| p-value: | 1e-60 |
| log p-value: | -1.393e+02 |
| Information Content per bp: | 1.719 |
| Number of Target Sequences with motif | 279.0 |
| Percentage of Target Sequences with motif | 8.62% |
| Number of Background Sequences with motif | 65.7 |
| Percentage of Background Sequences with motif | 1.27% |
| Average Position of motif in Targets | 144.4 +/- 87.5bp |
| Average Position of motif in Background | 167.2 +/- 87.0bp |
| Strand Bias (log2 ratio + to - strand density) | -0.3 |
| Multiplicity (# of sites on avg that occur together) | 1.03 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
ZNF143|STAF(Zf)/CUTLL-ZNF143-ChIP-Seq(GSE29600)/Homer
| Match Rank: | 1 |
| Score: | 0.95
| | Offset: | -2
| | Orientation: | forward strand |
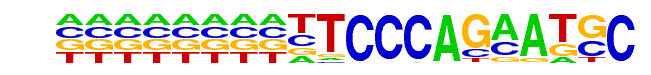
| Alignment: | --TTCCCAGAATGC-
ATTTCCCAGVAKSCY |
|


|
|
ZNF143/MA0088.2/Jaspar
| Match Rank: | 2 |
| Score: | 0.79
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TTCCCAGAATGC----
TACCCACAATGCATTG |
|


|
|
STAT3/MA0144.2/Jaspar
| Match Rank: | 3 |
| Score: | 0.75
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TTCCCAGAATGC
TTTCCCAGAAN-- |
|


|
|
Rbpj1(?)/Panc1-Rbpj1-ChIP-Seq(GSE47459)/Homer
| Match Rank: | 4 |
| Score: | 0.73
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --TTCCCAGAATGC
HTTTCCCASG---- |
|


|
|
GFY-Staf(?,Zf)/Promoter/Homer
| Match Rank: | 5 |
| Score: | 0.72
| | Offset: | -8
| | Orientation: | forward strand |
| Alignment: | --------TTCCCAGAATGC
AACTACAATTCCCAGAATGC |
|

|
|
Stat5a::Stat5b/MA0519.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.72
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --TTCCCAGAATGC
ATTTCCAAGAA--- |
|


|
|
ZNF416(Zf)/HEK293-ZNF416.GFP-ChIP-Seq(GSE58341)/Homer
| Match Rank: | 7 |
| Score: | 0.69
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TTCCCAGAATGC
TGCCCAGNHW-- |
|


|
|
STAT1/MA0137.3/Jaspar
| Match Rank: | 8 |
| Score: | 0.68
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TTCCCAGAATGC
TTTCCTGGAAA-- |
|


|
|
Su(H)/dmmpmm(Bergman)/fly
| Match Rank: | 9 |
| Score: | 0.67
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TTCCCAGAATGC
CTCCCAC----- |
|


|
|
PB0206.1_Zic2_2/Jaspar
| Match Rank: | 10 |
| Score: | 0.66
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TTCCCAGAATGC--
CCACACAGCAGGAGA |
|


|
|





















