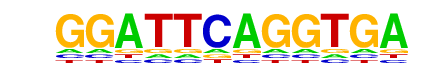
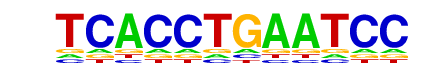
| p-value: | 1e-1 |
| log p-value: | -2.410e+00 |
| Information Content per bp: | 1.530 |
| Number of Target Sequences with motif | 7.0 |
| Percentage of Target Sequences with motif | 0.02% |
| Number of Background Sequences with motif | 3.0 |
| Percentage of Background Sequences with motif | 0.01% |
| Average Position of motif in Targets | 624.4 +/- 362.6bp |
| Average Position of motif in Background | 847.8 +/- 178.4bp |
| Strand Bias (log2 ratio + to - strand density) | 10.0 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
RAV1(2)(AP2/EREBP)/Arabidopsis thaliana/AthaMap
| Match Rank: | 1 |
| Score: | 0.75
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | GGATTCAGGTGA-
-GNNTCAGGTGAN |
|


|
|
MA0583.1_RAV1_(var.2)/Jaspar
| Match Rank: | 2 |
| Score: | 0.75
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | GGATTCAGGTGA-
-GNNTCAGGTGAN |
|


|
|
ARR1(MacIsaac)/Yeast
| Match Rank: | 3 |
| Score: | 0.72
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | GGATTCAGGTGA
--ATTCAAGT-- |
|
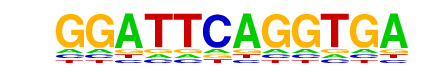
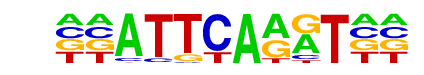
|
|
MA0086.1_sna/Jaspar
| Match Rank: | 4 |
| Score: | 0.71
| | Offset: | 5
| | Orientation: | forward strand |
| Alignment: | GGATTCAGGTGA
-----CAGGTG- |
|

|
|
MA0274.1_ARR1/Jaspar
| Match Rank: | 5 |
| Score: | 0.70
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | GGATTCAGGTGA
--ATTCAANT-- |
|

|
|
MA0103.2_ZEB1/Jaspar
| Match Rank: | 6 |
| Score: | 0.69
| | Offset: | 5
| | Orientation: | reverse strand |
| Alignment: | GGATTCAGGTGA--
-----CAGGTGAGG |
|


|
|
E2A(bHLH),near_PU.1/Bcell-PU.1-ChIP-Seq(GSE21512)/Homer
| Match Rank: | 7 |
| Score: | 0.68
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | GGATTCAGGTGA-
---NNCAGGTGNN |
|


|
|
TYE7/TYE7_YPD/39-CBF1(Harbison)/Yeast
| Match Rank: | 8 |
| Score: | 0.67
| | Offset: | 4
| | Orientation: | forward strand |
| Alignment: | GGATTCAGGTGA-
----TCACGTGAT |
|


|
|
MA0580.1_DYT1/Jaspar
| Match Rank: | 9 |
| Score: | 0.66
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --GGATTCAGGTGA
CGTGAGTCACGTGA |
|
|
|
Cbf1(bHLH)/Yeast-Cbf1-ChIP-Seq(GSE29506)/Homer
| Match Rank: | 10 |
| Score: | 0.65
| | Offset: | 4
| | Orientation: | forward strand |
| Alignment: | GGATTCAGGTGA--
----TCACGTGAYH |
|


|
|