| p-value: | 1e-2 |
| log p-value: | -5.038e+00 |
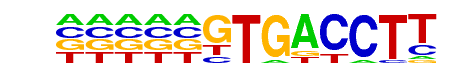
| Information Content per bp: | 1.330 |
| Number of Target Sequences with motif | 50.0 |
| Percentage of Target Sequences with motif | 10.57% |
| Number of Background Sequences with motif | 28.0 |
| Percentage of Background Sequences with motif | 5.93% |
| Average Position of motif in Targets | 11.9 +/- 3.6bp |
| Average Position of motif in Background | 11.2 +/- 3.3bp |
| Strand Bias (log2 ratio + to - strand density) | 0.3 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
bZIP911/MA0097.1/Jaspar
| Match Rank: | 1 |
| Score: | 0.57
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -NNNHNSDVDBYD
GATGACGTGGCC- |
|


|
|
MTF1/MA0863.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.56
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD--
GTGCCGTGTGCAAA |
|


|
|
NR4A2/MA0160.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.55
| | Offset: | 5
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD-
-----GTGACCTT |
|

|
|
bZIP911(1)(bZIP)/Antirrhinum majus/AthaMap
| Match Rank: | 4 |
| Score: | 0.55
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -NNNHNSDVDBYD
GGTGACGTGGCC- |
|


|
|
RTG3/Literature(Harbison)/Yeast
| Match Rank: | 5 |
| Score: | 0.52
| | Offset: | 5
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD
-----GTGACC- |
|


|
|
HAC1(MacIsaac)/Yeast
| Match Rank: | 6 |
| Score: | 0.52
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD
--TACGTGGCCT |
|


|
|
MGA1/MA0336.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.52
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -NNNHNSDVDBYD--------
NTTNNAGTGTTCTATANNNNN |
|


|
|
lin-14/MA0261.1/Jaspar
| Match Rank: | 8 |
| Score: | 0.51
| | Offset: | 5
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD
-----GTGTTC- |
|


|
|
MA0261.1_lin-14/Jaspar
| Match Rank: | 9 |
| Score: | 0.51
| | Offset: | 5
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD
-----GTGTTC- |
|


|
|
Gfi1b/MA0483.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.51
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | NNNHNSDVDBYD
-TGCTGTGATTT |
|


|
|





















