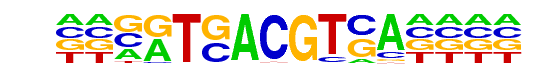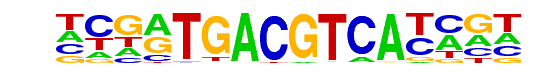| p-value: | 1e-26 |
| log p-value: | -6.210e+01 |
| Information Content per bp: | 1.721 |
| Number of Target Sequences with motif | 379.0 |
| Percentage of Target Sequences with motif | 11.71% |
| Number of Background Sequences with motif | 264.2 |
| Percentage of Background Sequences with motif | 5.12% |
| Average Position of motif in Targets | 135.6 +/- 82.4bp |
| Average Position of motif in Background | 145.9 +/- 156.7bp |
| Strand Bias (log2 ratio + to - strand density) | 0.0 |
| Multiplicity (# of sites on avg that occur together) | 1.14 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
TGA7/MA1070.1/Jaspar
| Match Rank: | 1 |
| Score: | 0.92
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA
GGTGACGTCA |
|


|
|
TGA6/MA1069.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.92
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA
GATGACGTCA |
|


|
|
CRE(bZIP)/Promoter/Homer
| Match Rank: | 3 |
| Score: | 0.91
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -SRTSACGTSA-
CGGTGACGTCAC |
|


|
|
Atf1(bZIP)/K562-ATF1-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 4 |
| Score: | 0.91
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA
GATGACGTCA |
|


|
|
CST6/MA0286.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.90
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA
-ATGACGTAA |
|


|
|
CREB1/MA0018.2/Jaspar
| Match Rank: | 6 |
| Score: | 0.90
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA
--TGACGTCA |
|


|
|
STF1(bZIP)/Glycine max/AthaMap
| Match Rank: | 7 |
| Score: | 0.89
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA--
AATGACGTCATT |
|


|
|
PB0004.1_Atf1_1/Jaspar
| Match Rank: | 8 |
| Score: | 0.89
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --SRTSACGTSA----
NCGATGACGTCATCGN |
|
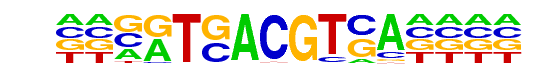
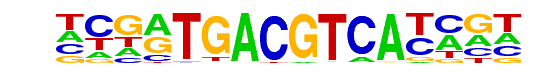
|
|
TGA1/MA0588.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.89
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -SRTSACGTSA
TGGTGACGTAA |
|


|
|
BZIP60/MA0967.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.89
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | SRTSACGTSA
--TGACGTCA |
|


|
|