BPNet tf-Modisco report¶
modisco_dir = "/users/avsec/workspace/basepair/data/processed/chipnexus/exp/models/oct-sox-nanog-klf/models/n_dil_layers=9/modisco/valid/new-hparams"
# Parameters
modisco_dir = "."
from basepair.modisco.results import ModiscoResult
from basepair.config import get_data_dir
from basepair.utils import read_json
from basepair.plot.vdom import vdom_modisco
from kipoi.readers import HDF5Reader
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
from plotnine import *
Using TensorFlow backend.
mr = ModiscoResult(f"{modisco_dir}/modisco.h5")
mr.open()
# load the data
modisco_kwargs = read_json(os.path.join(modisco_dir, "kwargs.json"))
d = HDF5Reader(modisco_kwargs['imp_scores'])
d.open()
strand_dist_file = f"{modisco_dir}/strand_distances.h5"
if modisco_kwargs.get("ignore_strand_dist", False) and os.path.exists(strand_dist_file):
included_samples = HDF5Reader.load(strand_dist_file)['included_samples']
else:
included_samples = np.ones(d.f['inputs'].shape[:1], dtype=bool)
if modisco_kwargs.get("filter_npy", None) is not None:
included_samples = np.load(modisco_kwargs['filter_npy']) * included_samples
id_hash = pd.DataFrame({"peak_id": d.f['/metadata/interval_from_task'][:][included_samples],
"example_idx": np.arange(d.f['/metadata/interval_from_task'][included_samples].shape[0])})
tasks = list(d.f["targets"]["profile"].keys())
# get all seqlet instances
dfp = mr.seqlet_df_instances().rename(columns=dict(seqname="example_idx"))
dfp = pd.merge(dfp, id_hash, on="example_idx")
TF-MoDISco is using the TensorFlow backend.
# row = example_idx
total_counts = pd.DataFrame({task: d.f[f"/targets/profile/{task}"][:][included_samples].sum(axis=-1).sum(axis=-1)
for task in tasks
})
len(mr.patterns())
176
# total number of seqlets
len(dfp)
97031
# Number of metaclusters
len(mr.metaclusters())
15
Number of seqlets per pattern¶
mc_stat = mr.metacluster_stats()
ggplot(aes(x="pattern", y='n'), mc_stat) + geom_bar(stat='identity') + \
facet_wrap("~metacluster", ncol=4, labeller='label_both') + \
ylab("Number of seqlets") + theme_classic()
<ggplot: (-9223363286739682125)>
Zoom-into the 500 seqlet range¶
ggplot(aes(x="pattern", y='n'), mc_stat) + geom_bar(stat='identity') + \
facet_wrap("~metacluster", ncol=4, labeller='label_both') + \
ylab("Number of seqlets") + theme_classic() + coord_cartesian(ylim=[0, 500])
<ggplot: (8750115236749)>
Important tasks per metacluster¶
mcs_grouped = mc_stat.groupby("metacluster").n.agg(["count", "sum"]).reset_index()
fig, ax = plt.subplots(2, 1, sharex=False, figsize=(18,6),
gridspec_kw={'height_ratios': [2,1]})
mcs_grouped.plot("metacluster", "count",
label="# patterns per metacluster", style="o--",
ax=ax[0],
yticks=range(mcs_grouped['count'].max()+1),
xticks=range(38),
fontsize='large',
xlim=(-.5, len(mr.metaclusters()) - .5 ))
mcs_grouped.plot("metacluster", "sum",
label="# seqlets per metacluster",
style="o--", ax=ax[0], secondary_y=True)
ax[0].grid(linewidth=0.2)
mr.plot_metacluster_activity(ax[1], cbar=False)
ax[1].set_title("Importance score activity: Red = positive, Blue = negative");
vdom_modisco(mr, "plots", total_counts, dfp, is_open=True, trim_frac=0.08, letter_width=0.15, height=0.5)
metacluster_0, # patterns: 27, # seqlets: 23880, important for: Klf4
pattern_0: # seqlets: 16233
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 15180 / 133644 regions (11.4%)
pattern_1: # seqlets: 1220
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1216 / 133644 regions (0.9%)
pattern_2: # seqlets: 1045
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1034 / 133644 regions (0.8%)
pattern_3: # seqlets: 568
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 567 / 133644 regions (0.4%)
pattern_4: # seqlets: 535
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 520 / 133644 regions (0.4%)
pattern_5: # seqlets: 484
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 477 / 133644 regions (0.4%)
pattern_6: # seqlets: 450
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 431 / 133644 regions (0.3%)
pattern_7: # seqlets: 354
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 347 / 133644 regions (0.3%)
pattern_8: # seqlets: 324
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 322 / 133644 regions (0.2%)
pattern_9: # seqlets: 317
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 317 / 133644 regions (0.2%)
pattern_10: # seqlets: 251
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 250 / 133644 regions (0.2%)
pattern_11: # seqlets: 201
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 192 / 133644 regions (0.1%)
pattern_12: # seqlets: 195
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 195 / 133644 regions (0.1%)
pattern_13: # seqlets: 177
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 177 / 133644 regions (0.1%)
pattern_14: # seqlets: 174
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 167 / 133644 regions (0.1%)
pattern_15: # seqlets: 161
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 160 / 133644 regions (0.1%)
pattern_16: # seqlets: 169
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 169 / 133644 regions (0.1%)
pattern_17: # seqlets: 152
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 145 / 133644 regions (0.1%)
pattern_18: # seqlets: 140
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 136 / 133644 regions (0.1%)
pattern_19: # seqlets: 133
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 131 / 133644 regions (0.1%)
pattern_20: # seqlets: 106
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 106 / 133644 regions (0.1%)
pattern_21: # seqlets: 104
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence
ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 101 / 133644 regions (0.1%)
pattern_22: # seqlets: 92
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 86 / 133644 regions (0.1%)
pattern_23: # seqlets: 82
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 81 / 133644 regions (0.1%)
pattern_24: # seqlets: 74
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 74 / 133644 regions (0.1%)
pattern_25: # seqlets: 69
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 69 / 133644 regions (0.1%)
pattern_26: # seqlets: 70
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 70 / 133644 regions (0.1%)
metacluster_1, # patterns: 26, # seqlets: 17559, important for: Nanog,Oct4,Sox2
pattern_0: # seqlets: 3933
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 3856 / 133644 regions (2.9%)
pattern_1: # seqlets: 3023
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 2956 / 133644 regions (2.2%)
pattern_2: # seqlets: 2232
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 2201 / 133644 regions (1.6%)
pattern_3: # seqlets: 1217
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1206 / 133644 regions (0.9%)
pattern_4: # seqlets: 901
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 898 / 133644 regions (0.7%)
pattern_5: # seqlets: 857
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 843 / 133644 regions (0.6%)
pattern_6: # seqlets: 866
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 860 / 133644 regions (0.6%)
pattern_7: # seqlets: 828
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 828 / 133644 regions (0.6%)
pattern_8: # seqlets: 460
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 460 / 133644 regions (0.3%)
pattern_9: # seqlets: 388
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 387 / 133644 regions (0.3%)
pattern_10: # seqlets: 340
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 322 / 133644 regions (0.2%)
pattern_11: # seqlets: 287
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 287 / 133644 regions (0.2%)
pattern_12: # seqlets: 357
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 357 / 133644 regions (0.3%)
pattern_13: # seqlets: 196
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 196 / 133644 regions (0.1%)
pattern_14: # seqlets: 203
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 203 / 133644 regions (0.2%)
pattern_15: # seqlets: 137
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)
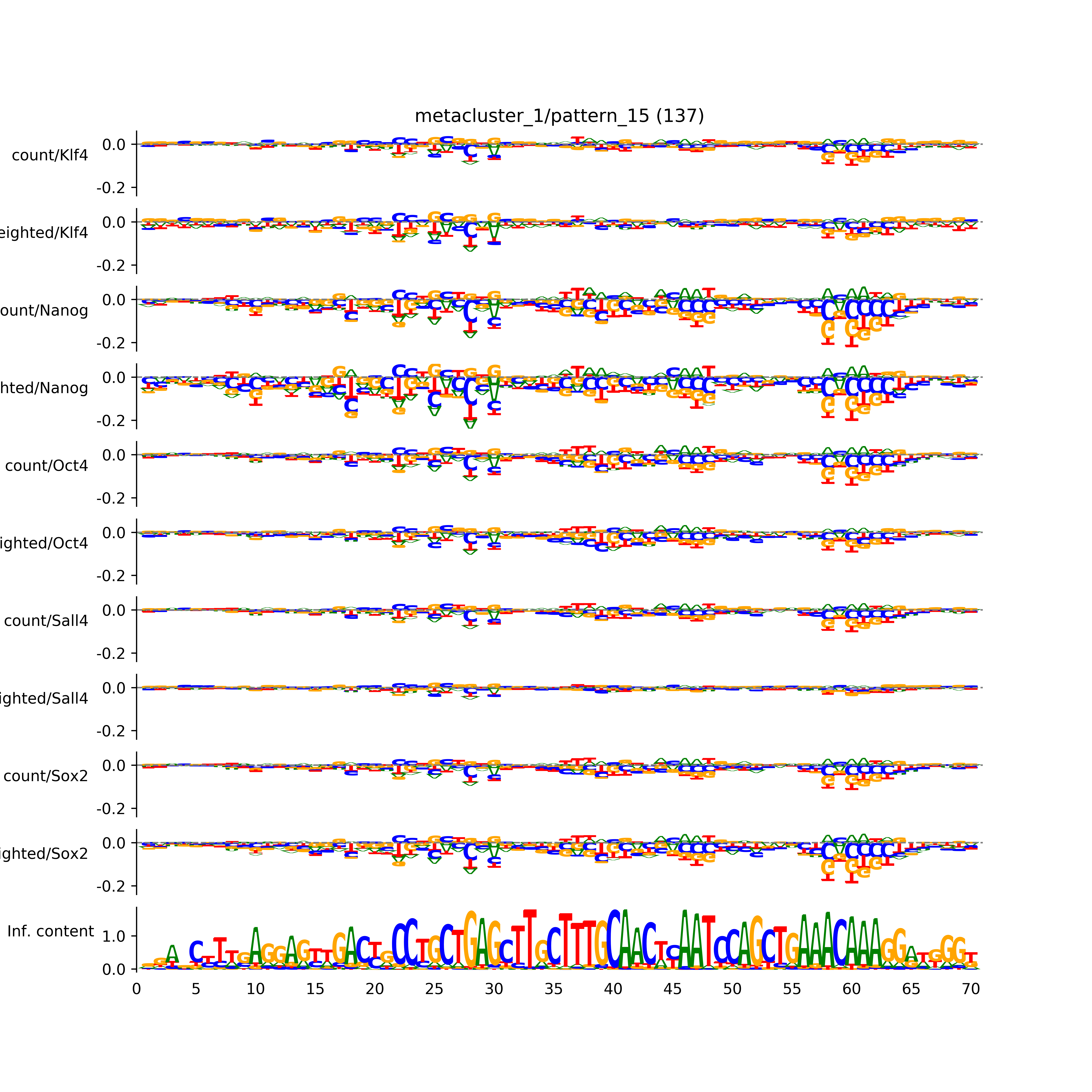
Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 137 / 133644 regions (0.1%)
pattern_16: # seqlets: 148
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 148 / 133644 regions (0.1%)
pattern_17: # seqlets: 136
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 136 / 133644 regions (0.1%)
pattern_18: # seqlets: 186
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)
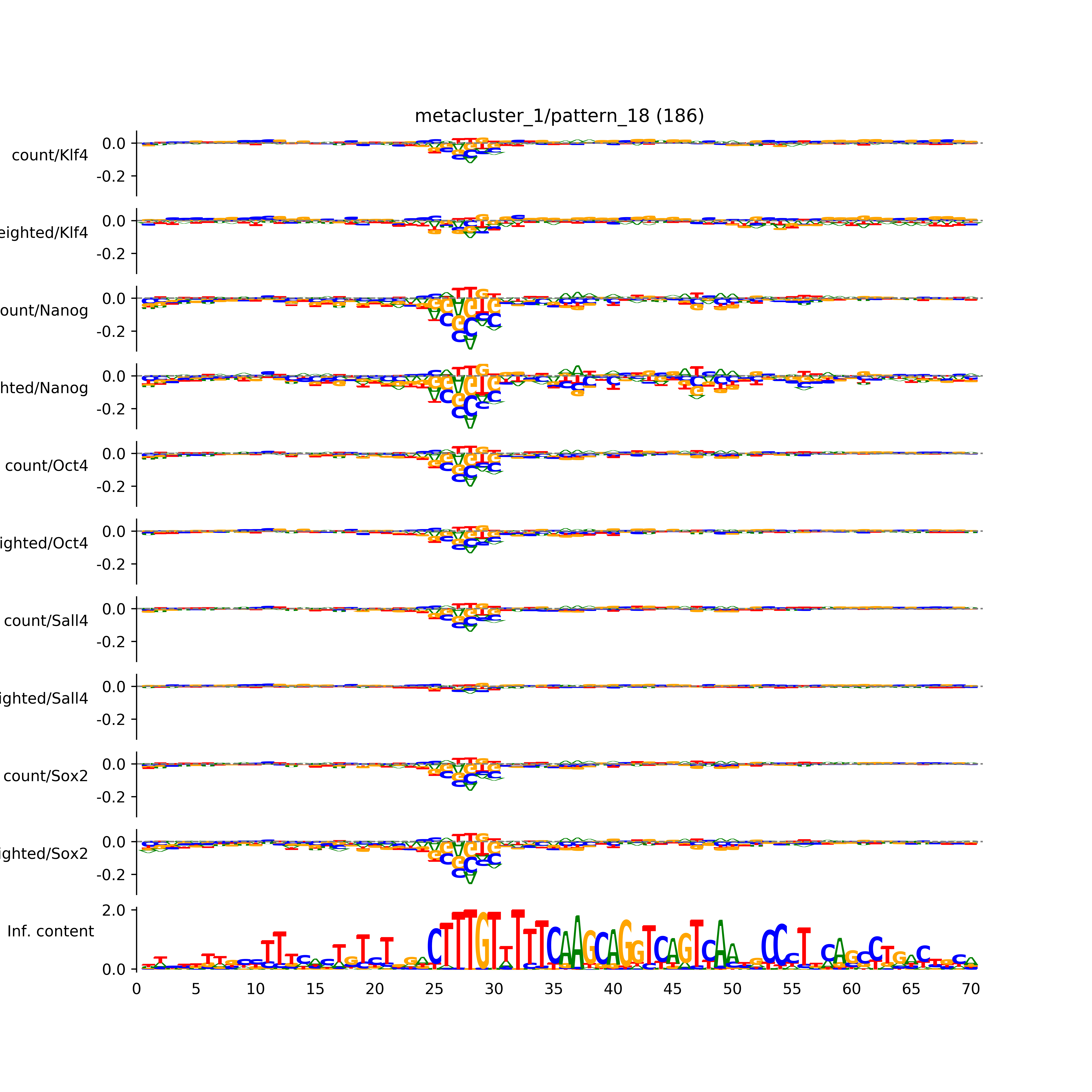
Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 186 / 133644 regions (0.1%)
pattern_19: # seqlets: 161
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 161 / 133644 regions (0.1%)
pattern_20: # seqlets: 105
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 105 / 133644 regions (0.1%)
pattern_21: # seqlets: 140
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 140 / 133644 regions (0.1%)
pattern_22: # seqlets: 104
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 104 / 133644 regions (0.1%)
pattern_23: # seqlets: 126
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 126 / 133644 regions (0.1%)
pattern_24: # seqlets: 126
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 126 / 133644 regions (0.1%)
pattern_25: # seqlets: 102
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 102 / 133644 regions (0.1%)
metacluster_2, # patterns: 22, # seqlets: 11042, important for: Nanog,Sox2
pattern_0: # seqlets: 3631
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 3539 / 133644 regions (2.6%)
pattern_1: # seqlets: 2179
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 2151 / 133644 regions (1.6%)
pattern_2: # seqlets: 641
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 629 / 133644 regions (0.5%)
pattern_3: # seqlets: 707
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 707 / 133644 regions (0.5%)
pattern_4: # seqlets: 614
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 614 / 133644 regions (0.5%)
pattern_5: # seqlets: 424
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 418 / 133644 regions (0.3%)
pattern_6: # seqlets: 385
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 383 / 133644 regions (0.3%)
pattern_7: # seqlets: 418
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 415 / 133644 regions (0.3%)
pattern_8: # seqlets: 425
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 425 / 133644 regions (0.3%)
pattern_9: # seqlets: 297
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 297 / 133644 regions (0.2%)
pattern_10: # seqlets: 122
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 122 / 133644 regions (0.1%)
pattern_11: # seqlets: 152
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 152 / 133644 regions (0.1%)
pattern_12: # seqlets: 114
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 114 / 133644 regions (0.1%)
pattern_13: # seqlets: 124
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 123 / 133644 regions (0.1%)
pattern_14: # seqlets: 117
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 117 / 133644 regions (0.1%)
pattern_15: # seqlets: 121
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 121 / 133644 regions (0.1%)
pattern_16: # seqlets: 140
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 140 / 133644 regions (0.1%)
pattern_17: # seqlets: 95
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 95 / 133644 regions (0.1%)
pattern_18: # seqlets: 90
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 90 / 133644 regions (0.1%)
pattern_19: # seqlets: 92
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 92 / 133644 regions (0.1%)
pattern_20: # seqlets: 80
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 79 / 133644 regions (0.1%)
pattern_21: # seqlets: 74
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 74 / 133644 regions (0.1%)
metacluster_3, # patterns: 32, # seqlets: 20129, important for: Klf4,Nanog,Oct4,Sall4,Sox2
pattern_0: # seqlets: 8325
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 8162 / 133644 regions (6.1%)
pattern_1: # seqlets: 1722
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1695 / 133644 regions (1.3%)
pattern_2: # seqlets: 1185
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1180 / 133644 regions (0.9%)
pattern_3: # seqlets: 1032
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1020 / 133644 regions (0.8%)
pattern_4: # seqlets: 773
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 711 / 133644 regions (0.5%)
pattern_5: # seqlets: 801
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 796 / 133644 regions (0.6%)
pattern_6: # seqlets: 586
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 585 / 133644 regions (0.4%)
pattern_7: # seqlets: 561
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 556 / 133644 regions (0.4%)
pattern_8: # seqlets: 518
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 507 / 133644 regions (0.4%)
pattern_9: # seqlets: 475
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 471 / 133644 regions (0.4%)
pattern_10: # seqlets: 399
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 399 / 133644 regions (0.3%)
pattern_11: # seqlets: 441
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 426 / 133644 regions (0.3%)
pattern_12: # seqlets: 383
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 379 / 133644 regions (0.3%)
pattern_13: # seqlets: 323
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 323 / 133644 regions (0.2%)
pattern_14: # seqlets: 229
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 229 / 133644 regions (0.2%)
pattern_15: # seqlets: 228
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 228 / 133644 regions (0.2%)
pattern_16: # seqlets: 208
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 208 / 133644 regions (0.2%)
pattern_17: # seqlets: 186
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 184 / 133644 regions (0.1%)
pattern_18: # seqlets: 183
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 183 / 133644 regions (0.1%)
pattern_19: # seqlets: 162
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 162 / 133644 regions (0.1%)
pattern_20: # seqlets: 145
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 145 / 133644 regions (0.1%)
pattern_21: # seqlets: 148
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 148 / 133644 regions (0.1%)
pattern_22: # seqlets: 125
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 125 / 133644 regions (0.1%)
pattern_23: # seqlets: 114
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 114 / 133644 regions (0.1%)
pattern_24: # seqlets: 110
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 110 / 133644 regions (0.1%)
pattern_25: # seqlets: 110
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 110 / 133644 regions (0.1%)
pattern_26: # seqlets: 120
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 120 / 133644 regions (0.1%)
pattern_27: # seqlets: 104
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 101 / 133644 regions (0.1%)
pattern_28: # seqlets: 100
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 100 / 133644 regions (0.1%)
pattern_29: # seqlets: 99
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 99 / 133644 regions (0.1%)
pattern_30: # seqlets: 110
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 110 / 133644 regions (0.1%)
pattern_31: # seqlets: 124
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 124 / 133644 regions (0.1%)
metacluster_4, # patterns: 8, # seqlets: 4400, important for: Oct4,Sox2
pattern_0: # seqlets: 2427
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 2390 / 133644 regions (1.8%)
pattern_1: # seqlets: 824
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 814 / 133644 regions (0.6%)
pattern_2: # seqlets: 427
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 421 / 133644 regions (0.3%)
pattern_3: # seqlets: 253
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 250 / 133644 regions (0.2%)
pattern_4: # seqlets: 244
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 244 / 133644 regions (0.2%)
pattern_5: # seqlets: 96
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 96 / 133644 regions (0.1%)
pattern_6: # seqlets: 69
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 69 / 133644 regions (0.1%)
pattern_7: # seqlets: 60
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 60 / 133644 regions (0.0%)
metacluster_5, # patterns: 15, # seqlets: 5152, important for: Nanog,Oct4,Sall4,Sox2
pattern_0: # seqlets: 2299
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 2275 / 133644 regions (1.7%)
pattern_1: # seqlets: 807
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 807 / 133644 regions (0.6%)
pattern_2: # seqlets: 605
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 605 / 133644 regions (0.5%)
pattern_3: # seqlets: 221
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 221 / 133644 regions (0.2%)
pattern_4: # seqlets: 265
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 265 / 133644 regions (0.2%)
pattern_5: # seqlets: 120
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 120 / 133644 regions (0.1%)
pattern_6: # seqlets: 127
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 127 / 133644 regions (0.1%)
pattern_7: # seqlets: 120
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 120 / 133644 regions (0.1%)
pattern_8: # seqlets: 89
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 89 / 133644 regions (0.1%)
pattern_9: # seqlets: 84
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 81 / 133644 regions (0.1%)
pattern_10: # seqlets: 83
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 83 / 133644 regions (0.1%)
pattern_11: # seqlets: 108
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 108 / 133644 regions (0.1%)
pattern_12: # seqlets: 84
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 84 / 133644 regions (0.1%)
pattern_13: # seqlets: 65
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 65 / 133644 regions (0.0%)
pattern_14: # seqlets: 75
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 75 / 133644 regions (0.1%)
metacluster_7, # patterns: 11, # seqlets: 4187, important for: Klf4,Nanog,Oct4,Sox2
pattern_0: # seqlets: 984
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 976 / 133644 regions (0.7%)
pattern_1: # seqlets: 700
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 699 / 133644 regions (0.5%)
pattern_2: # seqlets: 521
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 516 / 133644 regions (0.4%)
pattern_3: # seqlets: 539
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 534 / 133644 regions (0.4%)
pattern_4: # seqlets: 398
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 395 / 133644 regions (0.3%)
pattern_5: # seqlets: 376
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 375 / 133644 regions (0.3%)
pattern_6: # seqlets: 177
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 177 / 133644 regions (0.1%)
pattern_7: # seqlets: 101
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 101 / 133644 regions (0.1%)
pattern_8: # seqlets: 171
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 171 / 133644 regions (0.1%)
pattern_9: # seqlets: 134
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 134 / 133644 regions (0.1%)
pattern_10: # seqlets: 86
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 83 / 133644 regions (0.1%)
metacluster_8, # patterns: 6, # seqlets: 1865, important for: Nanog
pattern_0: # seqlets: 1070
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1064 / 133644 regions (0.8%)
pattern_1: # seqlets: 262
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 261 / 133644 regions (0.2%)
pattern_2: # seqlets: 158
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 158 / 133644 regions (0.1%)
pattern_3: # seqlets: 185
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 185 / 133644 regions (0.1%)
pattern_4: # seqlets: 100
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 100 / 133644 regions (0.1%)
pattern_5: # seqlets: 90
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 90 / 133644 regions (0.1%)
metacluster_10, # patterns: 7, # seqlets: 2745, important for: Klf4,Sall4
pattern_0: # seqlets: 1305
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1296 / 133644 regions (1.0%)
pattern_1: # seqlets: 734
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 725 / 133644 regions (0.5%)
pattern_2: # seqlets: 276
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 276 / 133644 regions (0.2%)
pattern_3: # seqlets: 139
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 139 / 133644 regions (0.1%)
pattern_4: # seqlets: 115
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 115 / 133644 regions (0.1%)
pattern_5: # seqlets: 92
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 90 / 133644 regions (0.1%)
pattern_6: # seqlets: 84
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 84 / 133644 regions (0.1%)
metacluster_11, # patterns: 1, # seqlets: 1125, important for: Sox2
pattern_0: # seqlets: 1125
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 1121 / 133644 regions (0.8%)
metacluster_12, # patterns: 8, # seqlets: 1282, important for: Klf4,Nanog
pattern_0: # seqlets: 316
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 316 / 133644 regions (0.2%)
pattern_1: # seqlets: 202
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 201 / 133644 regions (0.2%)
pattern_2: # seqlets: 145
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 145 / 133644 regions (0.1%)
pattern_3: # seqlets: 110
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 110 / 133644 regions (0.1%)
pattern_4: # seqlets: 216
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 213 / 133644 regions (0.2%)
pattern_5: # seqlets: 89
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 89 / 133644 regions (0.1%)
pattern_6: # seqlets: 127
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 125 / 133644 regions (0.1%)
pattern_7: # seqlets: 77
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 77 / 133644 regions (0.1%)
metacluster_13, # patterns: 6, # seqlets: 1881, important for: Klf4,Nanog,Sall4
pattern_0: # seqlets: 950
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 939 / 133644 regions (0.7%)
pattern_1: # seqlets: 295
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 295 / 133644 regions (0.2%)
pattern_2: # seqlets: 281
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 281 / 133644 regions (0.2%)
pattern_3: # seqlets: 155
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 154 / 133644 regions (0.1%)
pattern_4: # seqlets: 95
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 95 / 133644 regions (0.1%)
pattern_5: # seqlets: 105
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 105 / 133644 regions (0.1%)
metacluster_14, # patterns: 7, # seqlets: 1784, important for: Klf4,Nanog,Sox2
pattern_0: # seqlets: 206
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 205 / 133644 regions (0.2%)
pattern_1: # seqlets: 388
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 386 / 133644 regions (0.3%)
pattern_2: # seqlets: 566
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 565 / 133644 regions (0.4%)
pattern_3: # seqlets: 173
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 172 / 133644 regions (0.1%)
pattern_4: # seqlets: 256
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 256 / 133644 regions (0.2%)
pattern_5: # seqlets: 93
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 93 / 133644 regions (0.1%)
pattern_6: # seqlets: 102
Aggregated profiles and contribution scores)

Aggregated hypothetical contribution scores)

Sequence

ChIP-nexus counts


Importance scores (profile)

Importance scores (counts)

Positional distribution
Total count distribution
Pattern occurs in 102 / 133644 regions (0.1%)
print("Metaclusters heatmap")
import seaborn as sns
activity_patterns = np.array(mr.f.f['metaclustering_results']['attribute_vectors'])[
np.array(
[x[0] for x in sorted(
enumerate(mr.f.f['metaclustering_results']['metacluster_indices']),
key=lambda x: x[1])])]
sns.heatmap(activity_patterns, center=0);
Metaclusters heatmap