
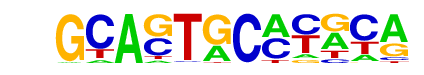











| p-value: | 1e-977 |
| log p-value: | -2.251e+03 |
| Information Content per bp: | 1.760 |
| Number of Target Sequences with motif | 5773.0 |
| Percentage of Target Sequences with motif | 13.77% |
| Number of Background Sequences with motif | 2224.9 |
| Percentage of Background Sequences with motif | 3.07% |
| Average Position of motif in Targets | 173.0 +/- 84.8bp |
| Average Position of motif in Background | 222.2 +/- 161.6bp |
| Strand Bias (log2 ratio + to - strand density) | 0.0 |
| Multiplicity (# of sites on avg that occur together) | 1.03 |
| Motif File: | file (matrix) reverse opposite |
| Rank | Match Score | Redundant Motif | P-value | log P-value | % of Targets | % of Background | Motif file |
| 1 | 0.942 |  | 1e-852 | -1963.083337 | 18.37% | 6.25% | motif file (matrix) |
| 2 | 0.796 |  | 1e-745 | -1716.188825 | 10.64% | 2.36% | motif file (matrix) |
| 3 | 0.923 |  | 1e-702 | -1617.562632 | 19.91% | 8.16% | motif file (matrix) |
| 4 | 0.836 |  | 1e-383 | -883.143333 | 28.62% | 17.87% | motif file (matrix) |
| 5 | 0.753 |  | 1e-374 | -862.134696 | 18.93% | 10.13% | motif file (matrix) |
| 6 | 0.656 |  | 1e-369 | -851.047629 | 3.41% | 0.35% | motif file (matrix) |
| 7 | 0.626 |  | 1e-322 | -741.948155 | 9.89% | 4.08% | motif file (matrix) |
| 8 | 0.818 |  | 1e-321 | -741.014798 | 16.74% | 9.00% | motif file (matrix) |
| 9 | 0.852 |  | 1e-197 | -453.854496 | 62.32% | 53.23% | motif file (matrix) |
| 10 | 0.710 |  | 1e-19 | -45.469087 | 6.36% | 5.06% | motif file (matrix) |
| 11 | 0.732 |  | 1e-3 | -8.157880 | 5.86% | 5.37% | motif file (matrix) |