Dup QC (all)
| Unpaired Reads | Paired Reads | Unmapped Reads | Unpaired Dupes | Paired Dupes | Paired Opt. Dupes | % Dupes | |
|---|---|---|---|---|---|---|---|
| rep1 | 16912203 | 0 | 0 | 13766775 | 0 | 0 | 0.814014 |
Flagstat QC (all, filtered)
| Reads (QC-passed) | Reads (QC-failed) | Dupes (QC-passed) | Dupes (QC-failed) | Mapped Reads | % Mapped | |
|---|---|---|---|---|---|---|
| rep1 | 3145428 | 0 | 0 | 0 | 3145428 | 100.00 |
PBC QC (all)
| Total Read Pairs | Distinct Read Pairs | One Read Pair | Two Read Pairs | NRF = Distinct/Total | PBC1 = OnePair/Distinct | PBC2 = OnePair/TwoPair | |
|---|---|---|---|---|---|---|---|
| rep1 | 16908105 | 3648884 | 1225028 | 402845 | 0.215807 | 0.335727 | 3.040941 |
NRF (non redundant fraction)
PBC1 (PCR Bottleneck coefficient 1)
PBC2 (PCR Bottleneck coefficient 2)
PBC1 is the primary measure. Provisionally
- 0-0.5 is severe bottlenecking
- 0.5-0.8 is moderate bottlenecking
- 0.8-0.9 is mild bottlenecking
- 0.9-1.0 is no bottlenecking
Cross-correlation QC (all)
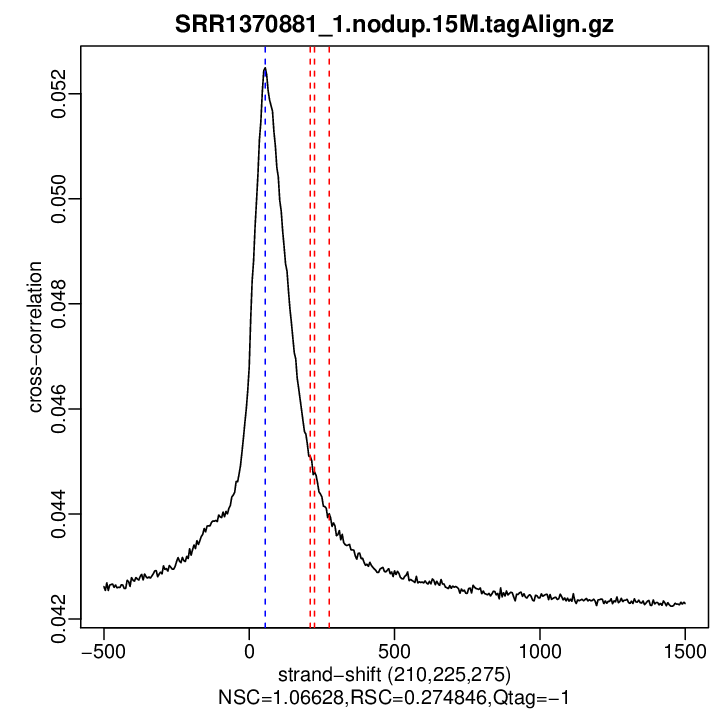
rep1
| numReads | estFragLen | corr_estFragLen | PhantomPeak | corr_phantomPeak | argmin_corr | min_corr | NSC | RSC | |
|---|---|---|---|---|---|---|---|---|---|
| rep1 | 3144688 | 210 | 0.0450985596418006 | 55 | 0.05249537 | 1500 | 0.04229503 | 1.066285 | 0.2748465 |
Normalized strand cross-correlation coefficient (NSC) = col9 in outFile
Relative strand cross-correlation coefficient (RSC) = col10 in outFile
Estimated fragment length = col3 in outFile, take the top value
Important columns highlighted, but all/whole file can be stored for display

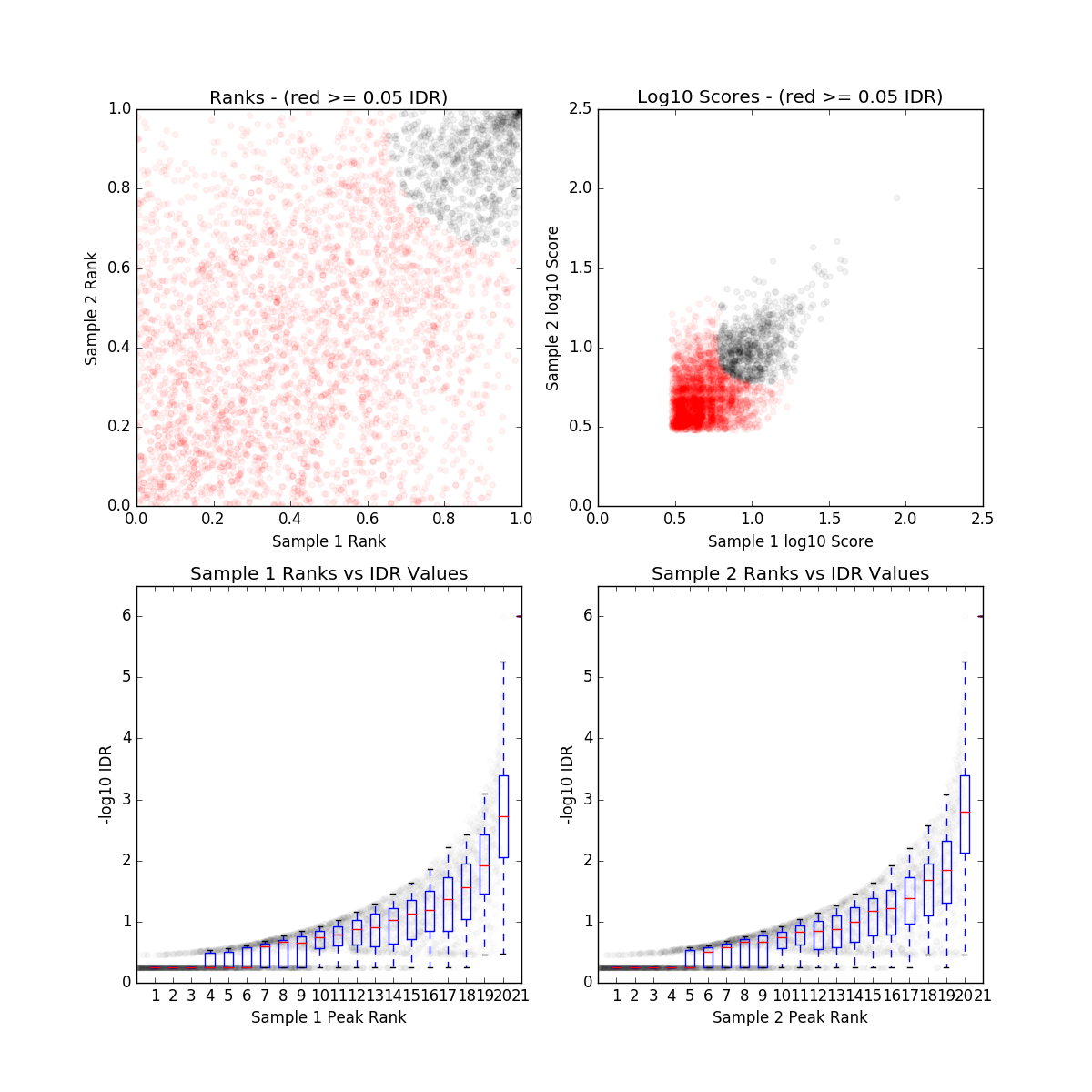
IDR QC (idr)
ZNF382_IDR_final.qc
| Nt | N1 | Np | conservative_set | optimal_set | rescue_ratio | self_consistency_ratio | reproducibility |
|---|---|---|---|---|---|---|---|
| 0 | 780 | 0 | N/A | N/A | NaN | 1.0 | 1 |