| p-value: | 1e-17 |
| log p-value: | -4.025e+01 |
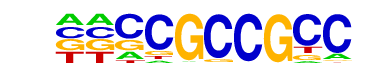
| Information Content per bp: | 1.768 |
| Number of Target Sequences with motif | 2390.0 |
| Percentage of Target Sequences with motif | 5.70% |
| Number of Background Sequences with motif | 3288.3 |
| Percentage of Background Sequences with motif | 4.54% |
| Average Position of motif in Targets | 238.3 +/- 150.4bp |
| Average Position of motif in Background | 227.5 +/- 109.9bp |
| Strand Bias (log2 ratio + to - strand density) | 0.2 |
| Multiplicity (# of sites on avg that occur together) | 1.47 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
SeqBias: CG bias
| Match Rank: | 1 |
| Score: | 0.75
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCSGSCGSC
CCCCCCCCCC |
|


|
|
Os05g0497200/MA1034.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.75
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCSGSCGSC
CGCCGCCA-- |
|


|
|
abi4/MA0123.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.73
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCSGSCGSC
CGGTGCCCCC |
|


|
|
ERF4/MA0992.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.73
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | CGCSGSCGSC
--CCGCCGCC |
|

|
|
ERF3/MA1005.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.72
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCSGSCGSC
CGCCGCCA-- |
|


|
|
RAP2-6/MA1052.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.72
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CGCSGSCGSC
GCGCCGCC--- |
|


|
|
Mad/MA0535.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.72
| | Offset: | -4
| | Orientation: | forward strand |
| Alignment: | ----CGCSGSCGSC-
CAGGCGCCGCCGCCG |
|


|
|
NtERF2(AP2/EREBP)/Nicotiana tabacum/AthaMap
| Match Rank: | 8 |
| Score: | 0.71
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCSGSCGSC
CGCCGCC--- |
|


|
|
CRF4/MA0976.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.71
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CGCSGSCGSC
GCGCCGCC--- |
|


|
|
RAP2-10/MA0980.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.71
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CGCSGSCGSC
GCGCCGCC--- |
|


|
|





















