| p-value: | 1e-8 |
| log p-value: | -1.974e+01 |
| Information Content per bp: | 1.811 |
| Number of Target Sequences with motif | 65.0 |
| Percentage of Target Sequences with motif | 0.14% |
| Number of Background Sequences with motif | 14.6 |
| Percentage of Background Sequences with motif | 0.03% |
| Average Position of motif in Targets | 13.6 +/- 3.6bp |
| Average Position of motif in Background | 13.5 +/- 3.9bp |
| Strand Bias (log2 ratio + to - strand density) | 1.1 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
SeqBias: polyA-repeat
| Match Rank: | 1 |
| Score: | 0.94
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | AAAAAAAAAAAAAA
--AAAAAAAAAA-- |
|


|
|
PB0182.1_Srf_2/Jaspar
| Match Rank: | 2 |
| Score: | 0.87
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --AAAAAAAAAAAAAA-
GTTAAAAAAAAAAATTA |
|


|
|
RLR1?/SacCer-Promoters/Homer
| Match Rank: | 3 |
| Score: | 0.81
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | AAAAAAAAAAAAAA
--AAAARRGAAAAW |
|


|
|
hb/dmmpmm(Noyes)/fly
| Match Rank: | 4 |
| Score: | 0.79
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | AAAAAAAAAAAAAA
ACNAAAAAAACA-- |
|


|
|
PB0116.1_Elf3_2/Jaspar
| Match Rank: | 5 |
| Score: | 0.78
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --AAAAAAAAAAAAAA-
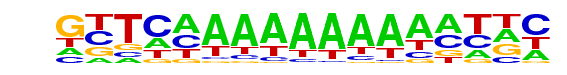
GTTCAAAAAAAAAATTC |
|

|
|
PB0093.1_Zfp105_1/Jaspar
| Match Rank: | 6 |
| Score: | 0.77
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -AAAAAAAAAAAAAA
AACAAACAACAAGAG |
|


|
|
SOC1/MA0554.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.75
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -AAAAAAAAAAAAAA------
AAAAAAAAAAAAAAAAAAAAA |
|


|
|
PB0192.1_Tcfap2e_2/Jaspar
| Match Rank: | 8 |
| Score: | 0.75
| | Offset: | -3
| | Orientation: | forward strand |
| Alignment: | ---AAAAAAAAAAAAAA
TACTGGAAAAAAAA--- |
|


|
|
hb/dmmpmm(Down)/fly
| Match Rank: | 9 |
| Score: | 0.75
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | AAAAAAAAAAAAAA
-CATAAAAAA---- |
|


|
|
hb/MA0049.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.73
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | AAAAAAAAAAAAAA
-GCATAAAAAA--- |
|


|
|





















