| p-value: | 1e-8 |
| log p-value: | -1.905e+01 |
| Information Content per bp: | 1.783 |
| Number of Target Sequences with motif | 34.0 |
| Percentage of Target Sequences with motif | 0.08% |
| Number of Background Sequences with motif | 8.7 |
| Percentage of Background Sequences with motif | 0.01% |
| Average Position of motif in Targets | 246.6 +/- 214.8bp |
| Average Position of motif in Background | 185.6 +/- 165.2bp |
| Strand Bias (log2 ratio + to - strand density) | 2.9 |
| Multiplicity (# of sites on avg that occur together) | 1.00 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
GFY(?)/Promoter/Homer
| Match Rank: | 1 |
| Score: | 0.68
| | Offset: | -1
| | Orientation: | reverse strand |
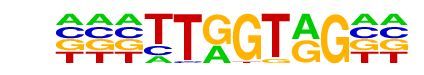
| Alignment: | -GGAGTAGTAGTR
GGGAATTGTAGT- |
|


|
|
SMP1/Literature(Harbison)/Yeast
| Match Rank: | 2 |
| Score: | 0.66
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---GGAGTAGTAGTR
CTAAAAATAGTAGT- |
|


|
|
SMP1(MacIsaac)/Yeast
| Match Rank: | 3 |
| Score: | 0.66
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---GGAGTAGTAGTR
CTAAAAATAGTAGT- |
|


|
|
schlank/MA0193.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.64
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | GGAGTAGTAGTR
---TTGGTAG-- |
|

|
|
br(var.2)/MA0011.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.63
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | GGAGTAGTAGTR
-AAATAGTA--- |
|


|
|
Aef1/dmmpmm(Pollard)/fly
| Match Rank: | 6 |
| Score: | 0.61
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | GGAGTAGTAGTR
--TGTTGTTG-- |
|


|
|
odd/MA0454.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.58
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -GGAGTAGTAGTR
AACAGTAGCAG-- |
|


|
|
br-Z2/dmmpmm(SeSiMCMC)/fly
| Match Rank: | 8 |
| Score: | 0.58
| | Offset: | 4
| | Orientation: | forward strand |
| Alignment: | GGAGTAGTAGTR
----TAGTAA-- |
|


|
|
ARR1/Literature(Harbison)/Yeast
| Match Rank: | 9 |
| Score: | 0.57
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | GGAGTAGTAGTR
---TTAGTAA-- |
|


|
|
YAP3/Literature(Harbison)/Yeast
| Match Rank: | 10 |
| Score: | 0.57
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | GGAGTAGTAGTR
---TTAGTAA-- |
|


|
|





















