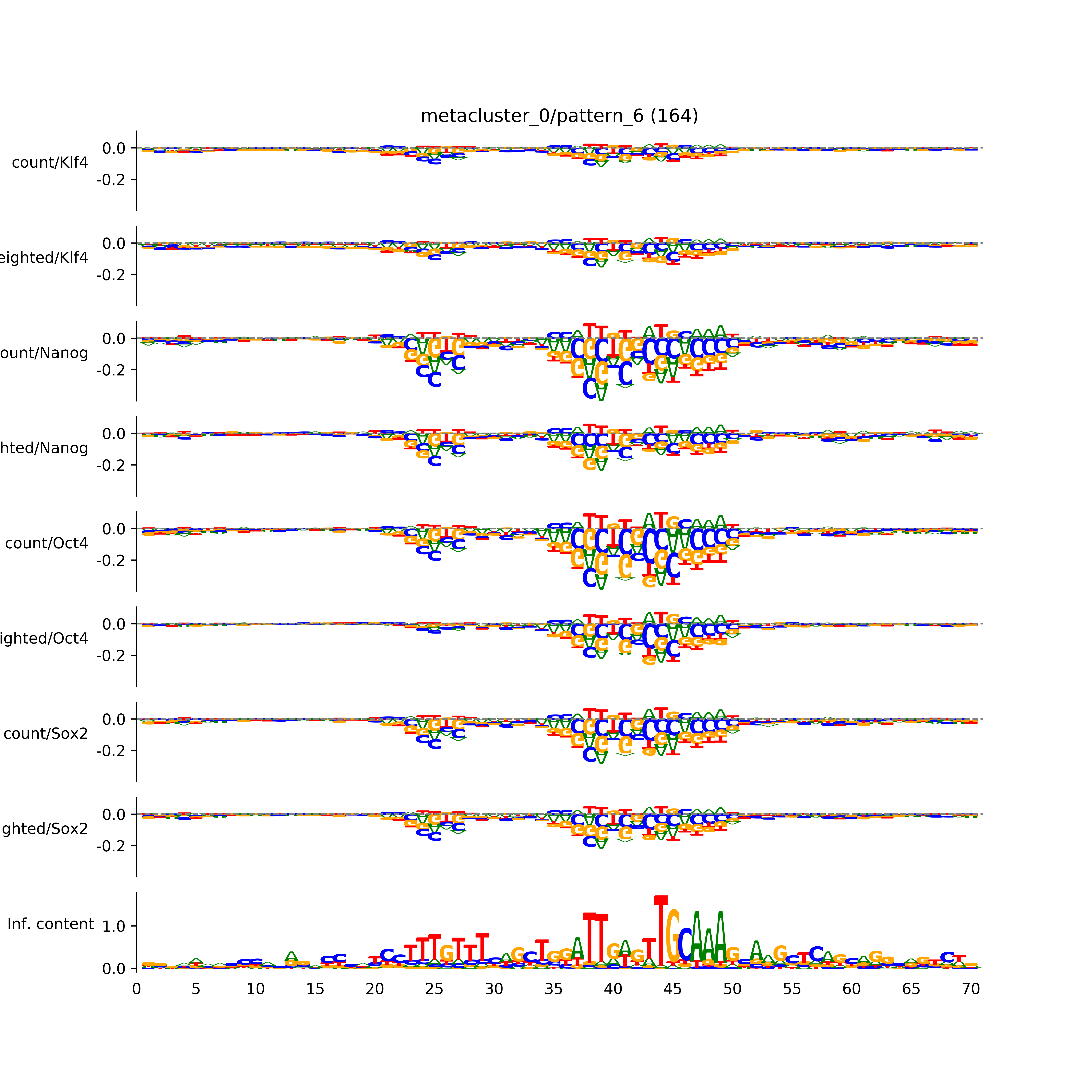
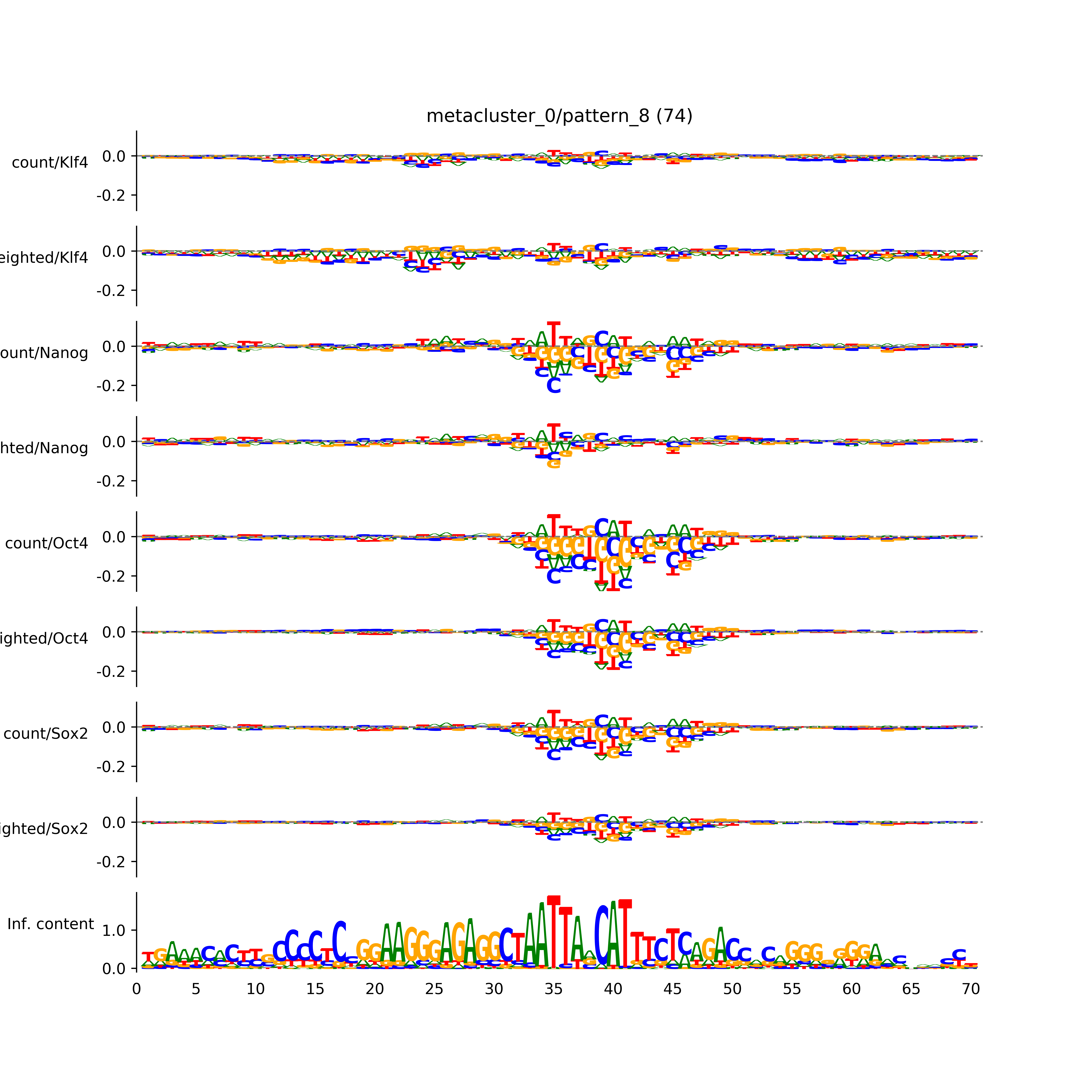
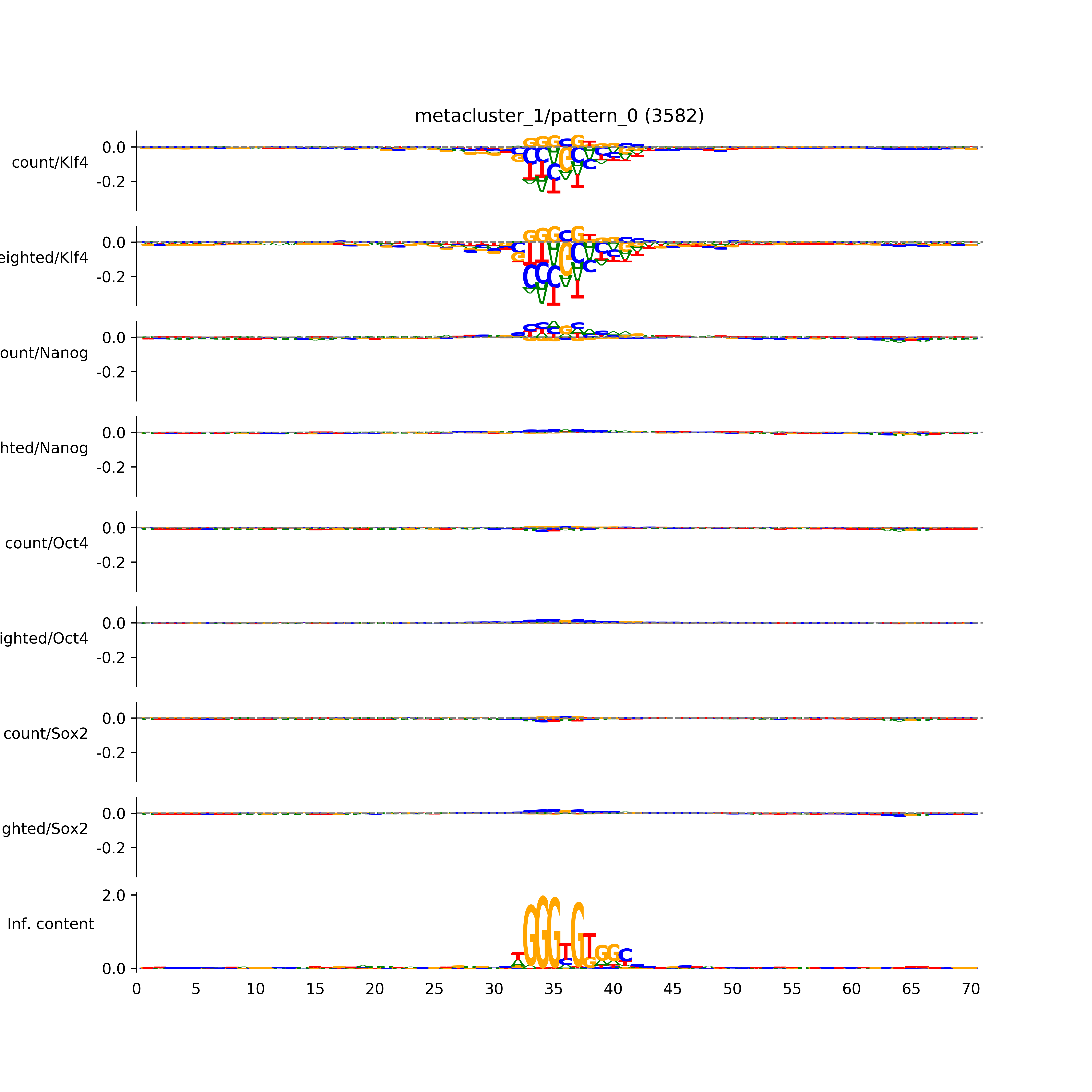
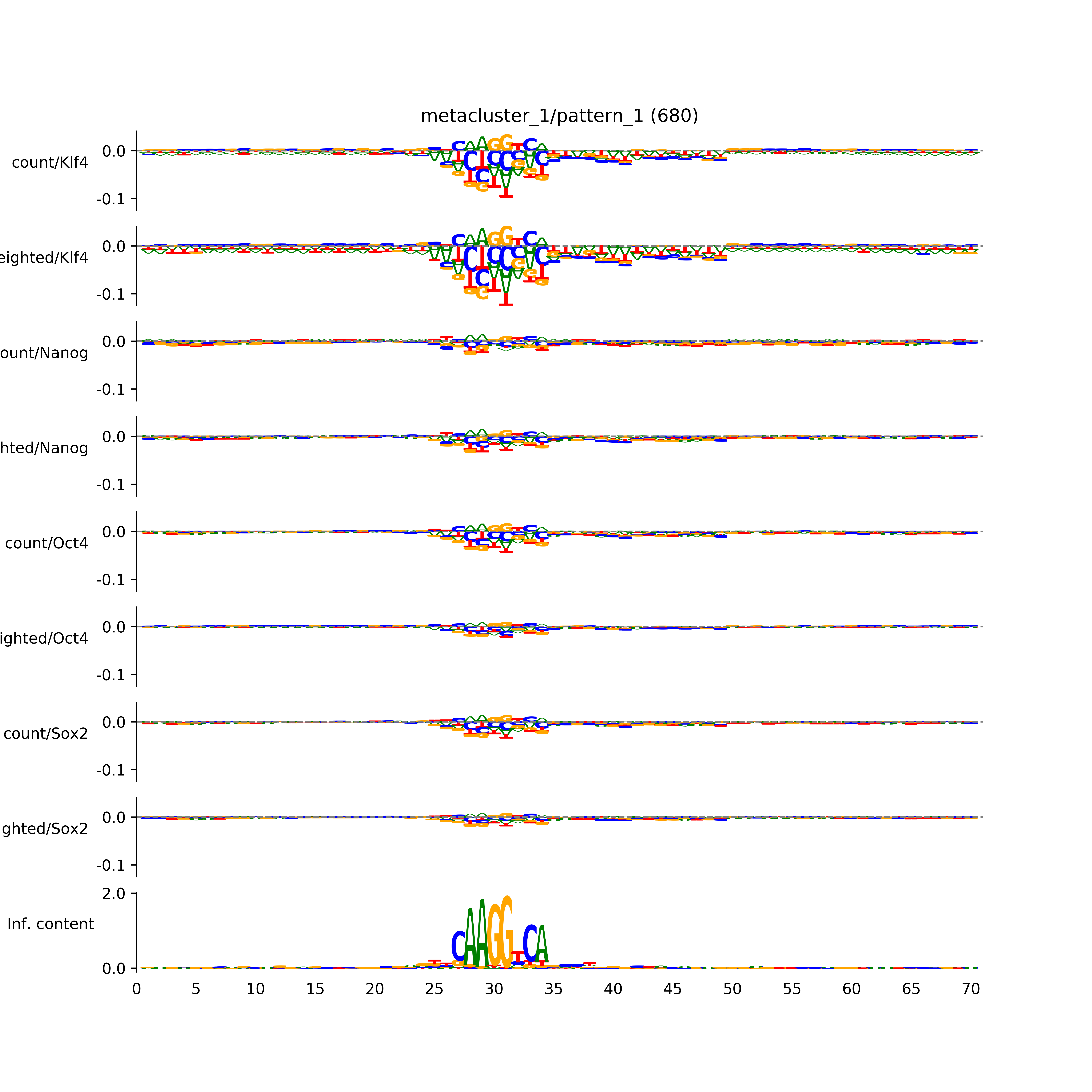
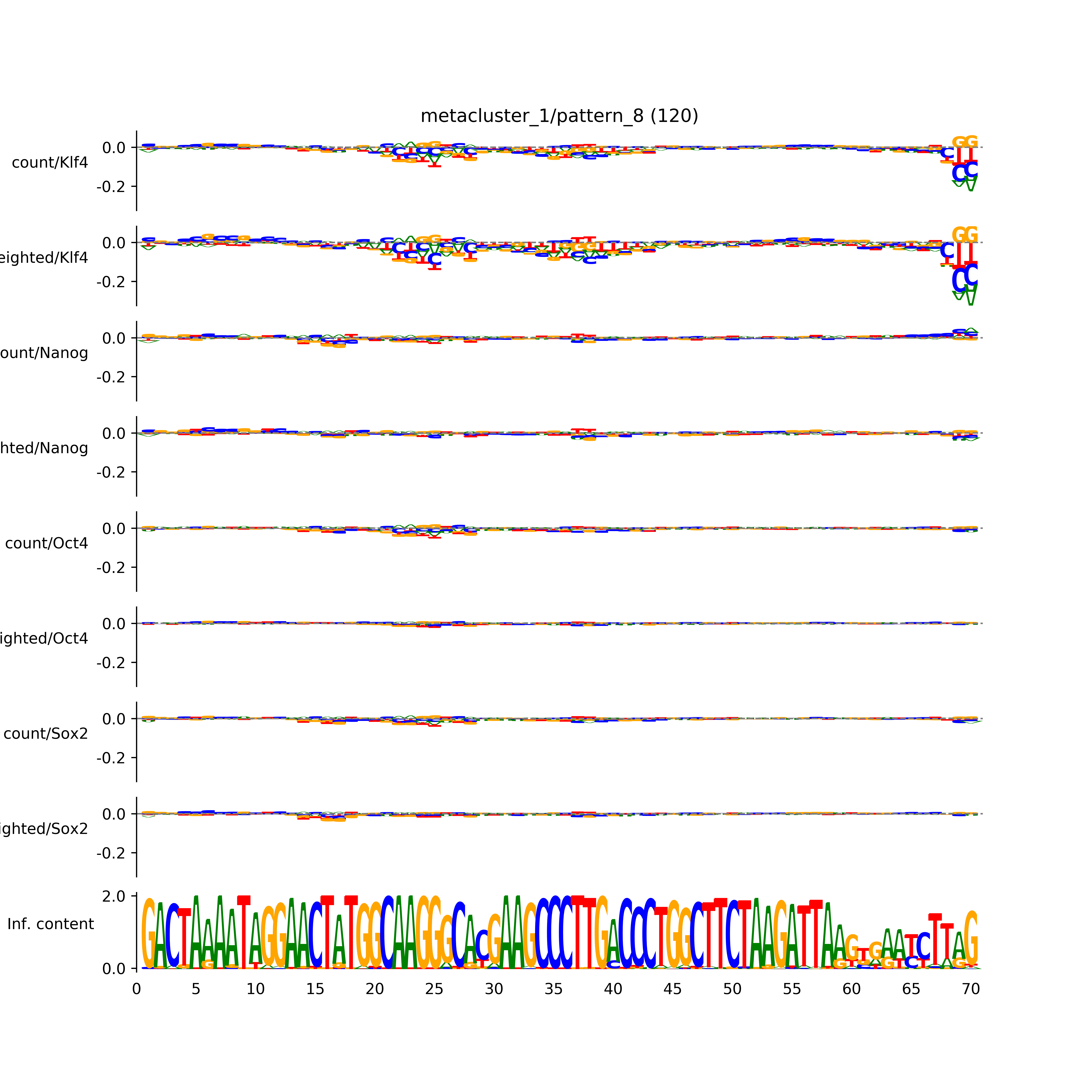
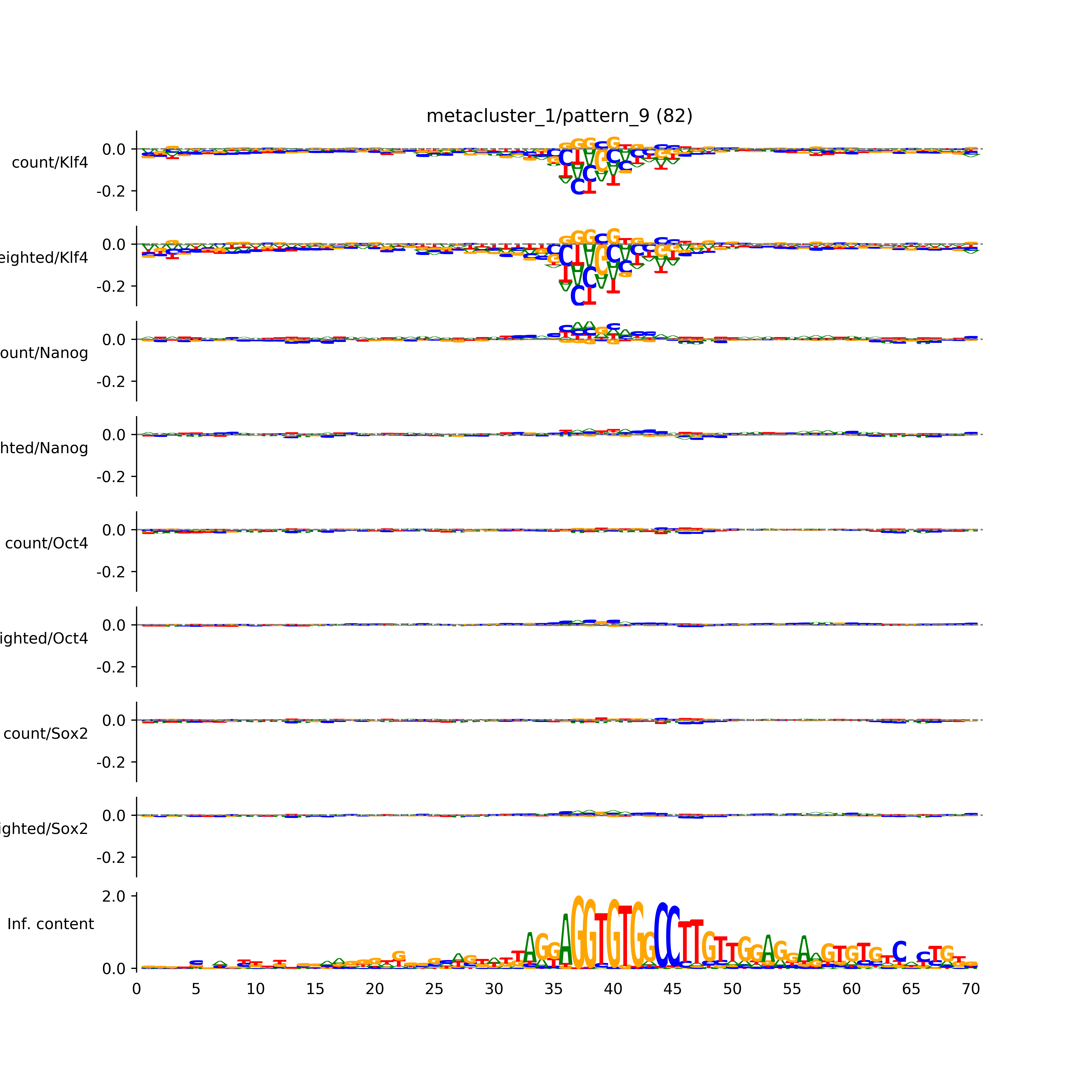
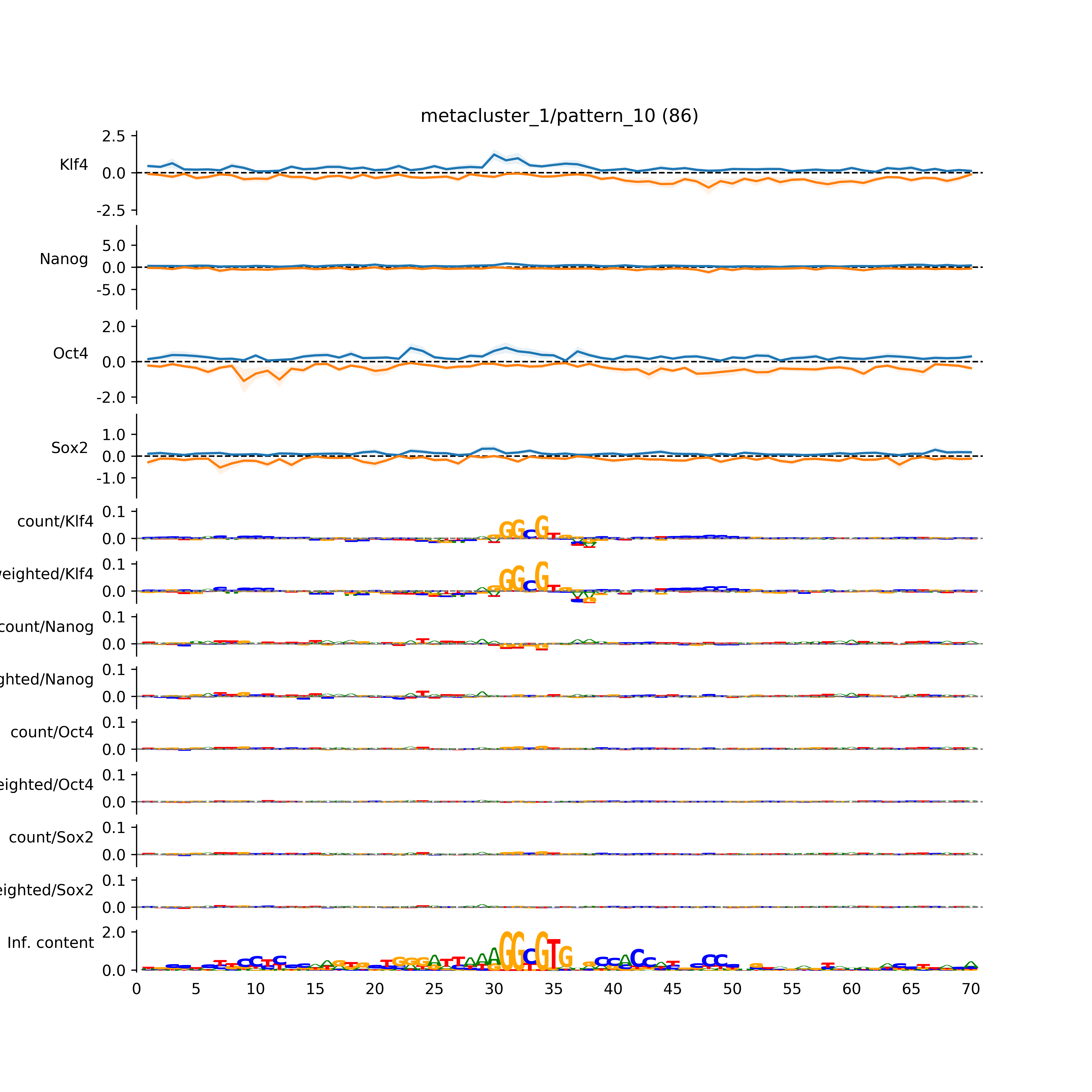
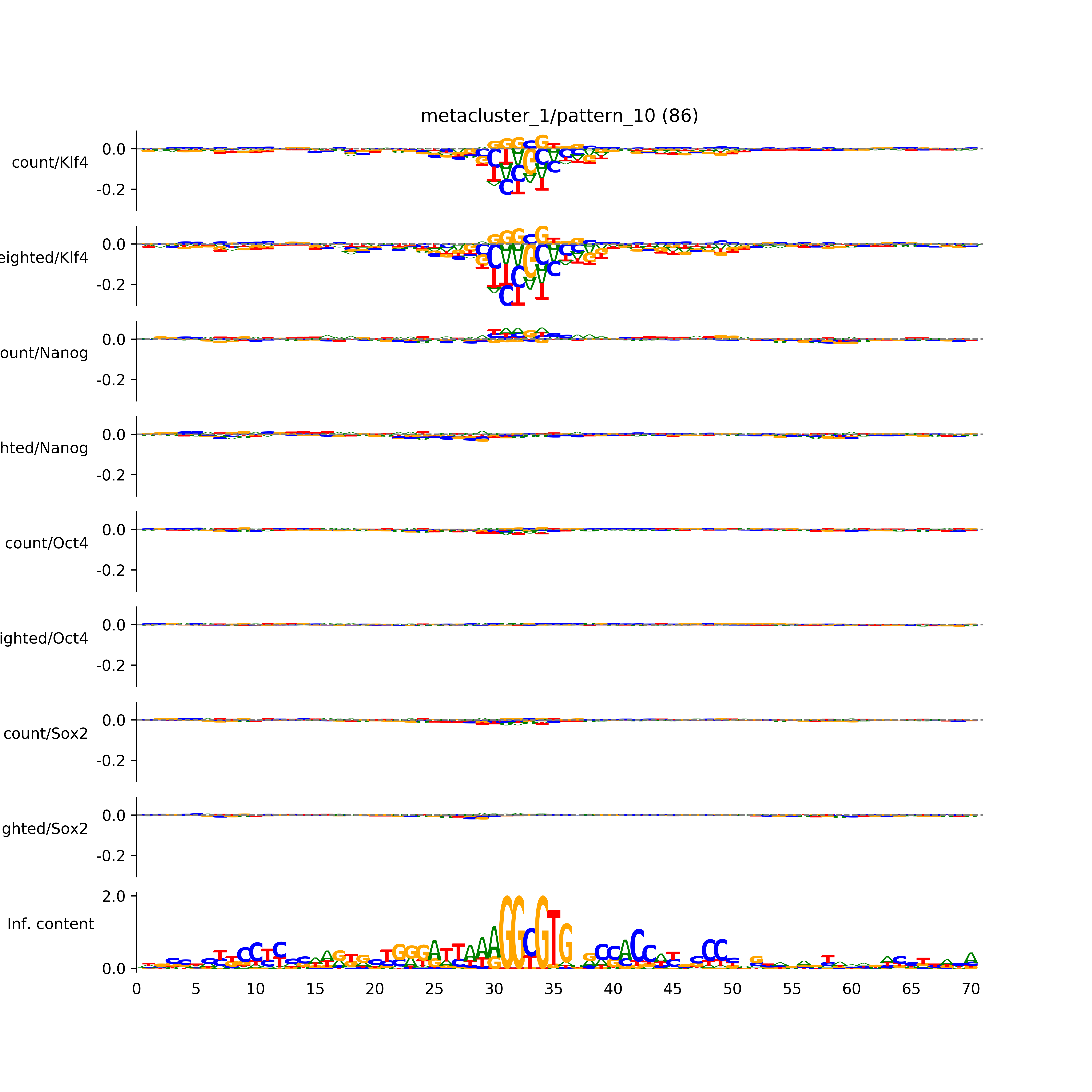
BPNet tf-Modisco report¶
In [1]:
url_dir = "http://mitra.stanford.edu/kundaje/avsec/chipnexus/oct-sox-nanog-klf/models/n_dil_layers=9/modisco/valid/new-hparams/plots/"
modisco_dir = "/users/avsec/workspace/basepair/data/processed/chipnexus/exp/models/oct-sox-nanog-klf/models/n_dil_layers=9/modisco/valid/new-hparams"
In [2]:
# Parameters
url_dir = "http://mitra.stanford.edu/kundaje/avsec/chipnexus/oct-sox-nanog-klf/models/n_dil_layers=9/modisco/clustered/weighted/0/plots/"
modisco_dir = "0"
In [3]:
from basepair.modisco import ModiscoResult
from basepair.config import get_data_dir
from basepair.utils import read_json
from basepair.plot.vdom import vdom_modisco
from kipoi.readers import HDF5Reader
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
from plotnine import *
In [4]:
mr = ModiscoResult(f"{modisco_dir}/modisco.h5")
mr.open()
In [5]:
# load the data
modisco_kwargs = read_json(os.path.join(modisco_dir, "kwargs.json"))
d = HDF5Reader(modisco_kwargs['imp_scores'])
d.open()
In [6]:
strand_dist_file = f"{modisco_dir}/strand_distances.h5"
if modisco_kwargs.get("ignore_strand_dist", False) and os.path.exists(strand_dist_file):
included_samples = HDF5Reader.load(strand_dist_file)['included_samples']
else:
included_samples = np.ones(d.f['inputs'].shape[:1], dtype=bool)
if modisco_kwargs.get("filter_npy", None) is not None:
included_samples = np.load(modisco_kwargs['filter_npy']) * included_samples
id_hash = pd.DataFrame({"peak_id": d.f['/metadata/interval_from_task'][:][included_samples],
"example_idx": np.arange(d.f['/metadata/interval_from_task'][included_samples].shape[0])})
tasks = list(d.f["targets"]["profile"].keys())
In [7]:
# get all seqlet instances
dfp = mr.seqlet_df_instances().rename(columns=dict(seqname="example_idx"))
dfp = pd.merge(dfp,id_hash, on="example_idx")
In [8]:
# row = example_idx
total_counts = pd.DataFrame({task: d.f[f"/targets/profile/{task}"][:][included_samples].sum(axis=-1).sum(axis=-1)
for task in tasks
})
In [9]:
len(mr.patterns())
Out[9]:
In [10]:
# Number of metaclusters
len(mr.metaclusters())
Out[10]:
Number of seqlets per pattern¶
In [11]:
mc_stat = mr.metacluster_stats()
In [12]:
ggplot(aes(x="pattern", y='n'), mc_stat) + geom_bar(stat='identity') + \
facet_wrap("~metacluster", ncol=4, labeller='label_both') + \
ylab("Number of seqlets") + theme_classic()
Out[12]:
Zoom-into the 500 seqlet range¶
In [13]:
ggplot(aes(x="pattern", y='n'), mc_stat) + geom_bar(stat='identity') + \
facet_wrap("~metacluster", ncol=4, labeller='label_both') + \
ylab("Number of seqlets") + theme_classic() + coord_cartesian(ylim=[0, 500])
Out[13]:
Important tasks per metacluster¶
In [14]:
mcs_grouped = mc_stat.groupby("metacluster").n.agg(["count", "sum"]).reset_index()
fig, ax = plt.subplots(2, 1, sharex=False, figsize=(18,6),
gridspec_kw={'height_ratios': [2,1]})
mcs_grouped.plot("metacluster", "count",
label="# patterns per metacluster", style="o--",
ax=ax[0],
yticks=range(mcs_grouped['count'].max()+1),
xticks=range(38),
fontsize='large',
xlim=(-.5, len(mr.metaclusters()) - .5 ))
mcs_grouped.plot("metacluster", "sum",
label="# seqlets per metacluster",
style="o--", ax=ax[0], secondary_y=True)
ax[0].grid(linewidth=0.2)
mr.plot_metacluster_activity(ax[1], cbar=False)
ax[1].set_title("Importance score activity: Red = positive, Blue = negative");
In [15]:
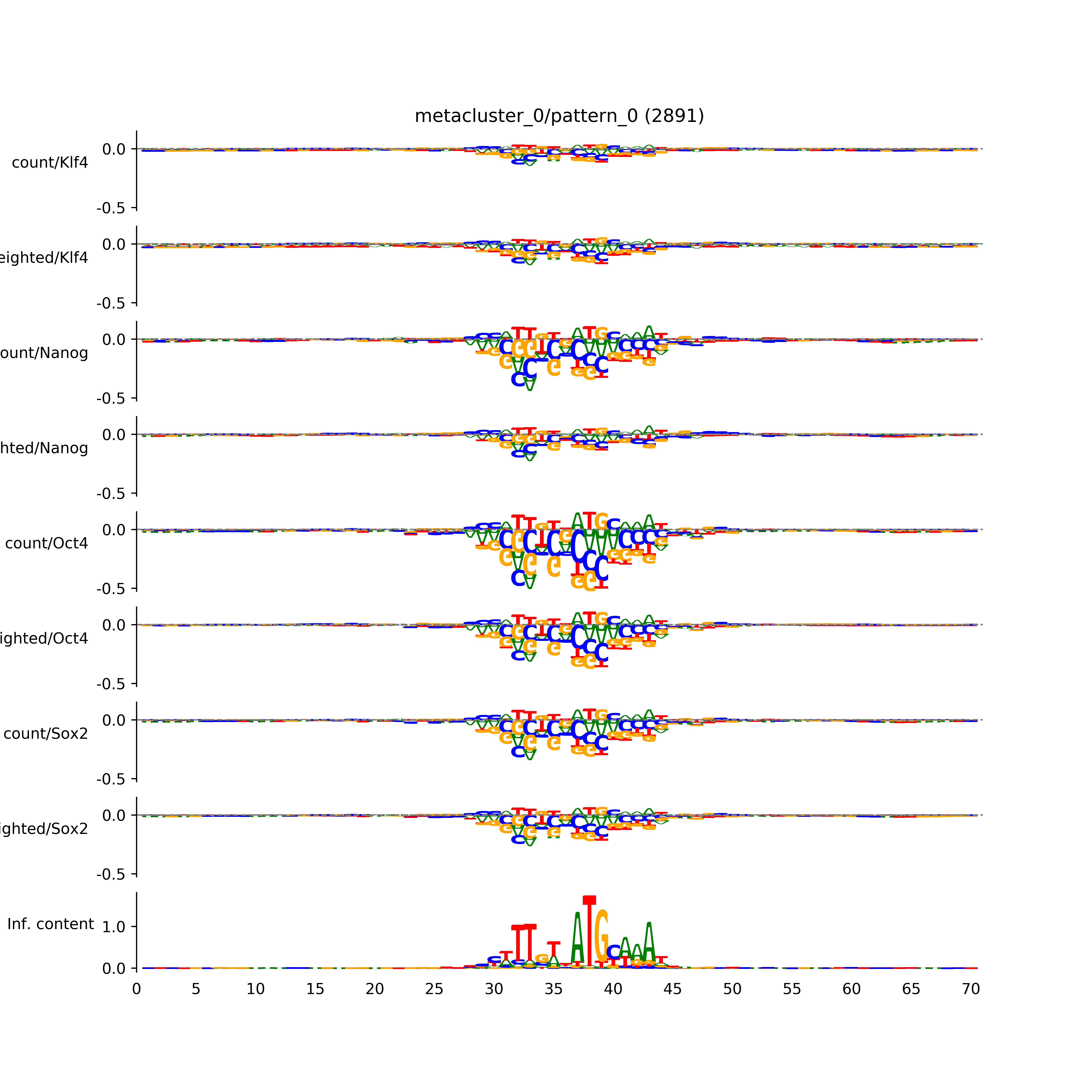
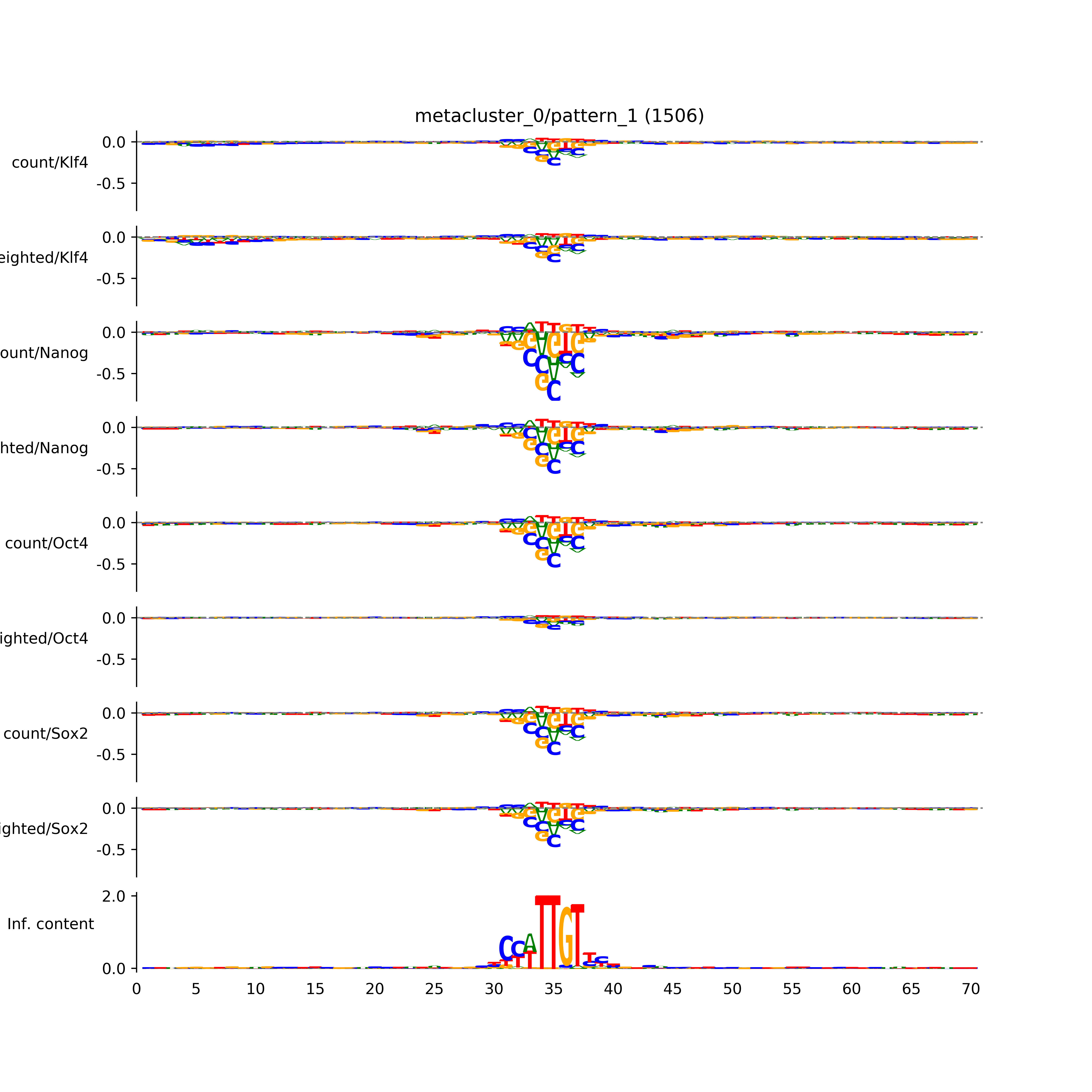
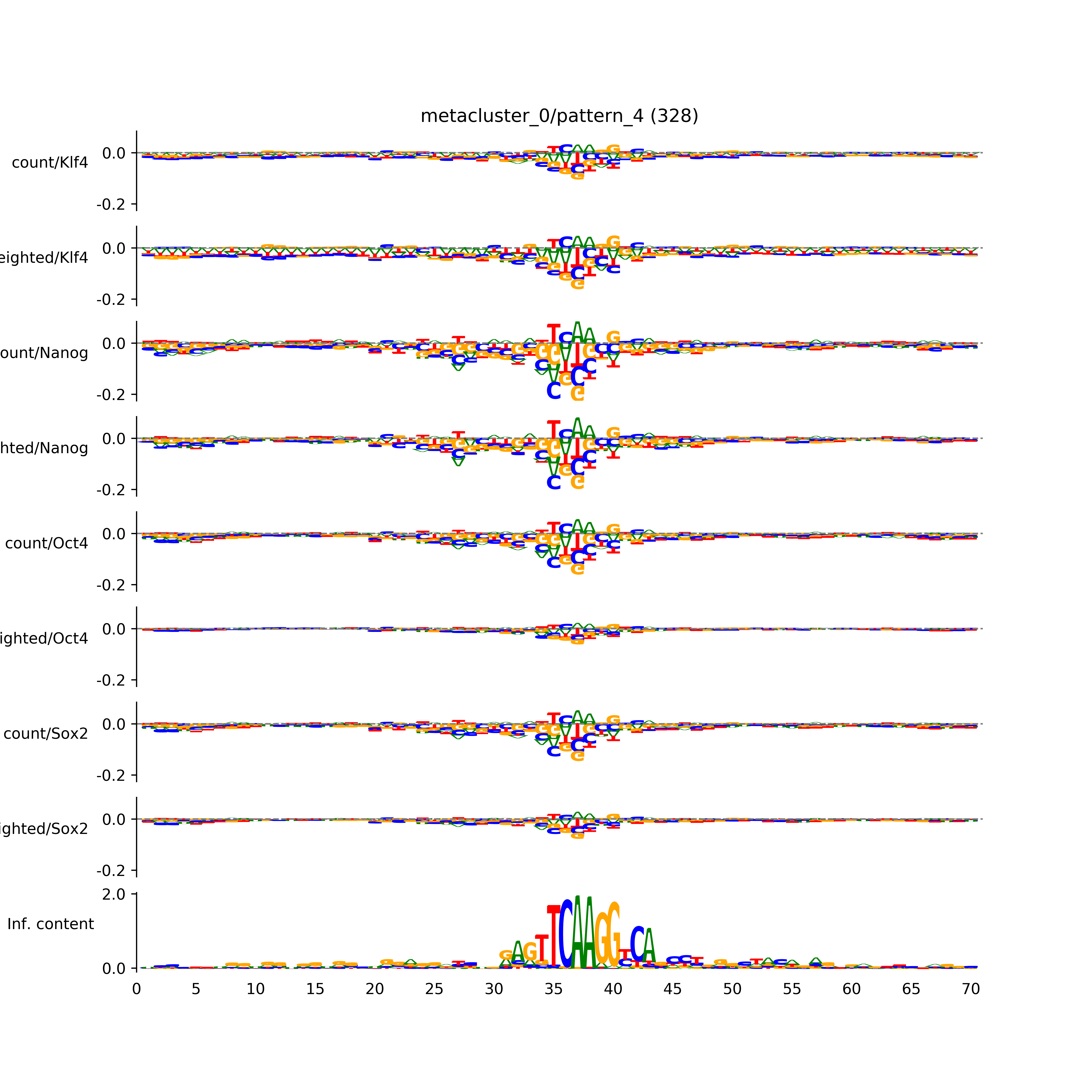
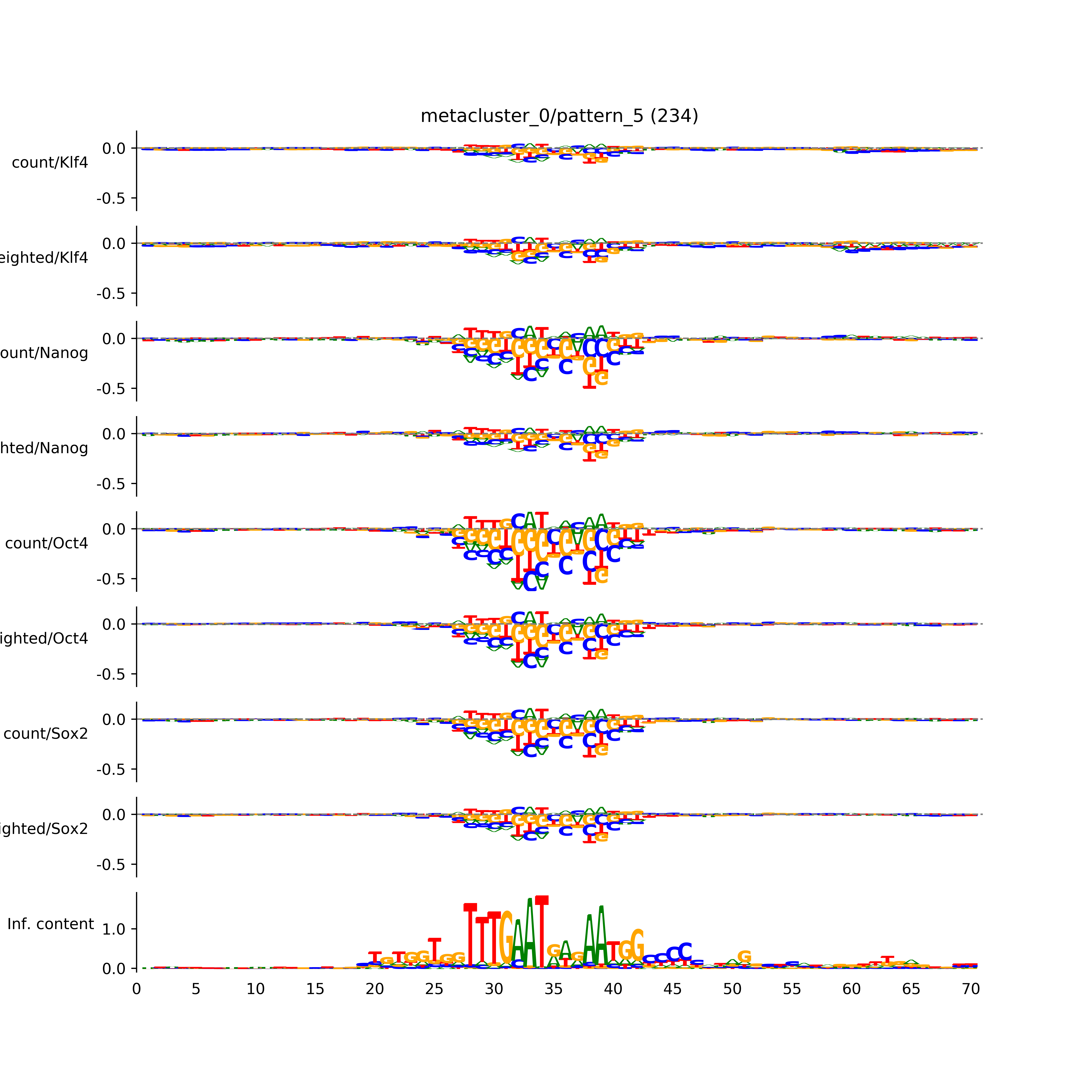
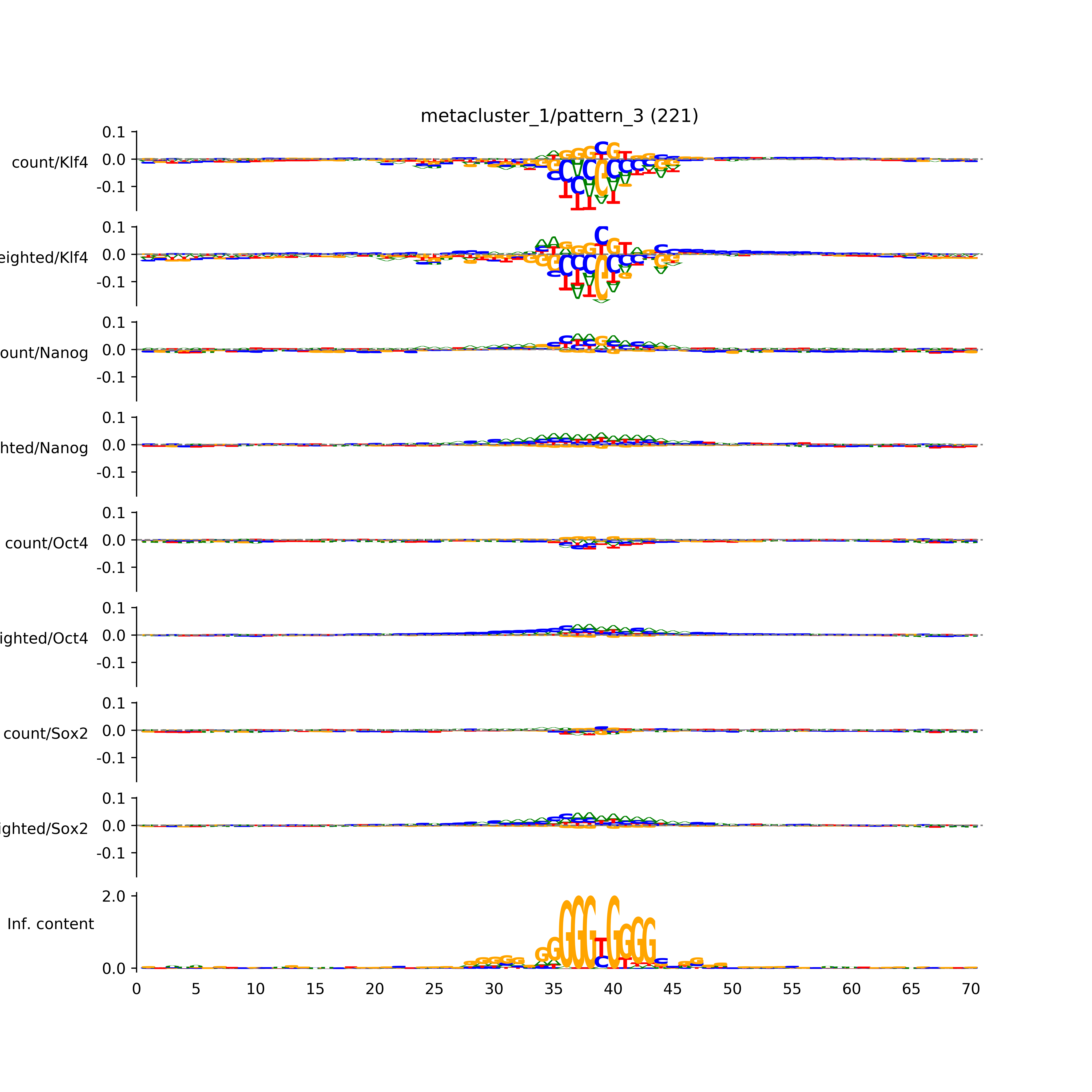
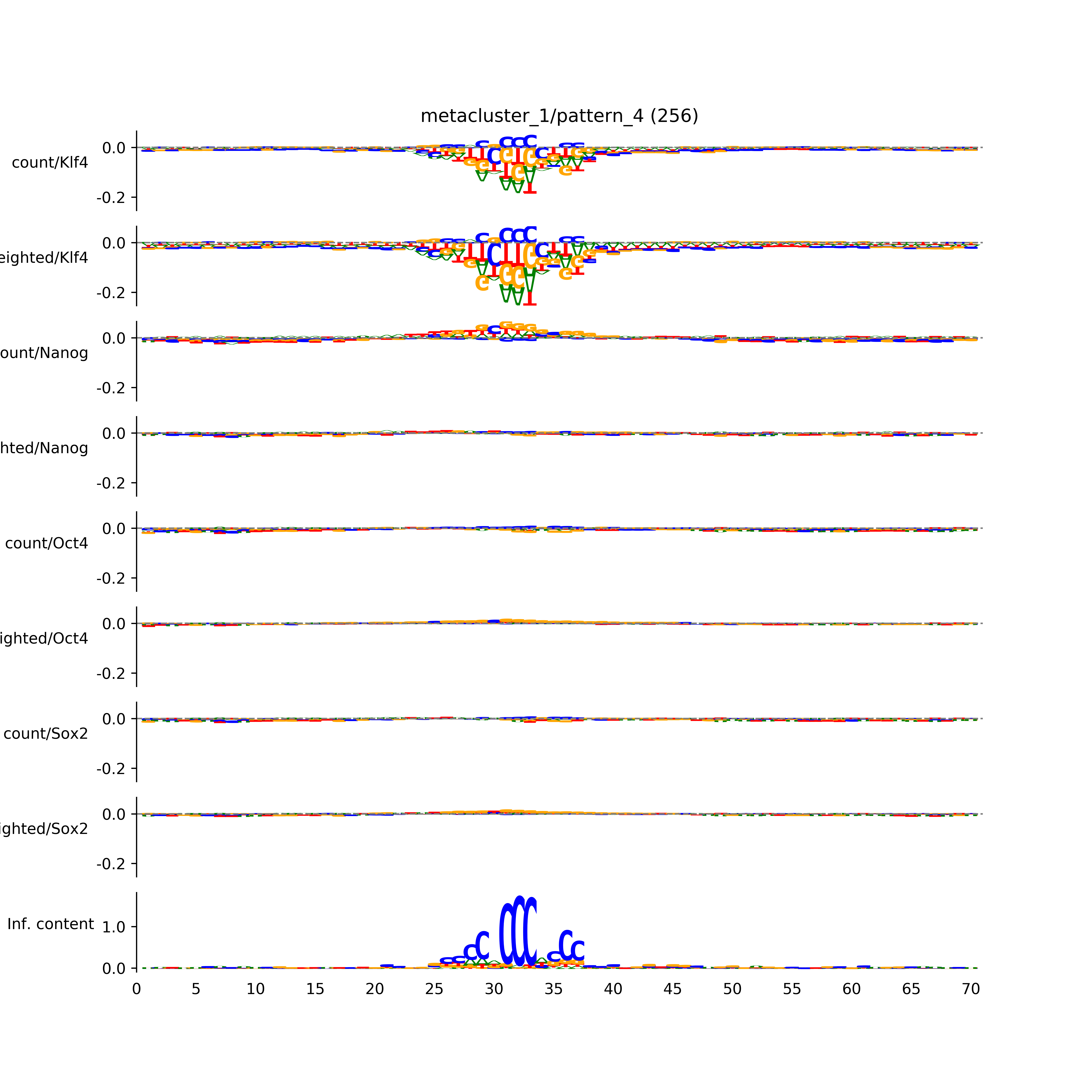
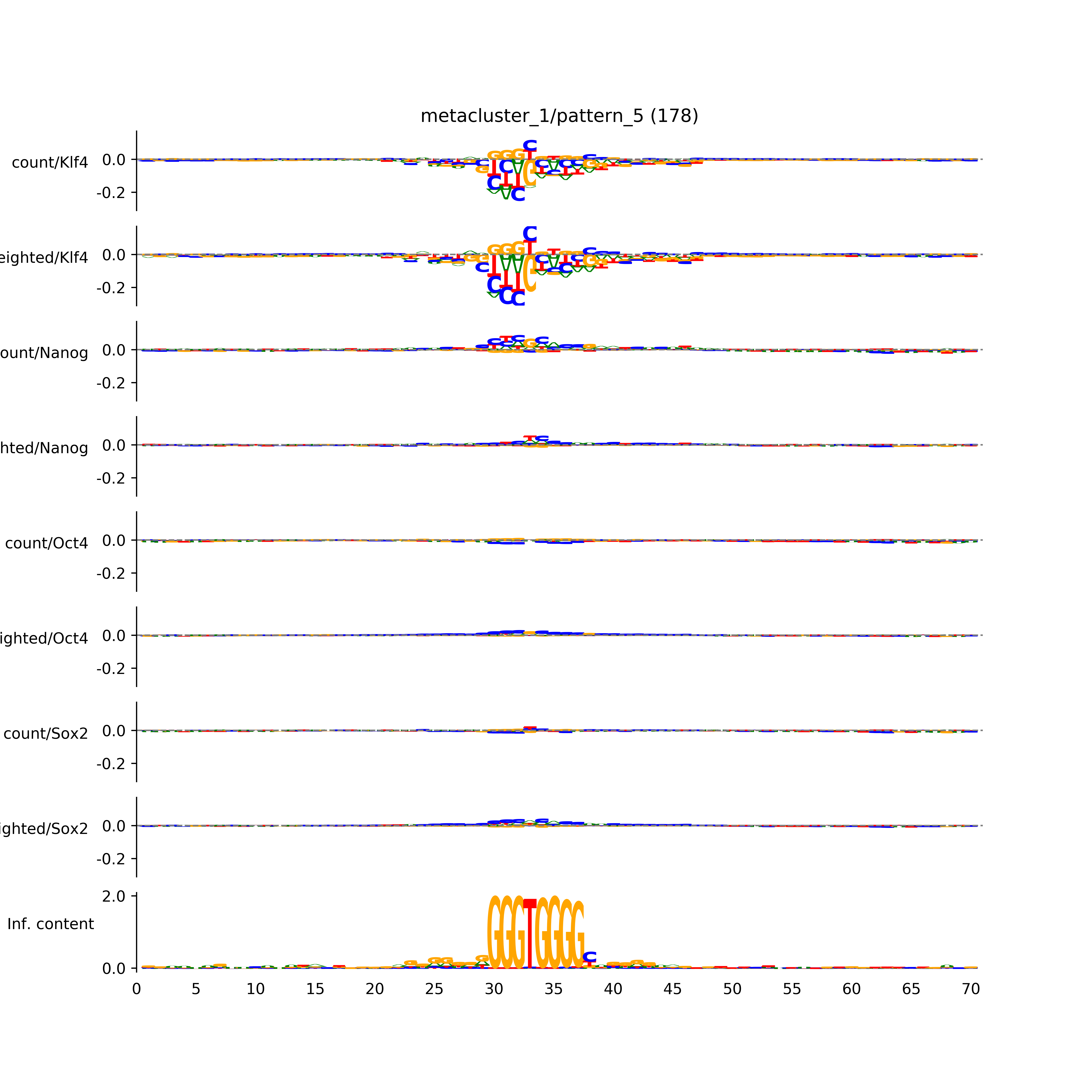
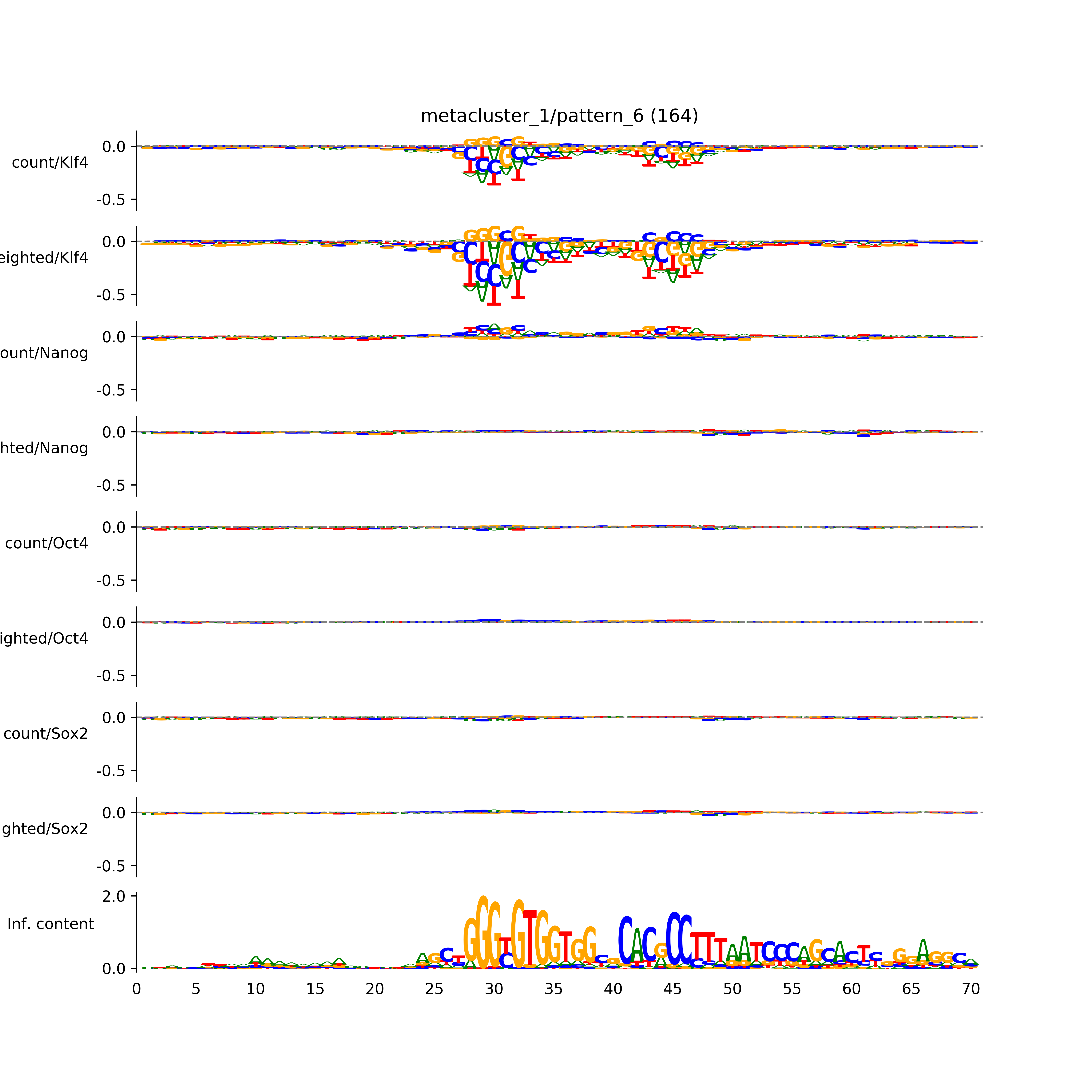
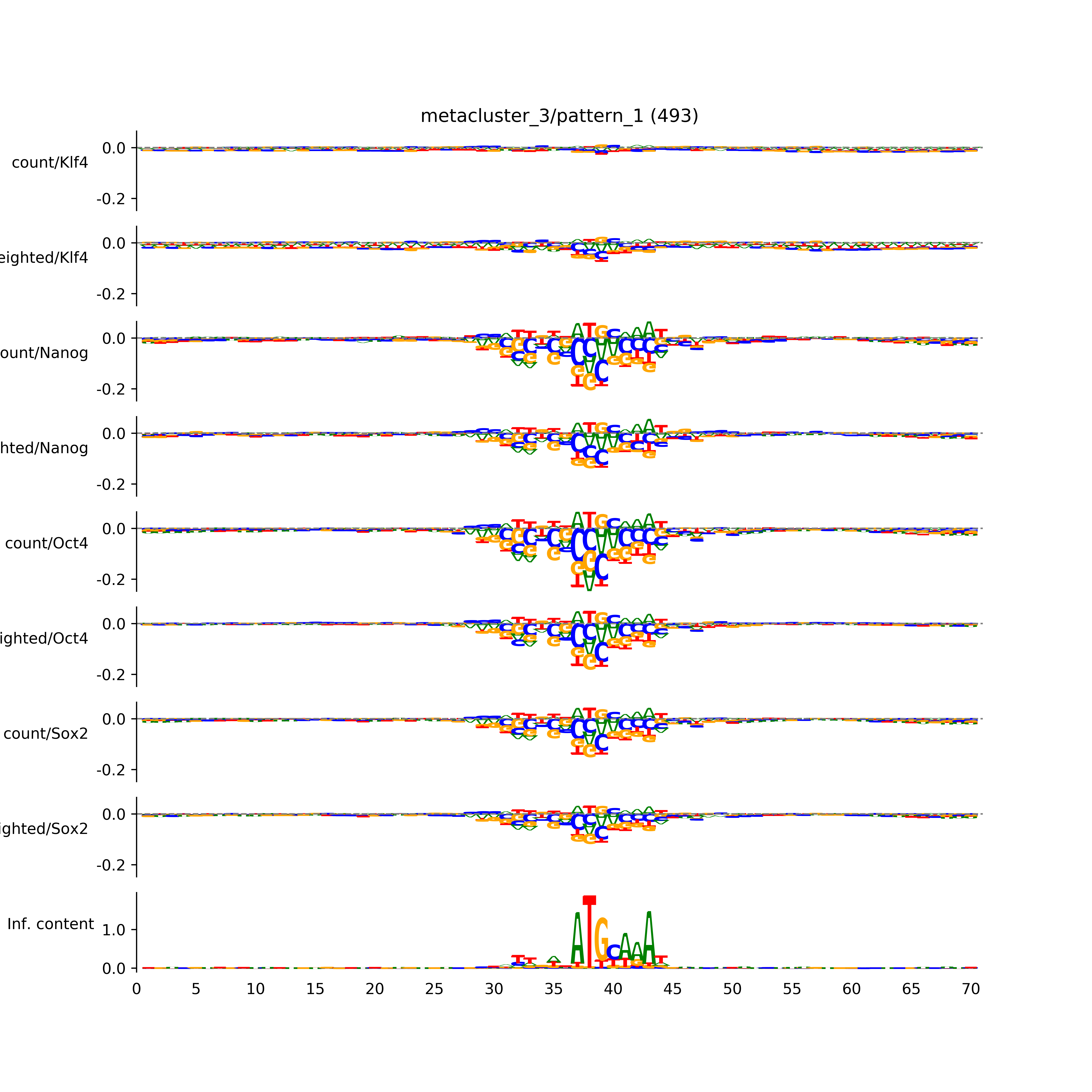
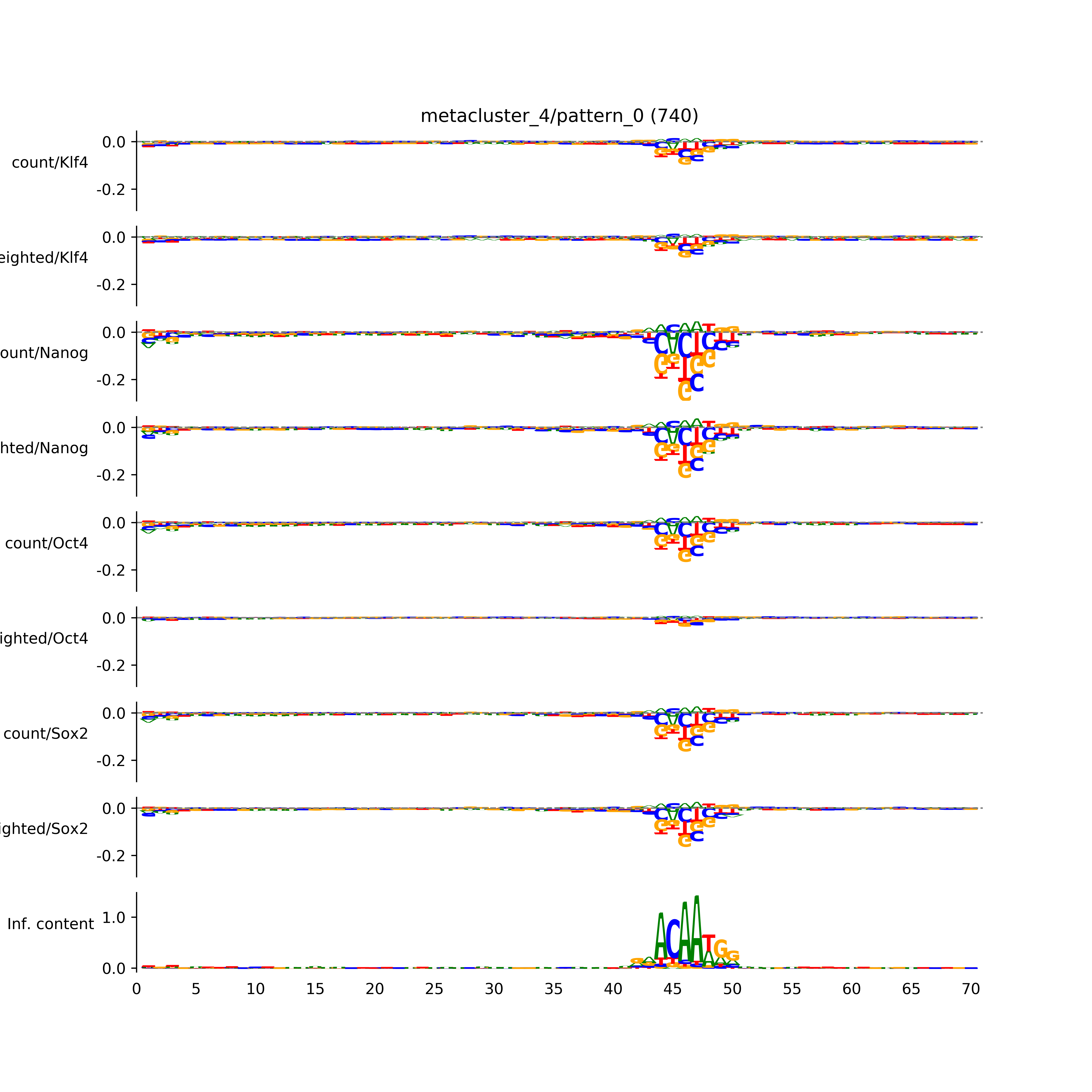
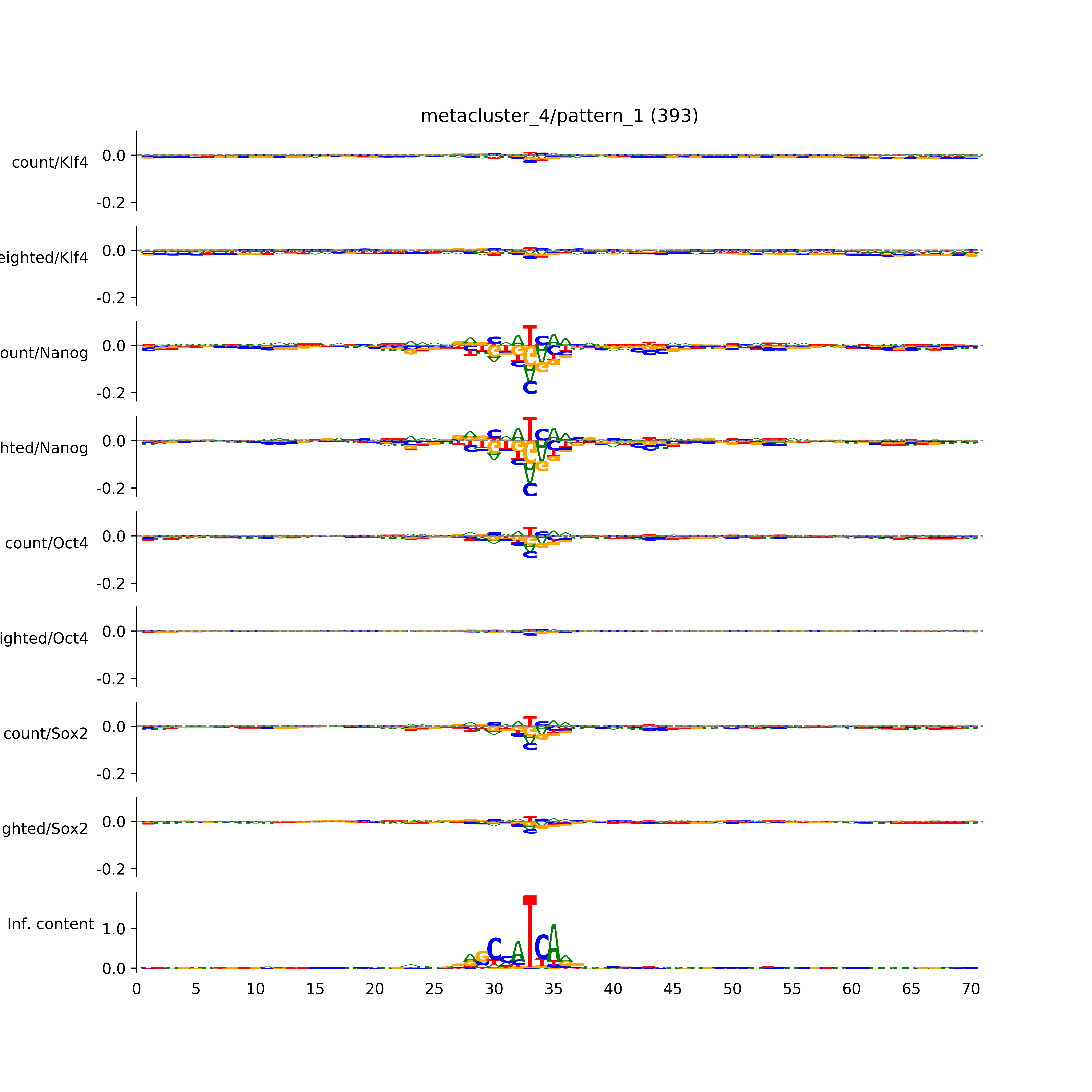
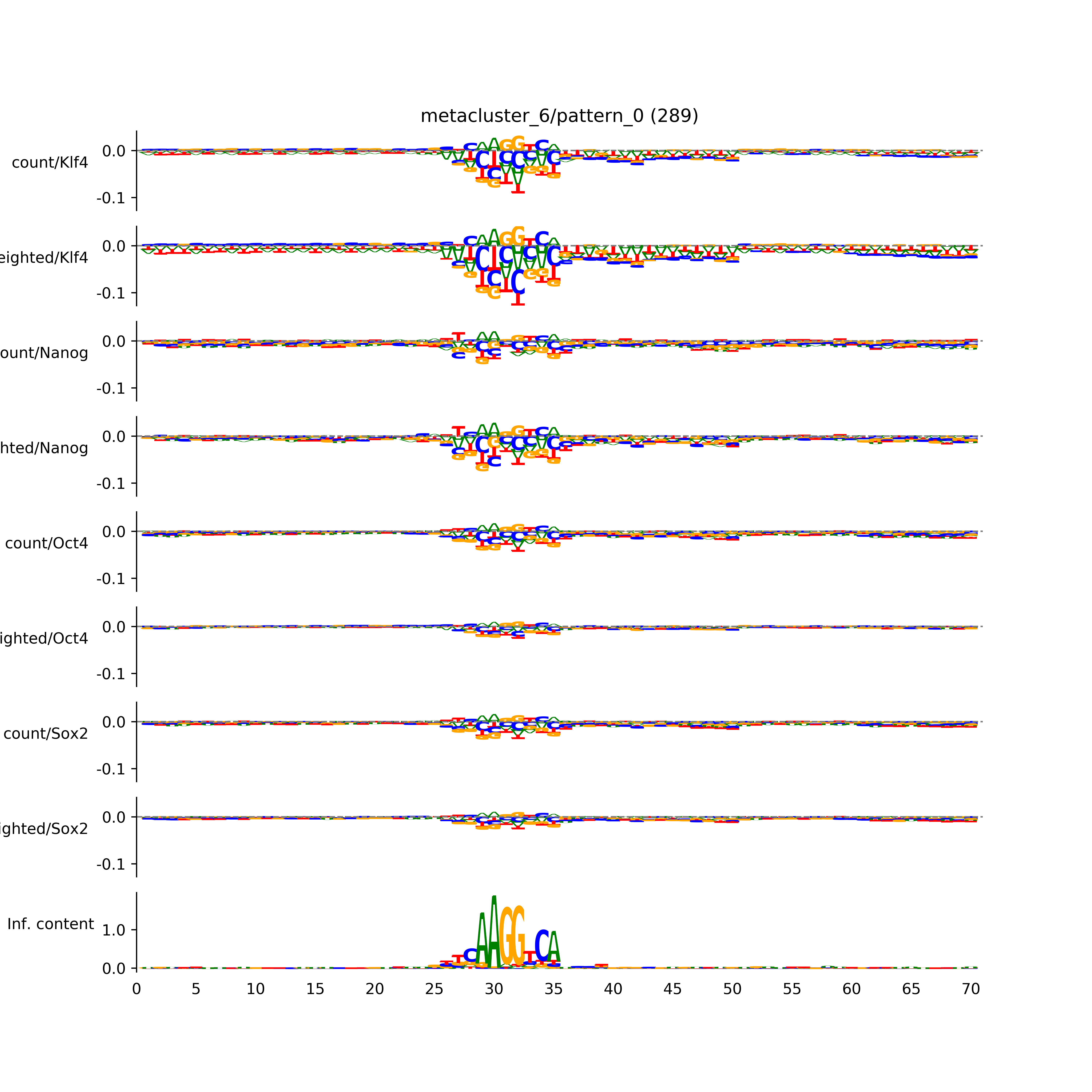
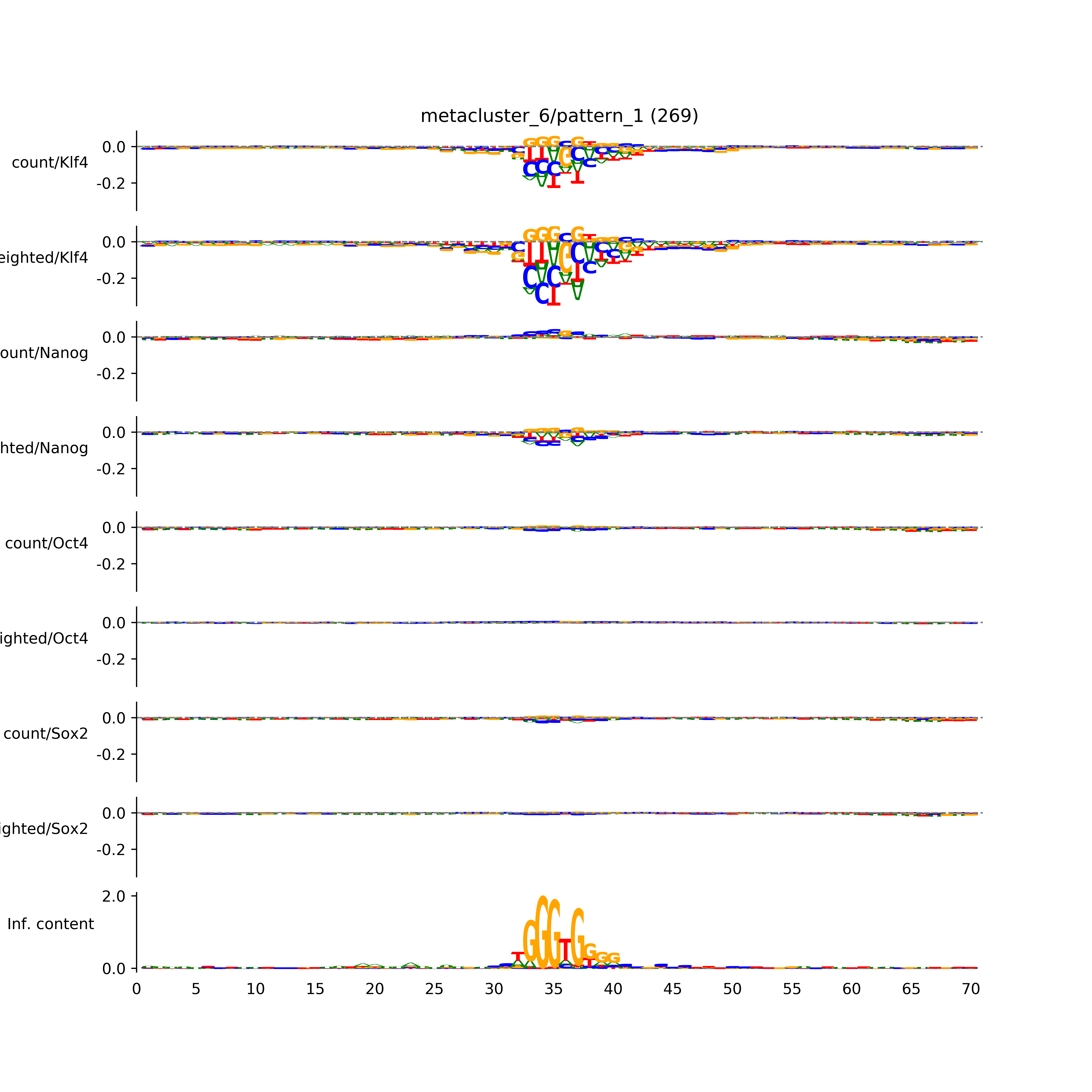
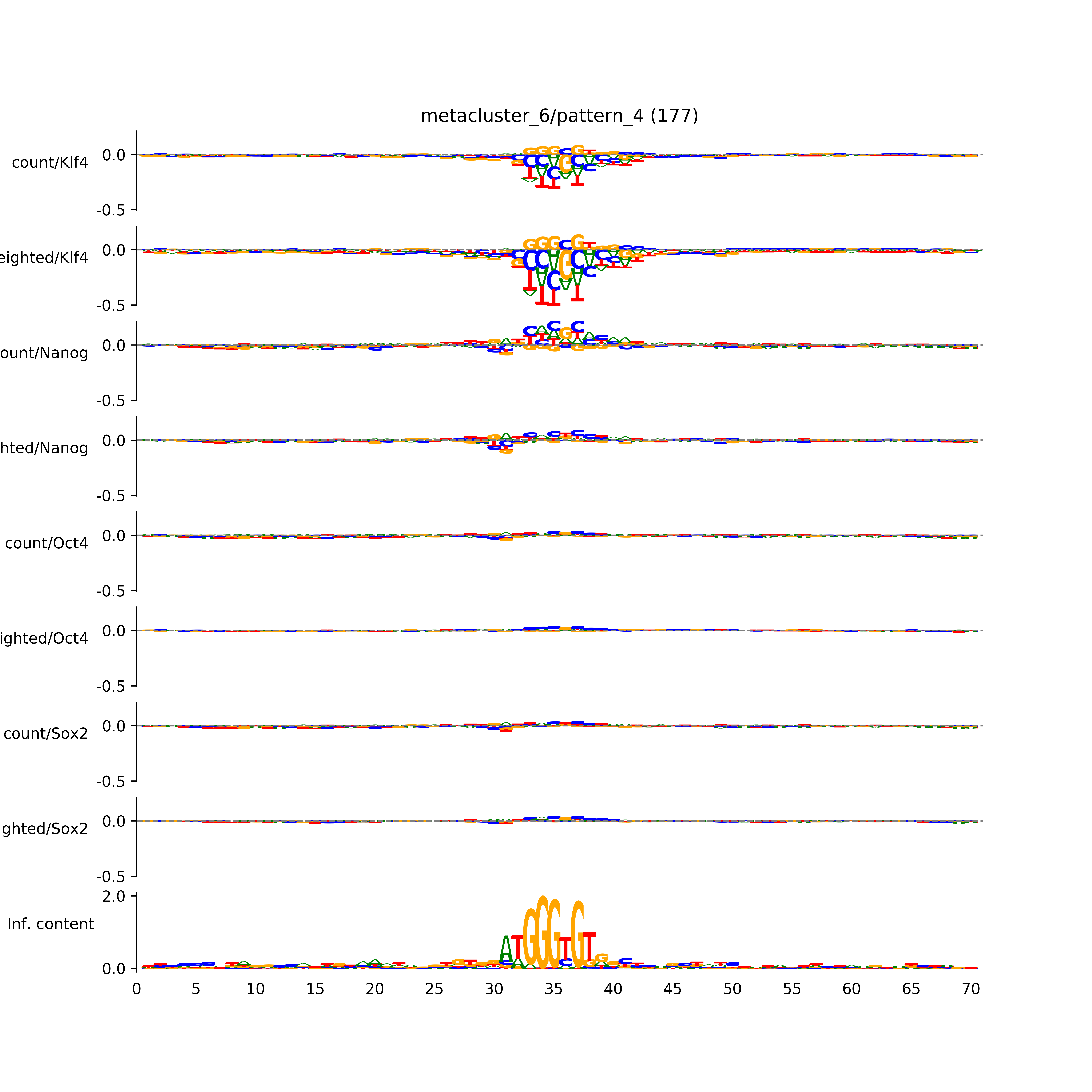
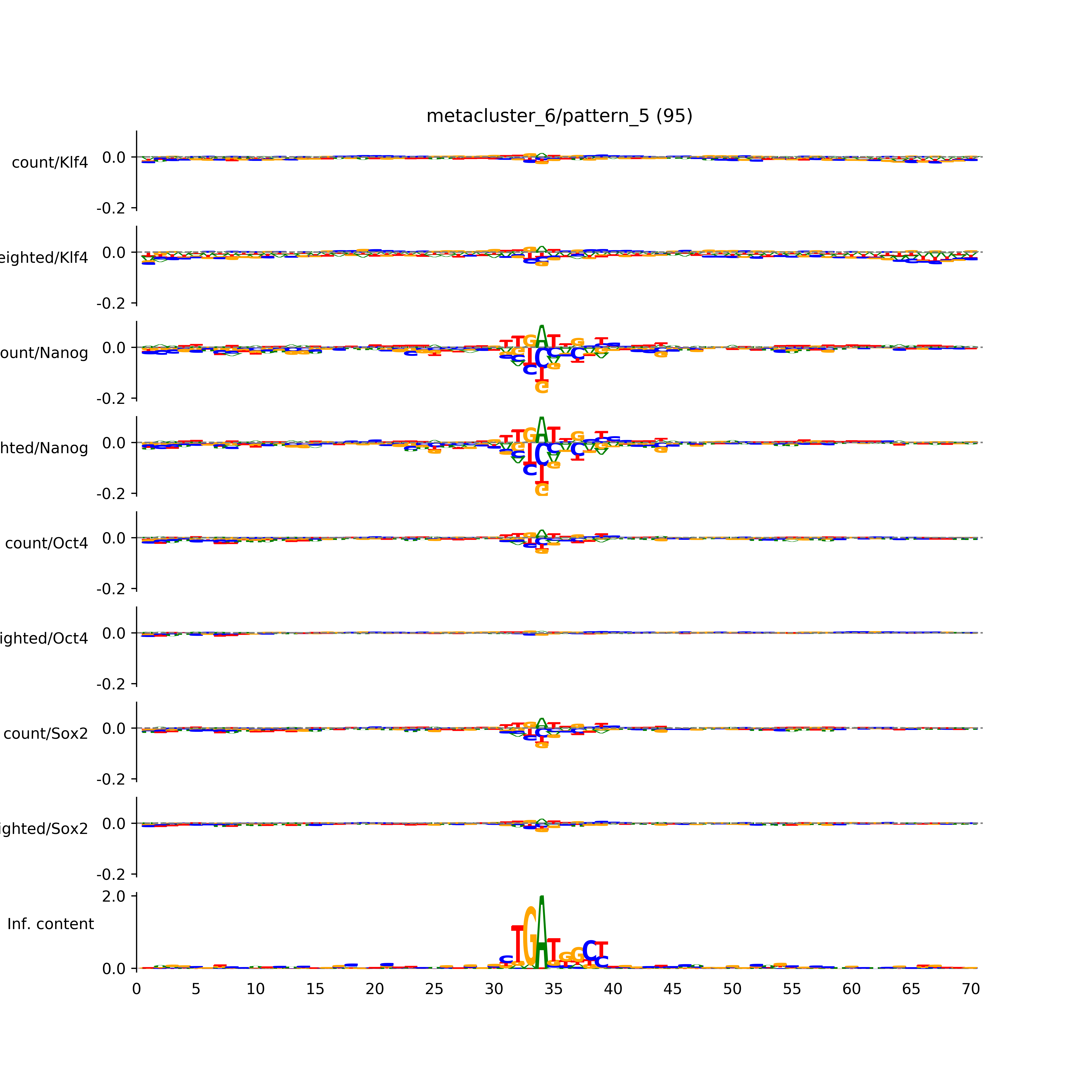
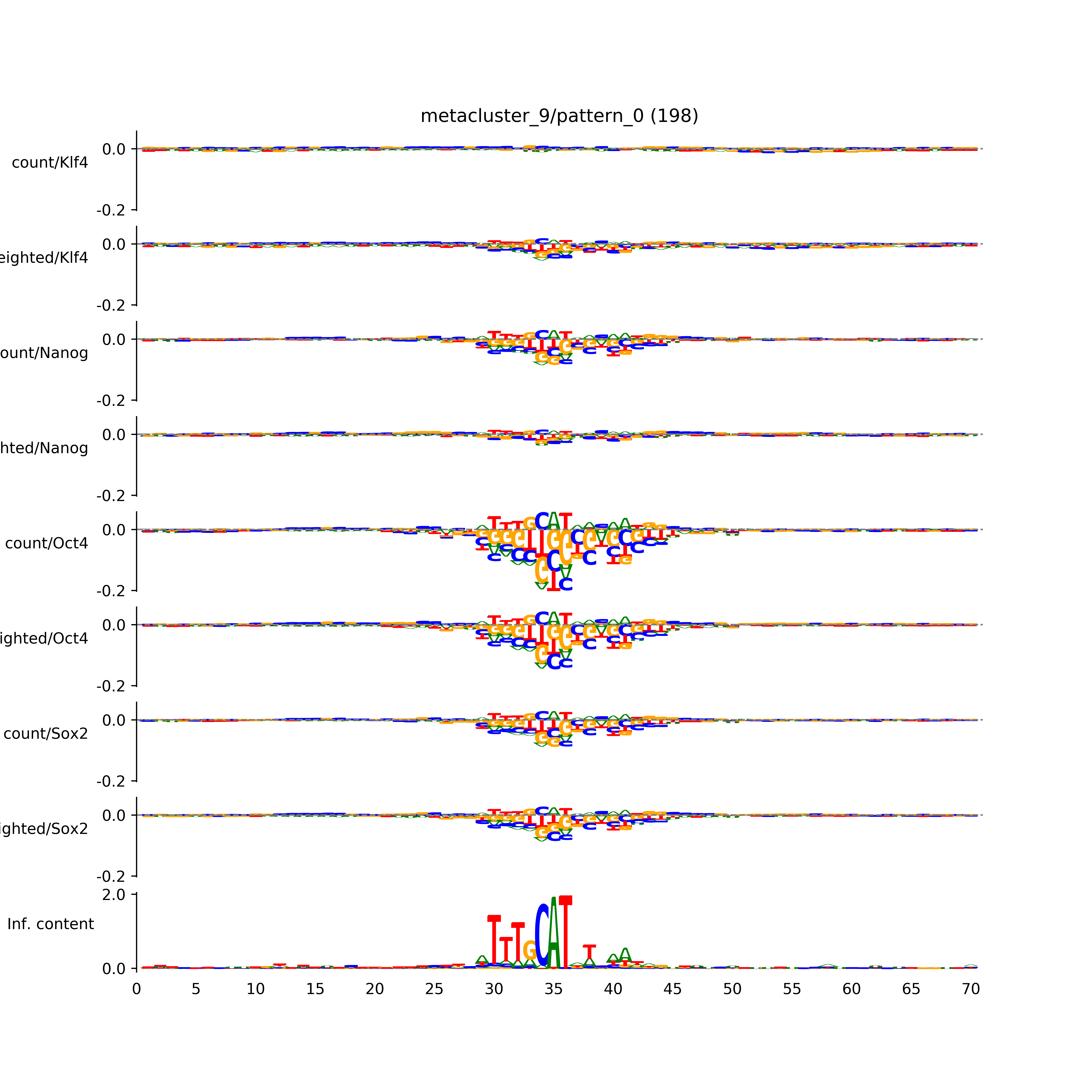
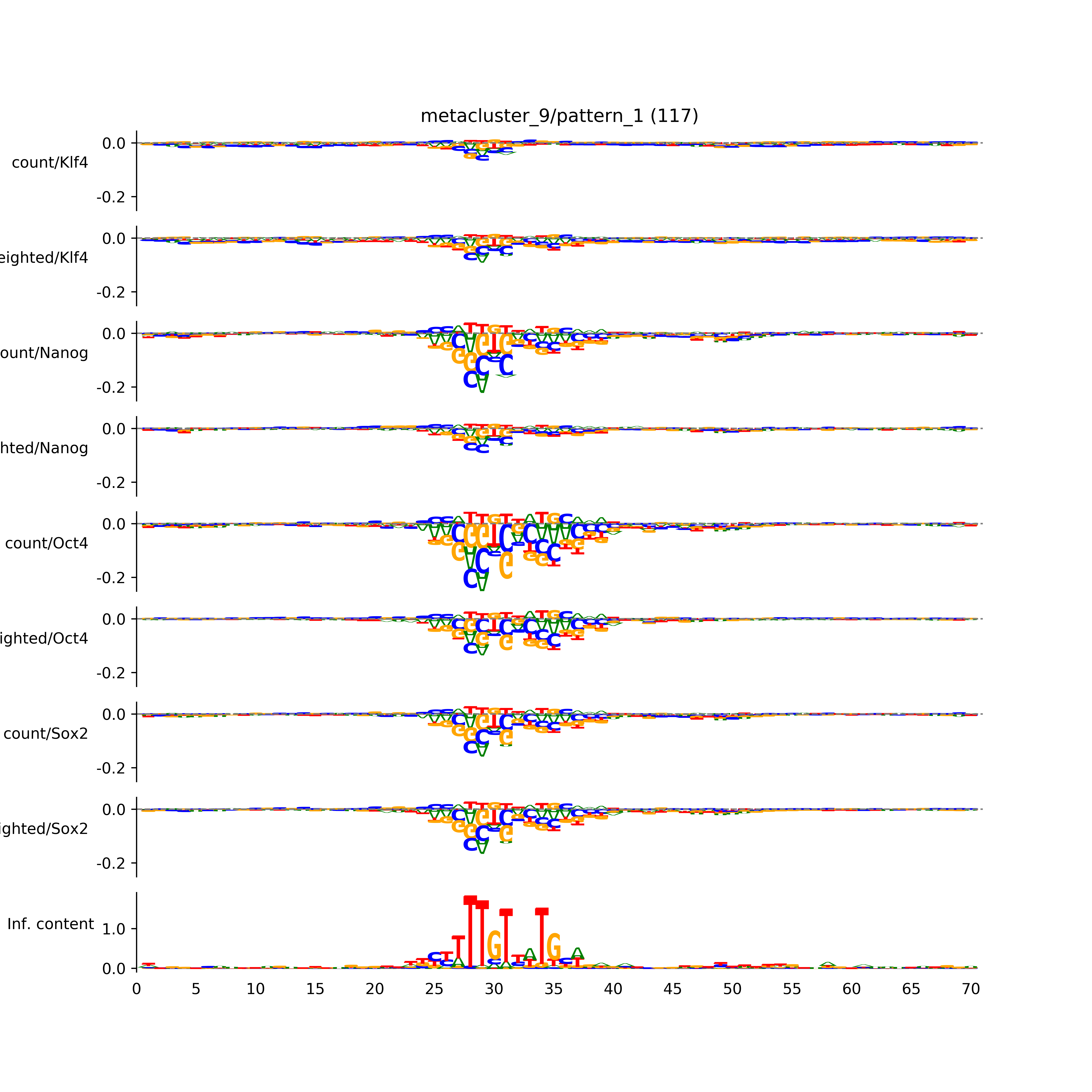
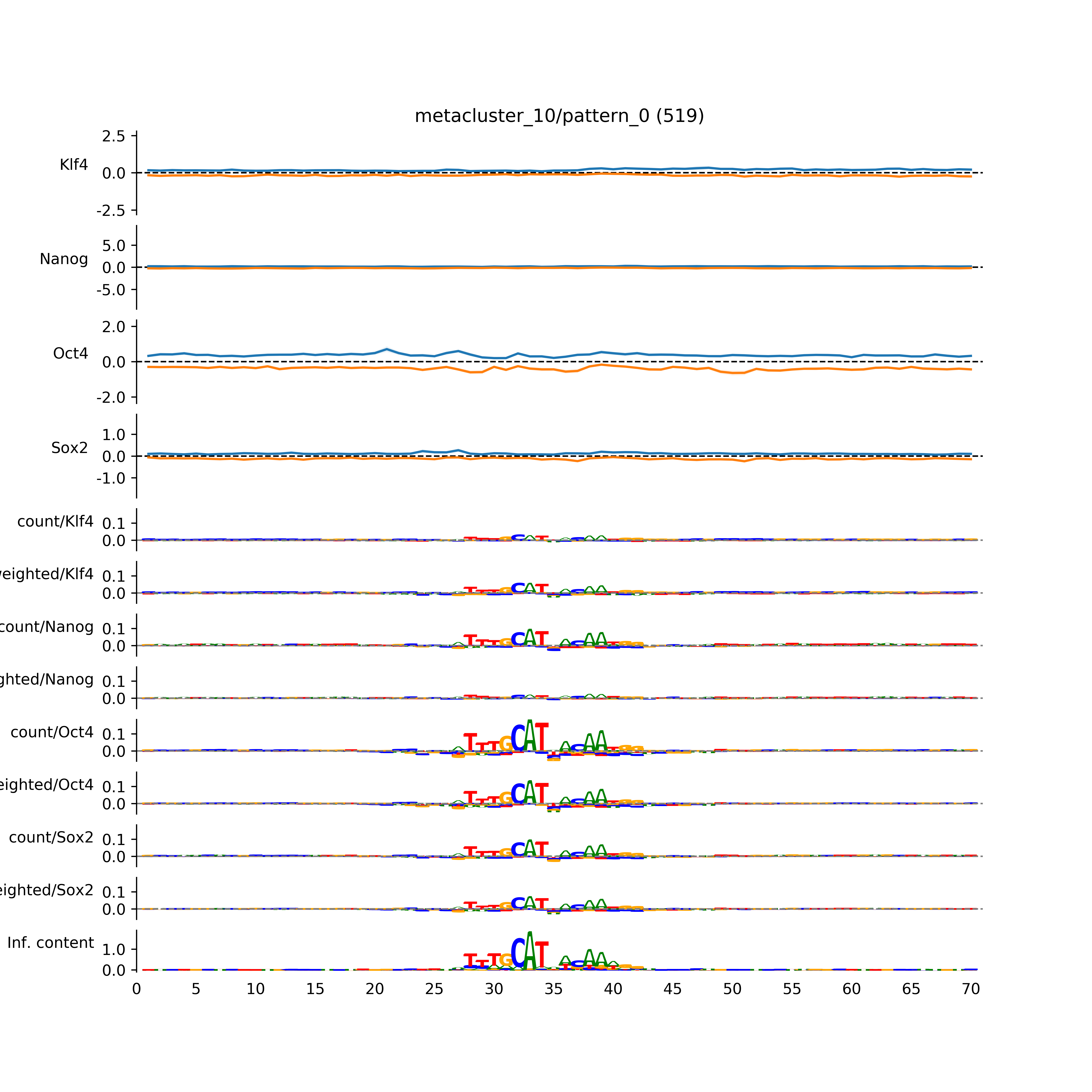
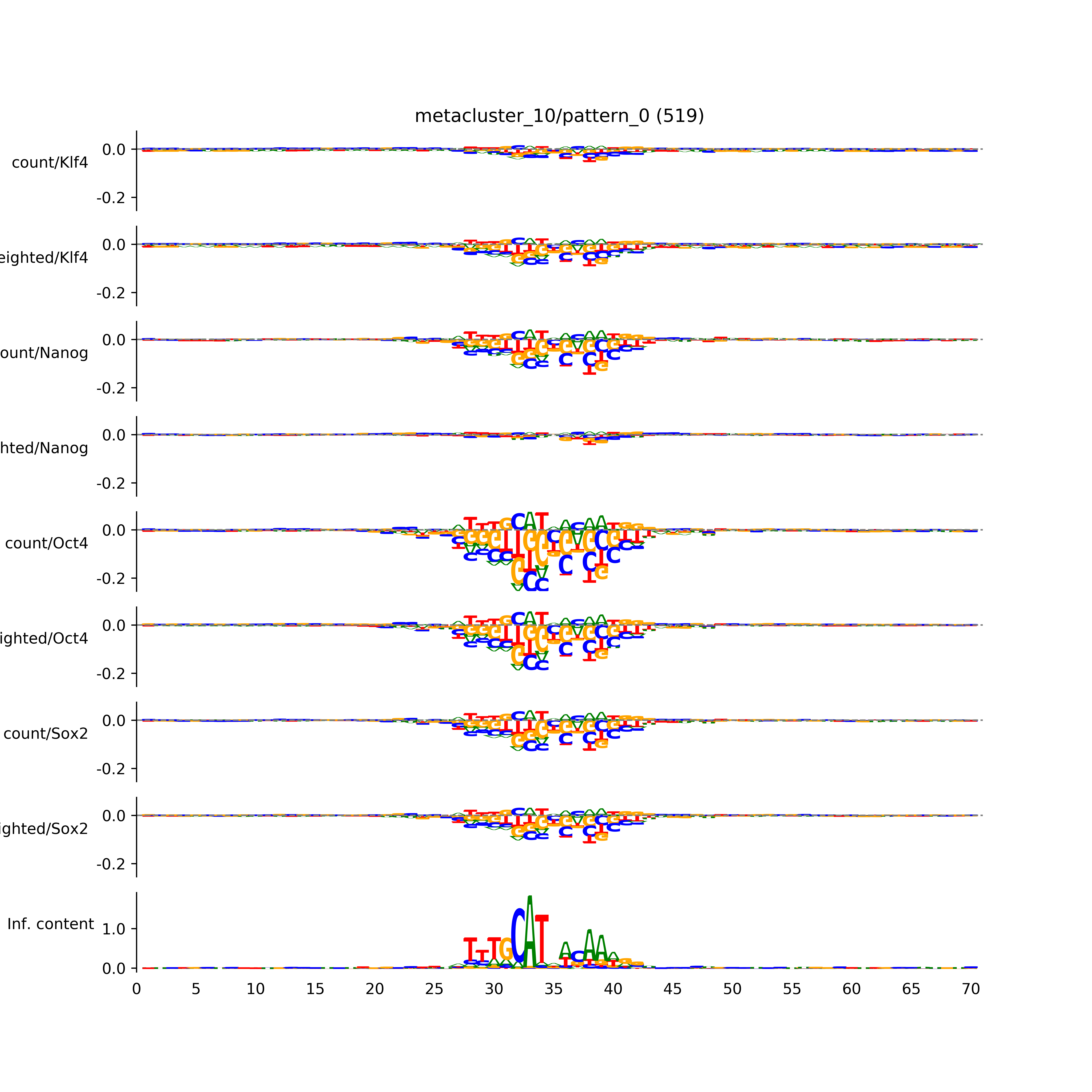
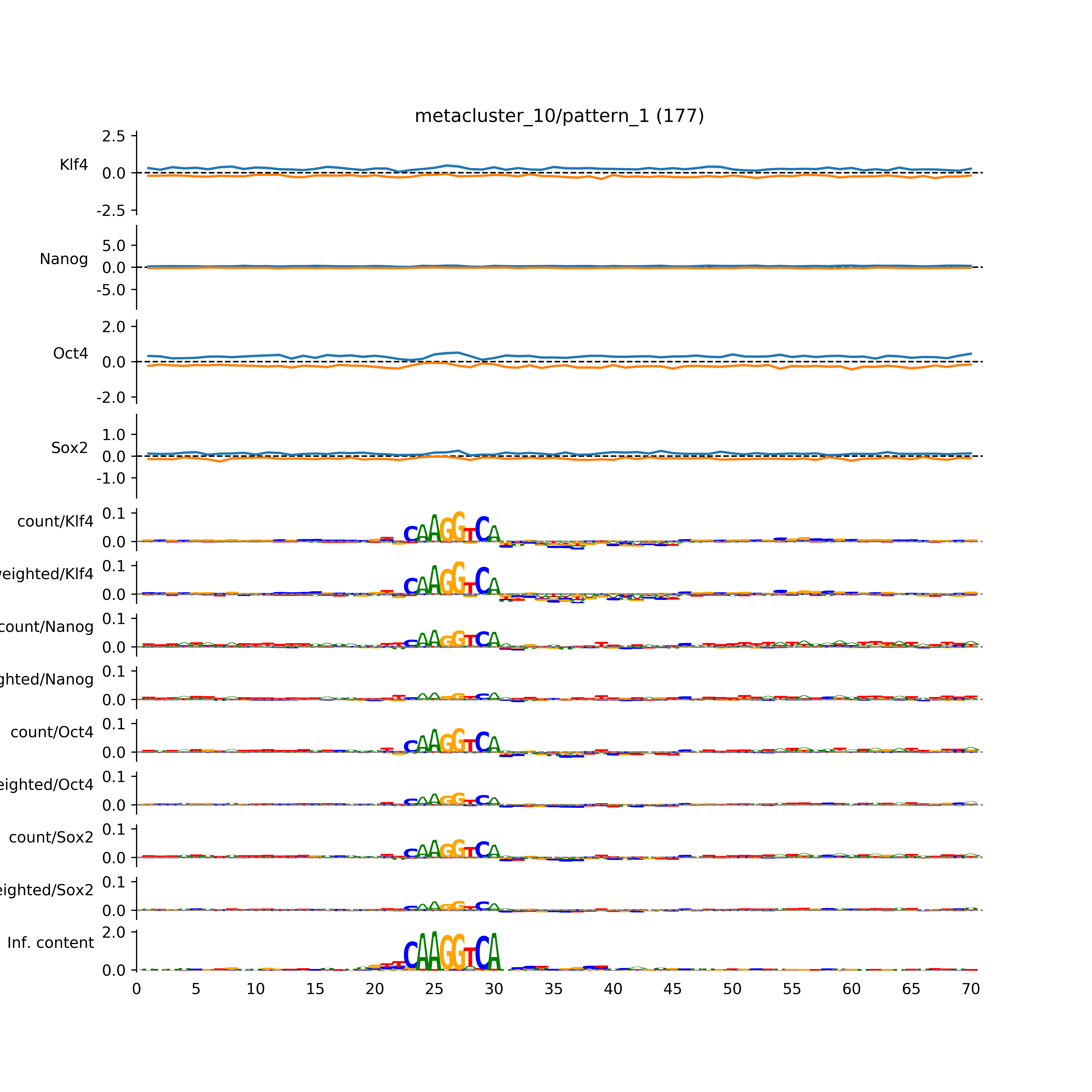
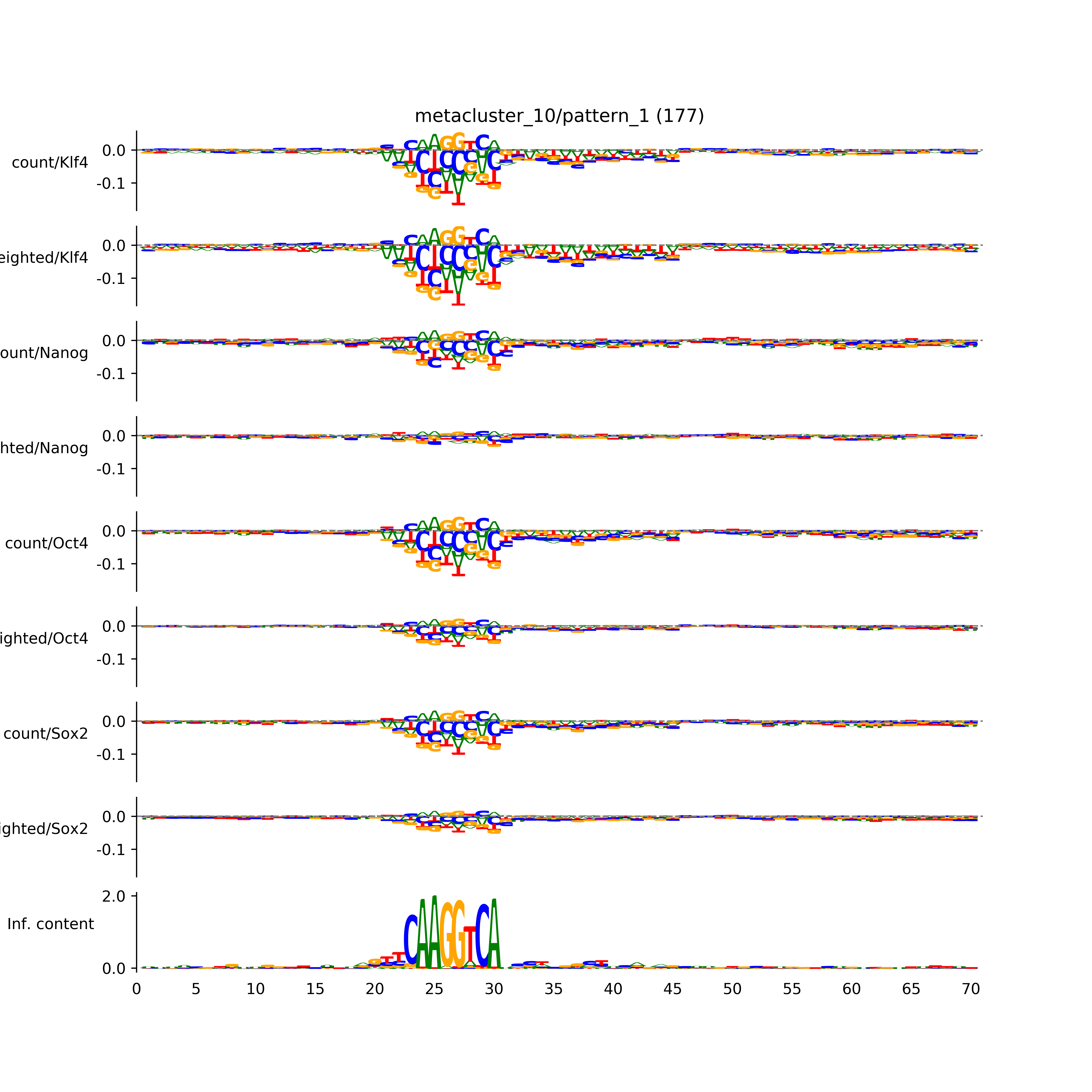
vdom_modisco(mr, url_dir, total_counts, dfp, is_open=True, trim_frac=0.08, letter_width=0.15, height=0.5)
Out[15]:
In [16]:
print("Metaclusters heatmap")
import seaborn as sns
activity_patterns = np.array(mr.f.f['metaclustering_results']['attribute_vectors'])[
np.array(
[x[0] for x in sorted(
enumerate(mr.f.f['metaclustering_results']['metacluster_indices']),
key=lambda x: x[1])])]
sns.heatmap(activity_patterns, center=0);