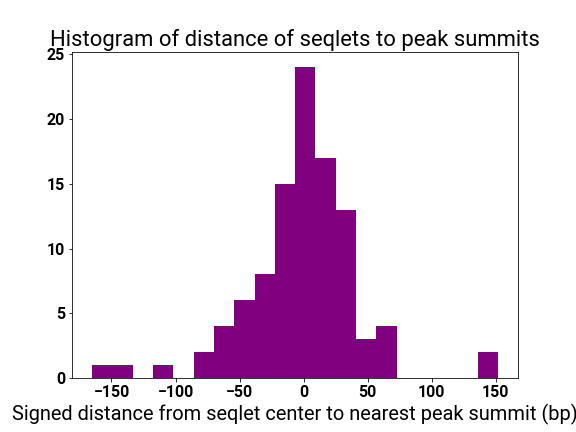
motif_distplot = '{}/motif_distances_pattern_{}.png'.format(self.mitra_web_results_profiles,str(pattern_i))
display(img(src=footprint_plot))
display(vdomh.h3("Pattern {}/{} Motif Distance Distribution".format(str(pattern_i + 1), num_patterns)))
display(img(src=motif_distplot))
def show_b1h_alignments(self):
try:
patterns_read_handle = get_read_handle(self.main_patterns)
except Exception as e:
print(e)
num_patterns = len(patterns_read_handle.keys())
header = vdomh.thead(
vdomh.tr(
vdomh.th("Pattern", style={"text-align": "center"}),
vdomh.th("N-Seqlets", style={"text-align": "center"}),
vdomh.th("Alignment", style={"text-align": "center"})
)
)
display(vdomh.h2("B1H alignments of all patterns"))
table_rows = []
for pattern_i in range(num_patterns):
total_seqlets = patterns_read_handle[str(pattern_i)]['n_seqlets'].value
alignment_img = '{}/pattern_{}_aligned.png'.format(self.mitra_web_results_b1h_aln,str(pattern_i))
table_rows.append(vdomh.tr(
vdomh.td('{}'.format(str(pattern_i+1))),
vdomh.td('{}'.format(str(total_seqlets))),
vdomh.td(img(src=alignment_img)),
))
close_h5file_read_handle(self.main_patterns,patterns_read_handle)
table = vdomh.table(header,vdomh.tbody(*table_rows))
display(table)
In [3]:
# Define parameters/fetch arguments
dataset = os.environ["DATASET"]
tf = os.environ["TF"]
In [4]:
# tf = 'ENCSR464KFG'
# dataset = 'ENCODE'
In [5]:
a = ReportGenerator(tf=tf,dataset=dataset)
In [6]:
# a.copy_to_mitra()
In [7]:
a.show_subcluster_table()
In [8]:
a.show_footprints_and_peak_dist()
In [9]:
a.show_b1h_alignments()
In [10]:
In [10]: