| p-value: | 1e0 |
| log p-value: | -1.935e+00 |
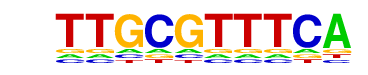
| Information Content per bp: | 1.530 |
| Number of Target Sequences with motif | 6.0 |
| Percentage of Target Sequences with motif | 0.20% |
| Number of Background Sequences with motif | 3.0 |
| Percentage of Background Sequences with motif | 0.10% |
| Average Position of motif in Targets | 432.2 +/- 272.1bp |
| Average Position of motif in Background | 332.6 +/- 251.3bp |
| Strand Bias (log2 ratio + to - strand density) | 0.0 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
MA0393.1_STE12/Jaspar
| Match Rank: | 1 |
| Score: | 0.77
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA
TGAAACA--- |
|


|
|
STE12(MacIsaac)/Yeast
| Match Rank: | 2 |
| Score: | 0.73
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA
TGAAACA--- |
|


|
|
STE12/STE12_Alpha/92-STE12(Harbison)/Yeast
| Match Rank: | 3 |
| Score: | 0.73
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA
TGAAACA--- |
|


|
|
DIG1/DIG1_YPD/32-STE12(Harbison)/Yeast
| Match Rank: | 4 |
| Score: | 0.72
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA
TGAAACA--- |
|


|
|
DIG1(MacIsaac)/Yeast
| Match Rank: | 5 |
| Score: | 0.68
| | Offset: | -9
| | Orientation: | forward strand |
| Alignment: | ---------TGAAACGCAA
ANACANTTNTGAAACA--- |
|


|
|
T1ISRE(IRF)/ThioMac-Ifnb-Expression/Homer
| Match Rank: | 6 |
| Score: | 0.66
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TGAAACGCAA--
AGAAACGAAAGT |
|


|
|
MA0242.1_run::Bgb/Jaspar
| Match Rank: | 7 |
| Score: | 0.66
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA
-TAACCGCAA |
|


|
|
CST6(MacIsaac)/Yeast
| Match Rank: | 8 |
| Score: | 0.65
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | TGAAACGCAA
-NAAATGCA- |
|


|
|
MA0204.1_Six4/Jaspar
| Match Rank: | 9 |
| Score: | 0.65
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA
TGATAC---- |
|


|
|
STB1/STB1_YPD/1-SWI4,1-SWI6(Harbison)/Yeast
| Match Rank: | 10 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | TGAAACGCAA--
--AAACGCGAAA |
|


|
|