| p-value: | 1e-3 |
| log p-value: | -7.273e+00 |
| Information Content per bp: | 1.751 |
| Number of Target Sequences with motif | 18.0 |
| Percentage of Target Sequences with motif | 4.03% |
| Number of Background Sequences with motif | 3.3 |
| Percentage of Background Sequences with motif | 0.73% |
| Average Position of motif in Targets | 10.9 +/- 3.0bp |
| Average Position of motif in Background | 12.0 +/- 6.3bp |
| Strand Bias (log2 ratio + to - strand density) | -0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.11 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
GAT3(MacIsaac)/Yeast
| Match Rank: | 1 |
| Score: | 0.73
| | Offset: | 0
| | Orientation: | reverse strand |
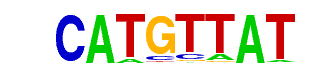
| Alignment: | CATGGCAT
CATGTTAT |
|

|
|
RIM101/Literature(Harbison)/Yeast
| Match Rank: | 2 |
| Score: | 0.71
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CATGGCAT
CTTGGCA- |
|


|
|
RIM101(MacIsaac)/Yeast
| Match Rank: | 3 |
| Score: | 0.71
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CATGGCAT
CTTGGCA- |
|


|
|
NAC92/MA1044.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.71
| | Offset: | -3
| | Orientation: | forward strand |
| Alignment: | ---CATGGCAT-
TAACACGCCACA |
|


|
|
MAFG::NFE2L1/MA0089.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.71
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CATGGCAT
CATGAC-- |
|


|
|
Tgif2(Homeobox)/mES-Tgif2-ChIP-Seq(GSE55404)/Homer
| Match Rank: | 6 |
| Score: | 0.70
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -CATGGCAT
ARNTGACA- |
|


|
|
pros/dmmpmm(Bergman)/fly
| Match Rank: | 7 |
| Score: | 0.69
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CATGGCAT
CATGNCT- |
|


|
|
pho/dmmpmm(Bergman)/fly
| Match Rank: | 8 |
| Score: | 0.67
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CATGGCAT
AATGGC-- |
|


|
|
INO2/MA0321.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.67
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CATGGCAT
GCATGTGAA |
|


|
|
MEIS3/MA0775.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.67
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CATGGCAT
CCTGTCAA |
|


|
|





















