| Alignment | .
|
| Replicate 1 | .
|
| Tag-align | ./align/rep1/ENCFF002ABA.nodup.tagAlign.gz
|
| Replicate 2 | .
|
| Tag-align | ./align/rep2/H3K27acRep2.nodup.tagAlign.gz
|
| Control 1 | .
|
| Tag-align |
|
| Pooled replicate | .
|
| Tag-align | ./align/pooled_rep/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign.gz
|
| Pseudo-replicates | .
|
| Replicate 1 | .
|
| Pseudo-replicate 1 | .
|
| Tag-align | ./align/pseudo_reps/rep1/pr1/ENCFF002ABA.nodup.pr1.tagAlign.gz
|
| Pseudo-replicate 2 | .
|
| Tag-align | ./align/pseudo_reps/rep1/pr2/ENCFF002ABA.nodup.pr2.tagAlign.gz
|
| Replicate 2 | .
|
| Pseudo-replicate 1 | .
|
| Tag-align | ./align/pseudo_reps/rep2/pr1/H3K27acRep2.nodup.pr1.tagAlign.gz
|
| Pseudo-replicate 2 | .
|
| Tag-align | ./align/pseudo_reps/rep2/pr2/H3K27acRep2.nodup.pr2.tagAlign.gz
|
| Pooled pseudo-replicates | .
|
| Pooled pseudo-replicate 1 | .
|
| Tag-align | ./align/pooled_pseudo_reps/ppr1/ENCFF002ABA.nodup.pr1_H3K27acRep2.nodup.pr1.tagAlign.gz
|
| Pooled pseudo-replicate 2 | .
|
| Tag-align | ./align/pooled_pseudo_reps/ppr2/ENCFF002ABA.nodup.pr2_H3K27acRep2.nodup.pr2.tagAlign.gz
|
| Signal tracks | .
|
| MACS2 | .
|
| Replicate 1 | .
|
| P-value | ./signal/macs2/rep1/ENCFF002ABA.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.pval.signal.bw
|
| Fold enrichment | ./signal/macs2/rep1/ENCFF002ABA.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.fc.signal.bw
|
| Replicate 2 | .
|
| P-value | ./signal/macs2/rep2/H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.pval.signal.bw
|
| Fold enrichment | ./signal/macs2/rep2/H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.fc.signal.bw
|
| Pooled replicate | .
|
| P-value | ./signal/macs2/pooled_rep/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.pval.signal.bw
|
| Fold enrichment | ./signal/macs2/pooled_rep/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.fc.signal.bw
|
| Peaks | .
|
| MACS2 | .
|
| Replicate 1 | .
|
| Narrow peak | ./peak/macs2/rep1/ENCFF002ABA.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.narrowPeak.gz
|
| Broad peak | ./peak/macs2/rep1/ENCFF002ABA.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.broadPeak.gz
|
| Gapped peak | ./peak/macs2/rep1/ENCFF002ABA.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.gappedPeak.gz
|
| Replicate 2 | .
|
| Narrow peak | ./peak/macs2/rep2/H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.narrowPeak.gz
|
| Broad peak | ./peak/macs2/rep2/H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.broadPeak.gz
|
| Gapped peak | ./peak/macs2/rep2/H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.gappedPeak.gz
|
| Pooled replicate | .
|
| Narrow peak | ./peak/macs2/pooled_rep/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.narrowPeak.gz
|
| Broad peak | ./peak/macs2/pooled_rep/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.broadPeak.gz
|
| Gapped peak | ./peak/macs2/pooled_rep/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.gappedPeak.gz
|
| Pooled pseudo-replicate | .
|
| Pooled pseudo-replicate 1 | .
|
| Broad peak | ./peak/macs2/pooled_pseudo_reps/ppr2/ENCFF002ABA.nodup.pr2_H3K27acRep2.nodup.pr2.tagAlign_x_ENCFF001HNC.nodup.tagAlign.broadPeak.gz
|
| Gapped peak | ./peak/macs2/pooled_pseudo_reps/ppr2/ENCFF002ABA.nodup.pr2_H3K27acRep2.nodup.pr2.tagAlign_x_ENCFF001HNC.nodup.tagAlign.gappedPeak.gz
|
| Pooled pseudo-replicate 2 | .
|
| Narrow peak | ./peak/macs2/pooled_pseudo_reps/ppr2/ENCFF002ABA.nodup.pr2_H3K27acRep2.nodup.pr2.tagAlign_x_ENCFF001HNC.nodup.tagAlign.narrowPeak.gz
|
| Naive overlap | .
|
| Broad peak | ./peak/macs2/overlap/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.naive_overlap.filt.gappedPeak.gz
|
| Gapped peak | ./peak/macs2/overlap/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.naive_overlap.filt.gappedPeak.gz
|
| Peak | ./peak/macs2/overlap/ENCFF002ABA.nodup_H3K27acRep2.nodup.tagAlign_x_ENCFF001HNC.nodup.tagAlign.naive_overlap.filt.narrowPeak.gz
|
| QC and logs | .
|
| Replicate 1 | .
|
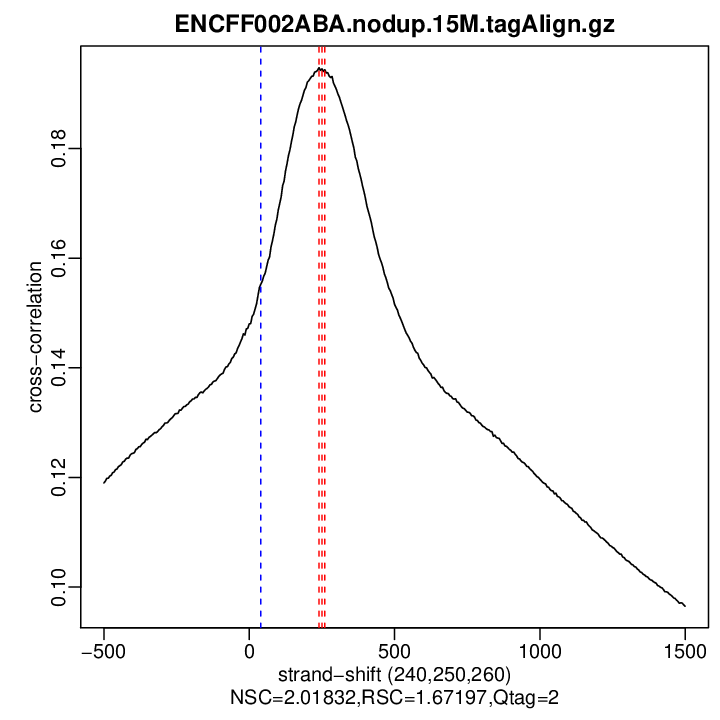
| Cross-corr. log | ./qc/rep1/ENCFF002ABA.nodup.15M.cc.qc
|
| Cross-corr. plot | ./qc/rep1/ENCFF002ABA.nodup.15M.cc.plot.pdf
|
| Replicate 2 | .
|
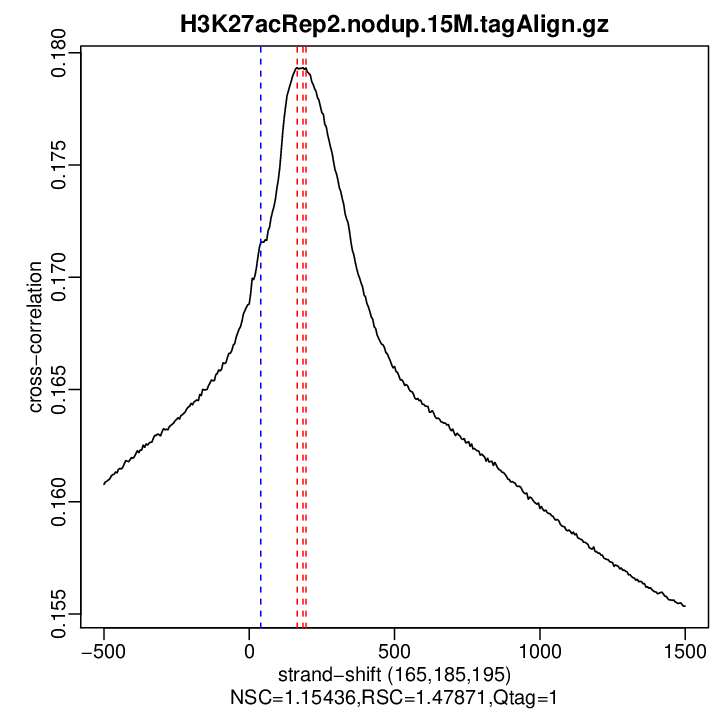
| Cross-corr. log | ./qc/rep2/H3K27acRep2.nodup.15M.cc.qc
|
| Cross-corr. plot | ./qc/rep2/H3K27acRep2.nodup.15M.cc.plot.pdf
|