| p-value: | 1e-11 |
| log p-value: | -2.732e+01 |
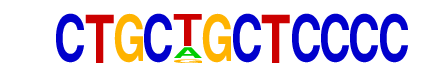
| Information Content per bp: | 1.954 |
| Number of Target Sequences with motif | 93.0 |
| Percentage of Target Sequences with motif | 0.19% |
| Number of Background Sequences with motif | 43.1 |
| Percentage of Background Sequences with motif | 0.05% |
| Average Position of motif in Targets | 214.2 +/- 135.6bp |
| Average Position of motif in Background | 214.4 +/- 125.1bp |
| Strand Bias (log2 ratio + to - strand density) | -0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.26 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
Znf263(Zf)/K562-Znf263-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 1 |
| Score: | 0.66
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | GGGGAGCAGCAG
-GGGAGGACNG- |
|


|
|
POL013.1_MED-1/Jaspar
| Match Rank: | 2 |
| Score: | 0.66
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | GGGGAGCAGCAG
-CGGAGC----- |
|


|
|
MZF1/MA0056.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.66
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -GGGGAGCAGCAG
TGGGGA------- |
|


|
|
SeqBias: GCW-triplet
| Match Rank: | 4 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | GGGGAGCAGCAG--
--GCAGCAGCAGCA |
|


|
|
ZNF189(Zf)/HEK293-ZNF189.GFP-ChIP-Seq(GSE58341)/Homer
| Match Rank: | 5 |
| Score: | 0.63
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | GGGGAGCAGCAG
-TGGAACAGMA- |
|


|
|
Spz1/MA0111.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.62
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -GGGGAGCAGCAG
AGGGTAACAGC-- |
|


|
|
GFY(?)/Promoter/Homer
| Match Rank: | 7 |
| Score: | 0.60
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | GGGGAGCAGCAG-
-GGGAATTGTAGT |
|


|
|
SCL(bHLH)/HPC7-Scl-ChIP-Seq(GSE13511)/Homer
| Match Rank: | 8 |
| Score: | 0.60
| | Offset: | 4
| | Orientation: | forward strand |
| Alignment: | GGGGAGCAGCAG
----ANCAGCTG |
|


|
|
MyoD(bHLH)/Myotube-MyoD-ChIP-Seq(GSE21614)/Homer
| Match Rank: | 9 |
| Score: | 0.59
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | GGGGAGCAGCAG--
--NNAGCAGCTGCT |
|


|
|
PRDM9(Zf)/Testis-DMC1-ChIP-Seq(GSE35498)/Homer
| Match Rank: | 10 |
| Score: | 0.59
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -GGGGAGCAGCAG--
ADGGYAGYAGCATCT |
|


|
|