



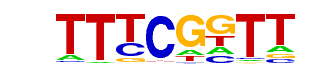






| p-value: | 1e-50 |
| log p-value: | -1.159e+02 |
| Information Content per bp: | 1.655 |
| Number of Target Sequences with motif | 1454.0 |
| Percentage of Target Sequences with motif | 3.21% |
| Number of Background Sequences with motif | 762.4 |
| Percentage of Background Sequences with motif | 1.69% |
| Average Position of motif in Targets | 13.0 +/- 4.7bp |
| Average Position of motif in Background | 13.1 +/- 6.0bp |
| Strand Bias (log2 ratio + to - strand density) | -0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix) reverse opposite |
| Rank | Match Score | Redundant Motif | P-value | log P-value | % of Targets | % of Background | Motif file |
| 1 | 0.893 |  | 1e-39 | -90.272203 | 1.91% | 0.89% | motif file (matrix) |
| 2 | 0.608 |  | 1e-34 | -79.837815 | 2.20% | 1.15% | motif file (matrix) |
| 3 | 0.623 |  | 1e-34 | -78.494961 | 2.39% | 1.30% | motif file (matrix) |
| 4 | 0.616 |  | 1e-33 | -78.002400 | 2.12% | 1.11% | motif file (matrix) |
| 5 | 0.730 |  | 1e-32 | -75.468659 | 4.46% | 2.96% | motif file (matrix) |
| 6 | 0.905 |  | 1e-29 | -68.265533 | 2.10% | 1.15% | motif file (matrix) |
| 7 | 0.839 |  | 1e-20 | -48.031362 | 1.02% | 0.48% | motif file (matrix) |
| 8 | 0.758 |  | 1e-19 | -45.832189 | 8.45% | 6.82% | motif file (matrix) |
| 9 | 0.788 |  | 1e-15 | -34.858840 | 2.67% | 1.88% | motif file (matrix) |