| p-value: | 1e-3 |
| log p-value: | -7.835e+00 |
| Information Content per bp: | 1.516 |
| Number of Target Sequences with motif | 46.0 |
| Percentage of Target Sequences with motif | 9.73% |
| Number of Background Sequences with motif | 19.9 |
| Percentage of Background Sequences with motif | 4.23% |
| Average Position of motif in Targets | 10.2 +/- 3.3bp |
| Average Position of motif in Background | 10.4 +/- 3.9bp |
| Strand Bias (log2 ratio + to - strand density) | -0.4 |
| Multiplicity (# of sites on avg that occur together) | 1.04 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
SeqBias: polyC-repeat
| Match Rank: | 1 |
| Score: | 0.75
| | Offset: | 0
| | Orientation: | forward strand |
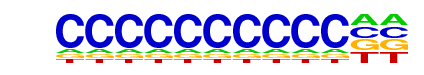
| Alignment: | CCCMCYSYCCBC
CCCCCCCCCC-- |
|

|
|
PB0097.1_Zfp281_1/Jaspar
| Match Rank: | 2 |
| Score: | 0.73
| | Offset: | -3
| | Orientation: | forward strand |
| Alignment: | ---CCCMCYSYCCBC
TCCCCCCCCCCCCCC |
|


|
|
Maz(Zf)/HepG2-Maz-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 3 |
| Score: | 0.72
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | CCCMCYSYCCBC
-CCCCCCCC--- |
|


|
|
SP2/MA0516.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.69
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CCCMCYSYCCBC--
GCCCCGCCCCCTCCC |
|


|
|
KLF5/MA0599.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.68
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CCCMCYSYCCBC
GCCCCGCCCC-- |
|


|
|
SP1/MA0079.3/Jaspar
| Match Rank: | 6 |
| Score: | 0.67
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CCCMCYSYCCBC
GCCCCGCCCCC-- |
|


|
|
EGR1/MA0162.2/Jaspar
| Match Rank: | 7 |
| Score: | 0.65
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CCCMCYSYCCBC-
CCCCCGCCCCCGCC |
|


|
|
SUT1/SUT1_YPD/[](Harbison)/Yeast
| Match Rank: | 8 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | CCCMCYSYCCBC
--CCGGCCCCGC |
|


|
|
Sp1(Zf)/Promoter/Homer
| Match Rank: | 9 |
| Score: | 0.63
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --CCCMCYSYCCBC
GGCCCCGCCCCC-- |
|


|
|
ABI4(1)(AP2/EREBP)/Zea mays/AthaMap
| Match Rank: | 10 |
| Score: | 0.63
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | CCCMCYSYCCBC
--CACCGCCCCC |
|


|
|





















