Flagstat QC (all)
Dup QC (all)
Flagstat QC (all, filtered)
PBC QC (all)
| Total Read Pairs | Distinct Read Pairs | One Read Pair | Two Read Pairs | NRF = Distinct/Total | PBC1 = OnePair/Distinct | PBC2 = OnePair/TwoPair | |
|---|---|---|---|---|---|---|---|
| rep1 | 8784426 | 7535313 | 6436973 | 965149 | 0.857804 | 0.854241 | 6.669409 |
| rep2 | 26235817 | 25032759 | 23881912 | 1103599 | 0.954144 | 0.954026 | 21.640027 |
| ctl1 | 11609614 | 11458969 | 11311780 | 144942 | 0.987024 | 0.987155 | 78.043493 |
NRF (non redundant fraction)
PBC1 (PCR Bottleneck coefficient 1)
PBC2 (PCR Bottleneck coefficient 2)
PBC1 is the primary measure. Provisionally
- 0-0.5 is severe bottlenecking
- 0.5-0.8 is moderate bottlenecking
- 0.8-0.9 is mild bottlenecking
- 0.9-1.0 is no bottlenecking
Cross-correlation QC (all)
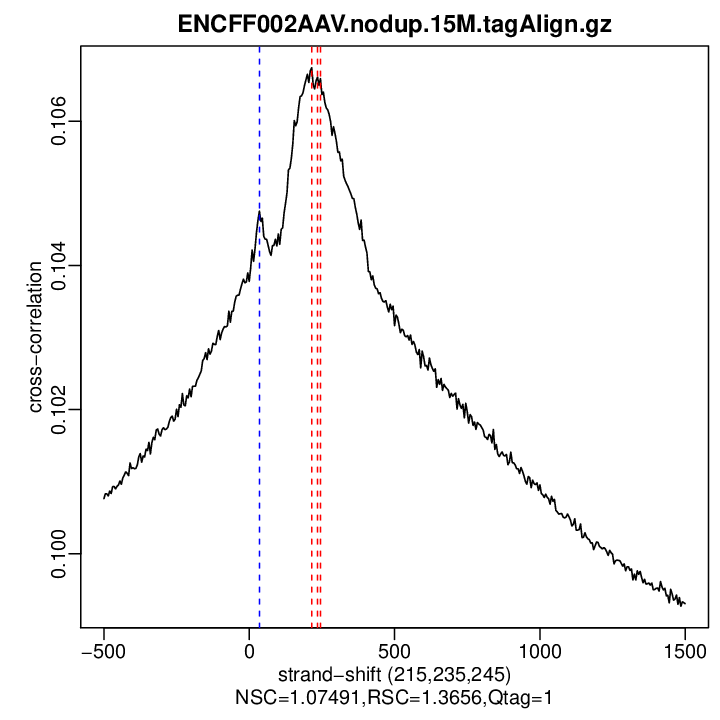
rep1 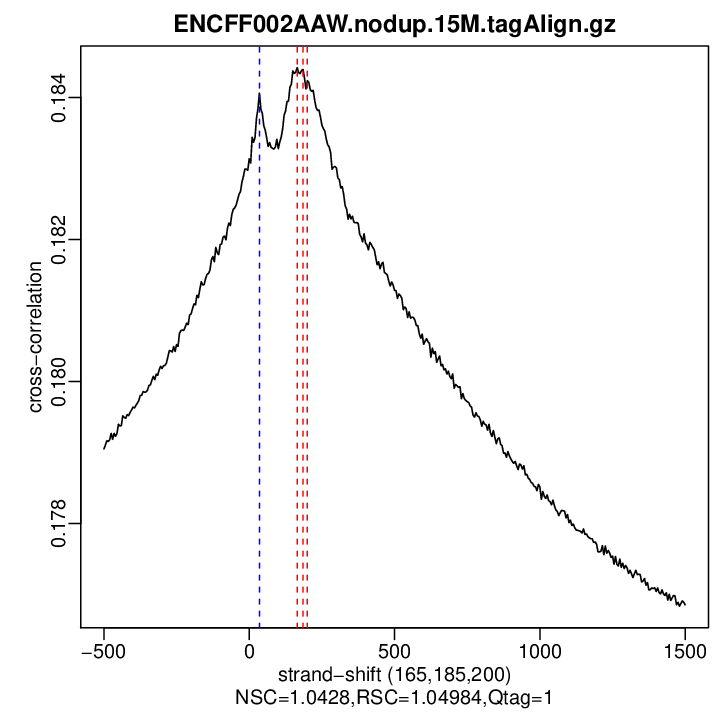
rep2
| numReads | estFragLen | corr_estFragLen | PhantomPeak | corr_phantomPeak | argmin_corr | min_corr | NSC | RSC | |
|---|---|---|---|---|---|---|---|---|---|
| rep1 | 7510000 | 215 | 0.106745496488178 | 35 | 0.104754 | 1500 | 0.09930667 | 1.074908 | 1.365596 |
| rep2 | 15000000 | 165 | 0.184421337122509 | 35 | 0.184062 | 1500 | 0.1768528 | 1.042796 | 1.049843 |
Normalized strand cross-correlation coefficient (NSC) = col9 in outFile
Relative strand cross-correlation coefficient (RSC) = col10 in outFile
Estimated fragment length = col3 in outFile, take the top value
Important columns highlighted, but all/whole file can be stored for display

