| p-value: | 1e-5 |
| log p-value: | -1.348e+01 |
| Information Content per bp: | 1.605 |
| Number of Target Sequences with motif | 970.0 |
| Percentage of Target Sequences with motif | 1.98% |
| Number of Background Sequences with motif | 1275.8 |
| Percentage of Background Sequences with motif | 1.62% |
| Average Position of motif in Targets | 204.5 +/- 131.6bp |
| Average Position of motif in Background | 239.5 +/- 175.3bp |
| Strand Bias (log2 ratio + to - strand density) | 0.0 |
| Multiplicity (# of sites on avg that occur together) | 1.01 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
TEAD(TEA)/Fibroblast-PU.1-ChIP-Seq(Unpublished)/Homer
| Match Rank: | 1 |
| Score: | 0.78
| | Offset: | -1
| | Orientation: | forward strand |
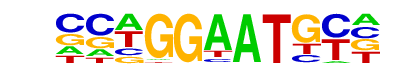
| Alignment: | -AAGGAWTSCT
NCTGGAATGC- |
|


|
|
TEAD4(TEA)/Tropoblast-Tead4-ChIP-Seq(GSE37350)/Homer
| Match Rank: | 2 |
| Score: | 0.74
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -AAGGAWTSCT
CCWGGAATGY- |
|

|
|
ZmHOX2a(1)(HD-HOX)/Zea mays/AthaMap
| Match Rank: | 3 |
| Score: | 0.73
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -AAGGAWTSCT
TTAGGAC---- |
|


|
|
TEC1/TEC1_YPD/[](Harbison)/Yeast
| Match Rank: | 4 |
| Score: | 0.72
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | AAGGAWTSCT
-AGGAATG-- |
|


|
|
TEAD3/MA0808.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.71
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | AAGGAWTSCT
-TGGAATGT- |
|


|
|
TEAD2(TEA)/Py2T-Tead2-ChIP-Seq(GSE55709)/Homer
| Match Rank: | 6 |
| Score: | 0.70
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -AAGGAWTSCT
CCWGGAATGY- |
|


|
|
TEAD4/MA0809.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.70
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | AAGGAWTSCT
NTGGAATGTN |
|


|
|
TEC1/MA0406.1/Jaspar
| Match Rank: | 8 |
| Score: | 0.69
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | AAGGAWTSCT
-GGGAATGT- |
|


|
|
TEAD1/MA0090.2/Jaspar
| Match Rank: | 9 |
| Score: | 0.68
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | AAGGAWTSCT
NTGGAATGTG |
|


|
|
Eip74EF/dmmpmm(SeSiMCMC)/fly
| Match Rank: | 10 |
| Score: | 0.67
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | AAGGAWTSCT
-AGGAAGTAT |
|


|
|





















