Pairwise perturbation¶
- This notebooks shows
TODO¶
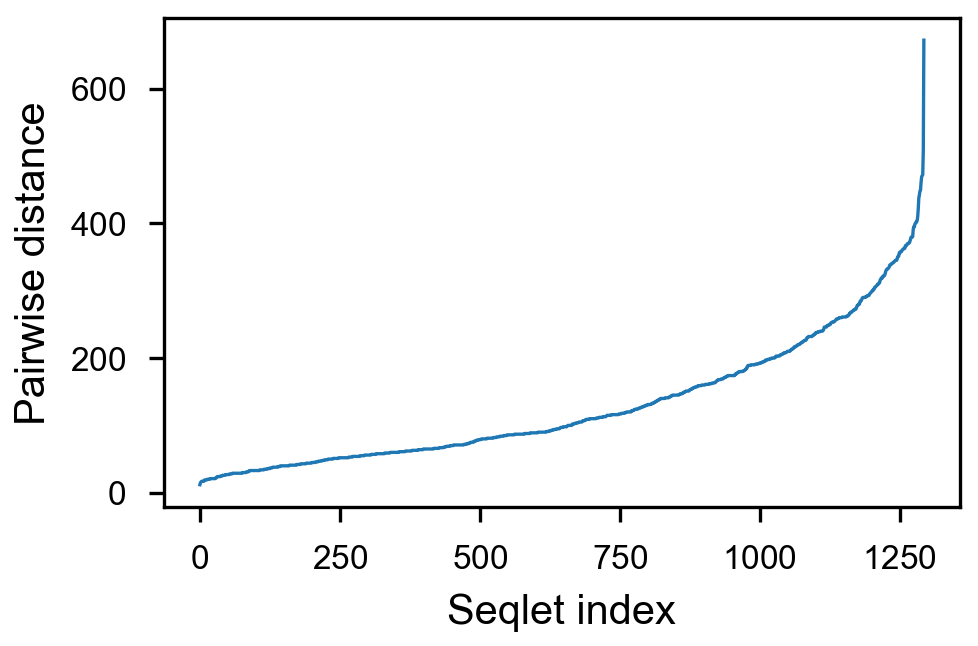
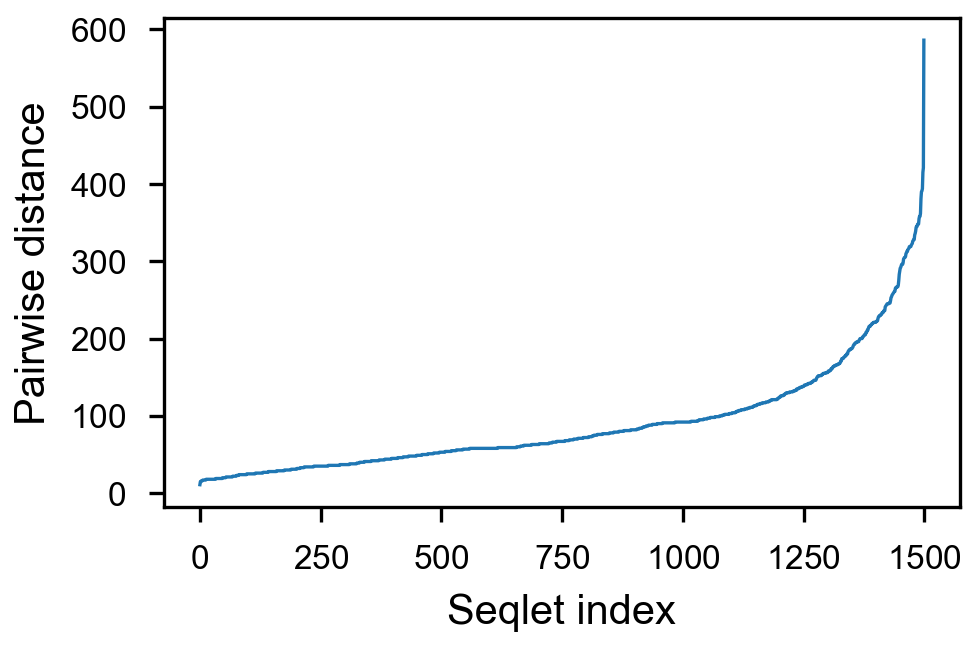
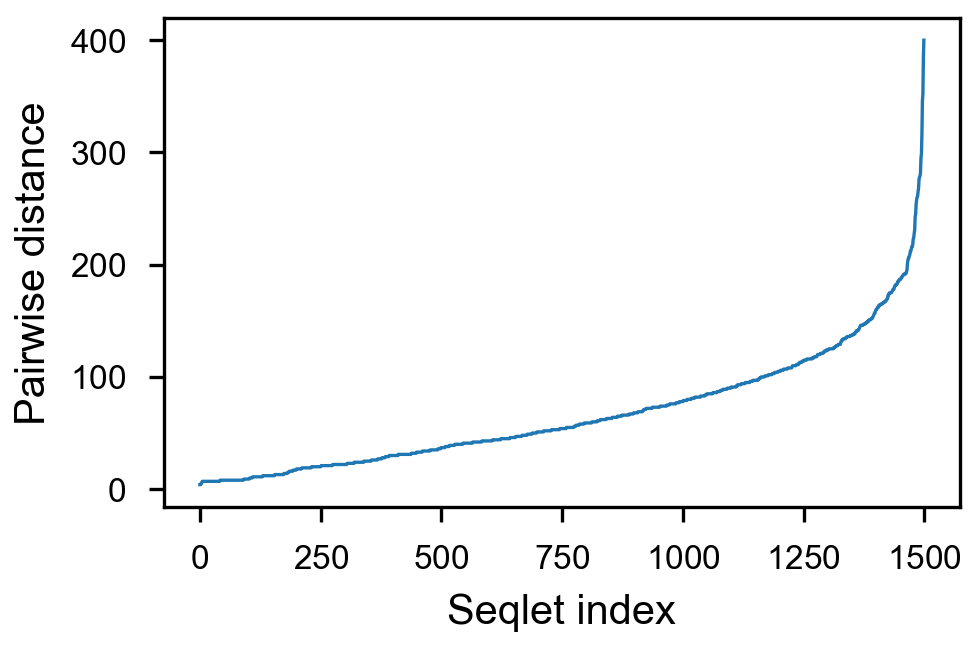
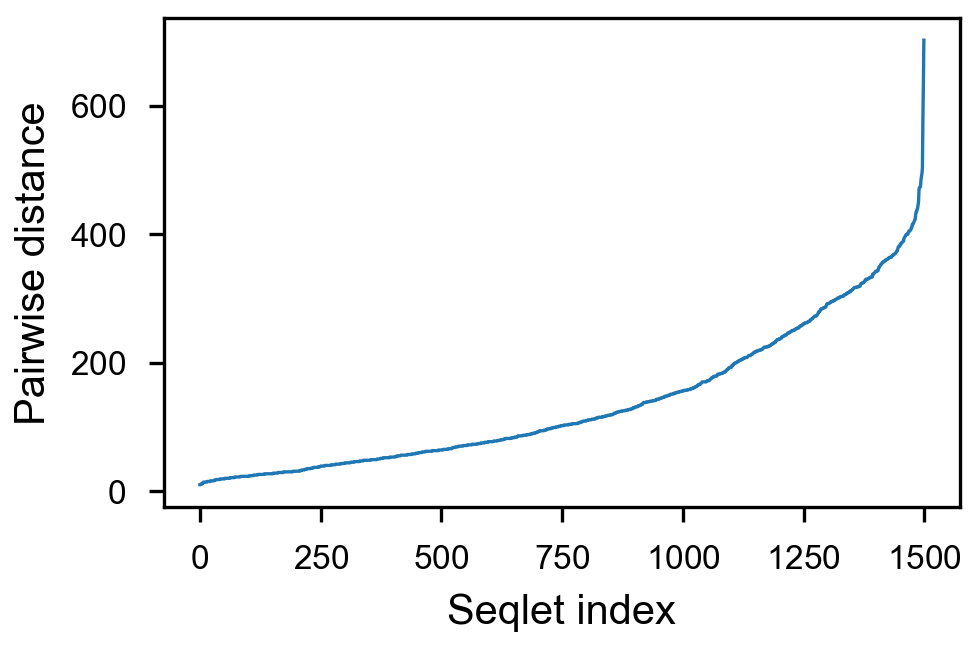
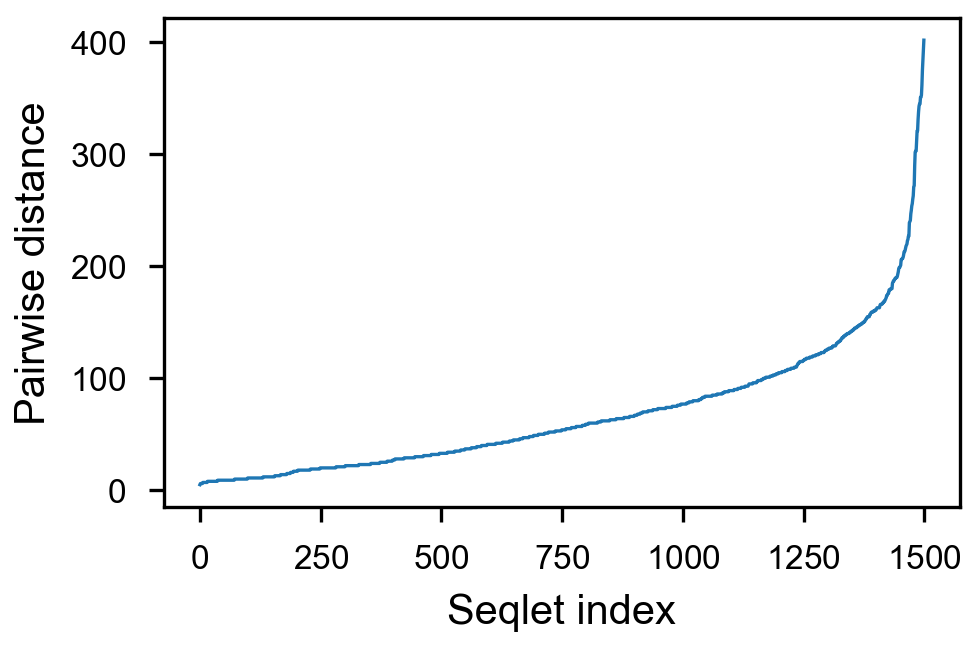
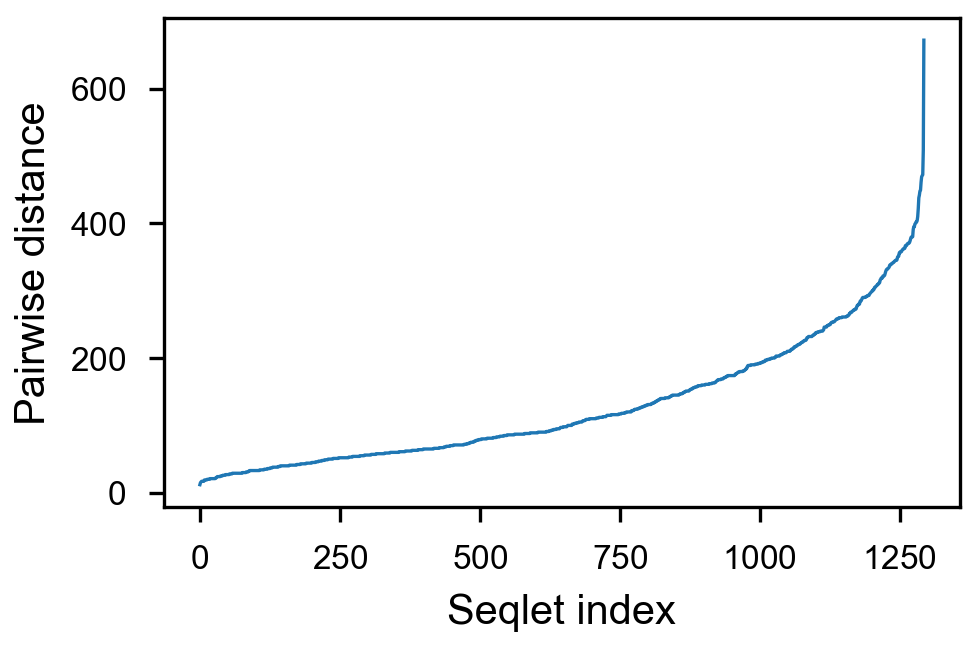
- [x] take top 1500 values per counts, sort by pairwise distance
- [x] try out different normalizers
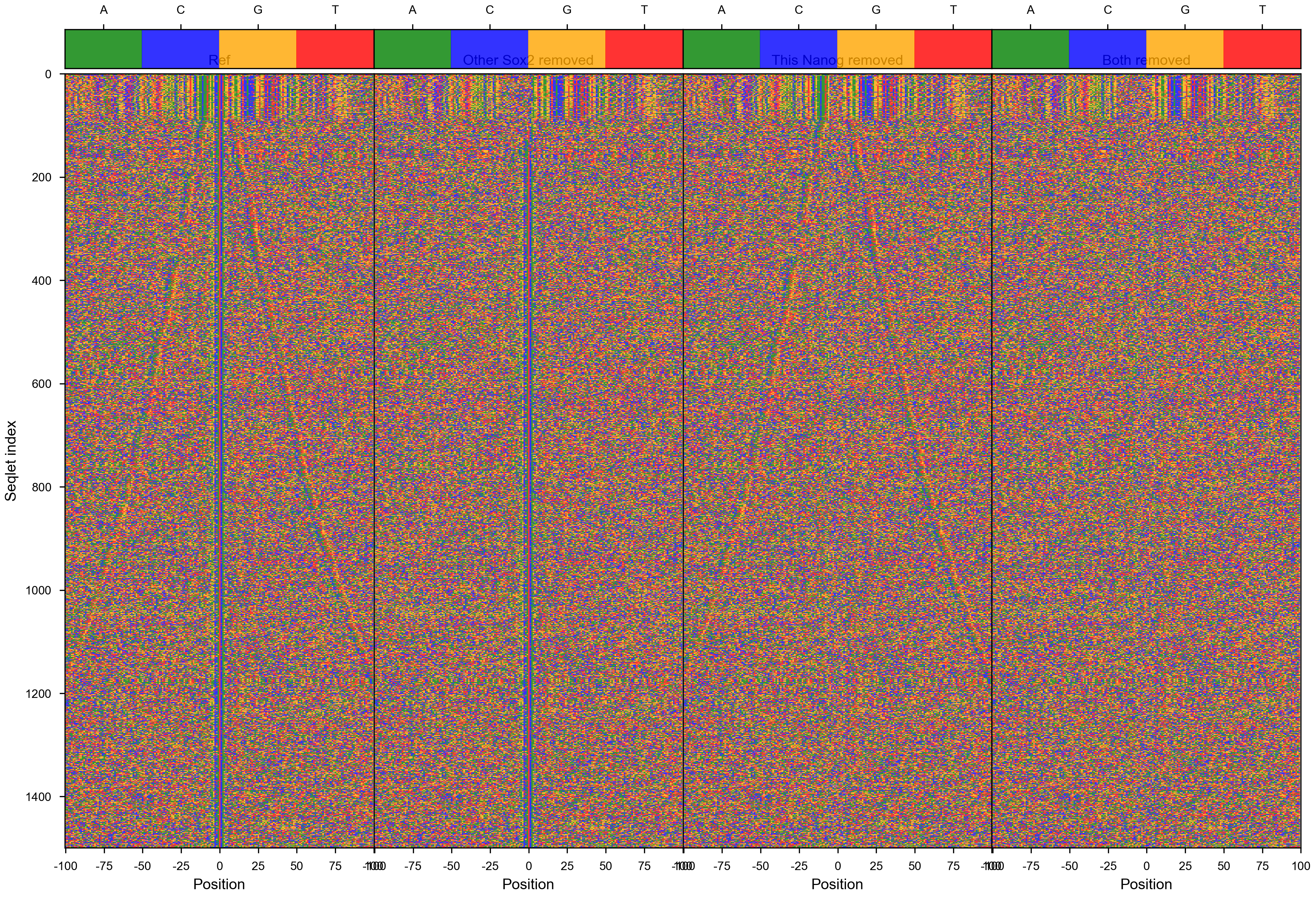
- [x] add also the sequence plot
- [x] eyeball the plots
- what would be a good way to quantify the effects? - importance scores
- [x] implement different feature functions quantifying the interactions
- [x] given a list of arrays (pred, imp, imp_count), quantify the metric
- [x] combine differnet metrics with each other
- [x] add also the max-count metric
- positions with maximal number of counts (averaged per strands)
- [x] add scatterplots quantifying the interactions
- [ ] re-compute motif_centers and motif_diff using proper rounding schemes
TODO¶
- [ ] generate all the pdf plots
- [ ] for scatterplots, show only the corresponding TF in the main (triangular) figure
- [ ] for the supplementary figures generate all 40 plots
- [ ] use max count
- [ ] add the heatmap of A->B type of analysis
- [ ] show <35 and >35 distance versions
- [ ] make sure colors look good and are consistent
- [ ] switch colors so that red means the 'interesting' effect
- [ ] simulation - generate the max score (same as in the perturbation analysis)
In [1]:
from basepair.utils import read_pkl
from pathlib import Path
from basepair.exp.paper.config import motifs, profile_mapping
from basepair.plot.config import paper_config, get_figsize
from basepair.exp.chipnexus.perturb.vdom import vdom_motif_pair, plot_spacing_hist
from basepair.exp.chipnexus.spacing import remove_edge_instances, get_motif_pairs
from basepair.plot.heatmaps import RowQuantileNormalizer, QuantileTruncateNormalizer
from tqdm import tqdm
from basepair.data import NumpyDataset
from basepair.exp.chipnexus.perturb.gen import *
from basepair.exp.chipnexus.perturb.scores import *
from copy import deepcopy
from plotnine import *
import plotnine
import warnings
warnings.filterwarnings("ignore")
from basepair.config import get_data_dir
paper_config()
ddir = get_data_dir()
In [2]:
# Common paths
model_dir = Path(f"{ddir}/processed/chipnexus/exp/models/oct-sox-nanog-klf/models/n_dil_layers=9/")
modisco_dir = model_dir / f"modisco/all/deeplift/profile/"
output_dir = modisco_dir
dataset_dir = output_dir / 'perturbation-analysis'
In [3]:
pairs = get_motif_pairs(motifs)
In [4]:
# load the data
motif_pair_lpdata = read_pkl(dataset_dir / 'motif_pair_lpdata.incl-whole.pkl')
dfab = pd.read_csv(dataset_dir / 'dfab.csv.gz')
In [5]:
dfi_subset = pd.read_csv(dataset_dir / 'dfi_subset.csv.gz')
In [6]:
# write_pkl(motif_pair_lpdata, dataset_dir / 'motif_pair_lpdata.incl-whole.pkl')
In [7]:
# TODO - de-hardcode
tasks = ['Oct4', 'Sox2', 'Nanog', 'Klf4']
Basic stats¶
In [ ]:
%matplotlib inline
In [ ]:
paper_config()
In [26]:
fig = plt.figure(figsize=get_figsize(.5))
dfab.groupby("motif_pair").size().plot.barh()
plt.axvline(1500) # displayed cutoff
plt.xlabel("Number of pairs");
Pairwise spacing¶
In [ ]:
dfab['motif_pair_cat'] = pd.Categorical(dfab.motif_pair, categories=dfab.groupby("motif_pair").size().sort_values(ascending=False).index)
dfab['strand_combination_cat'] = pd.Categorical(dfab['strand_combination'], )
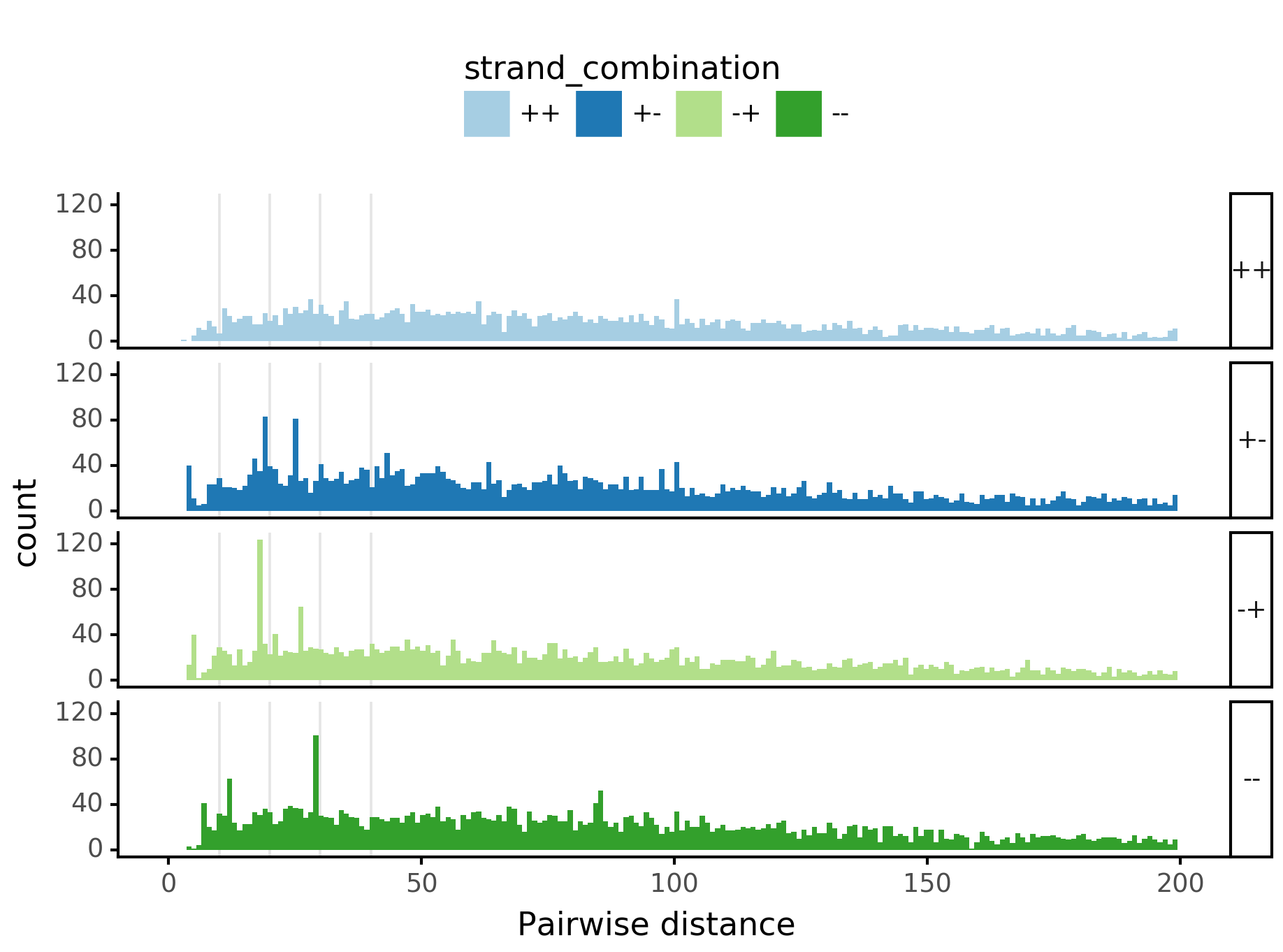
In [28]:
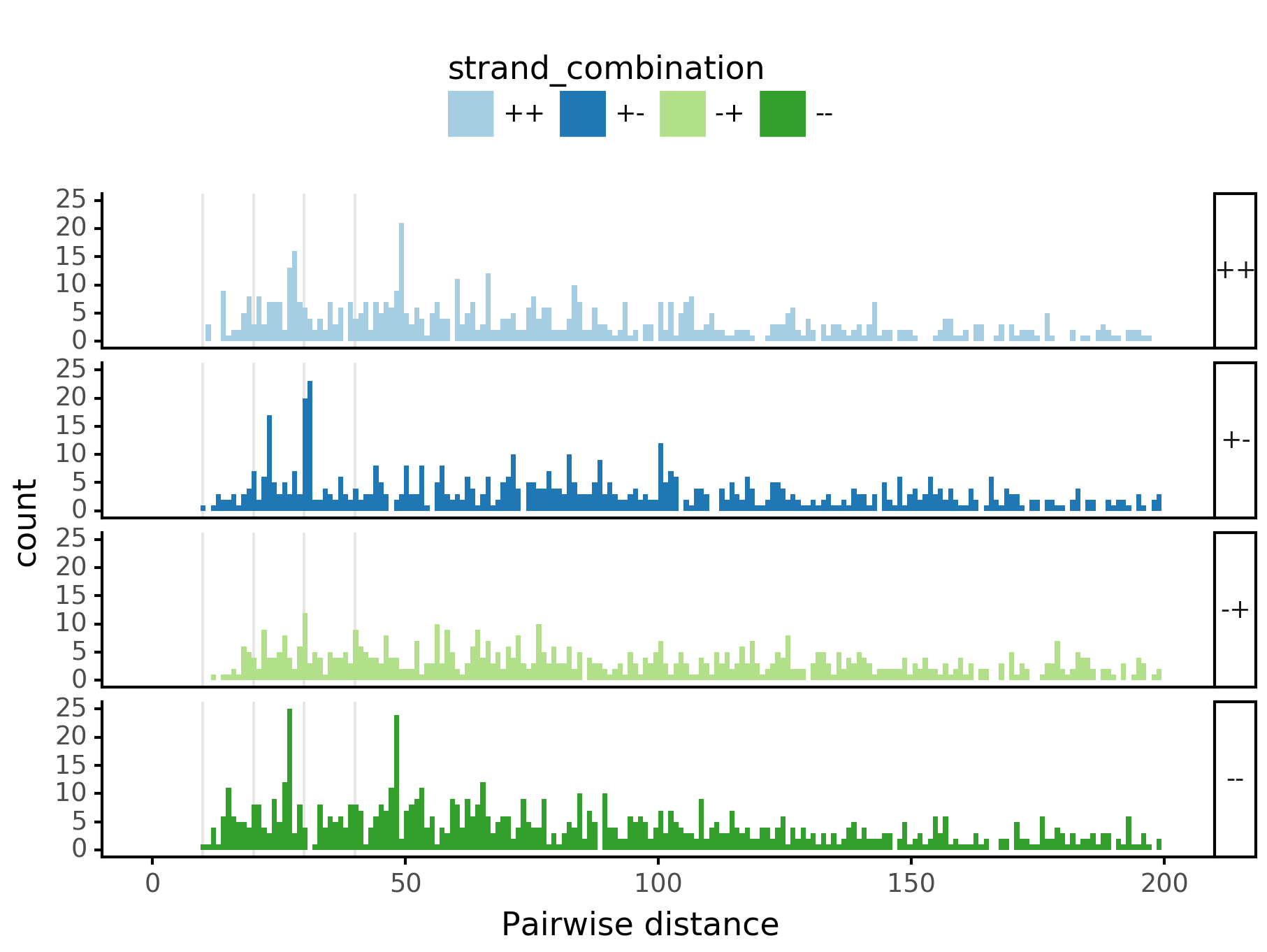
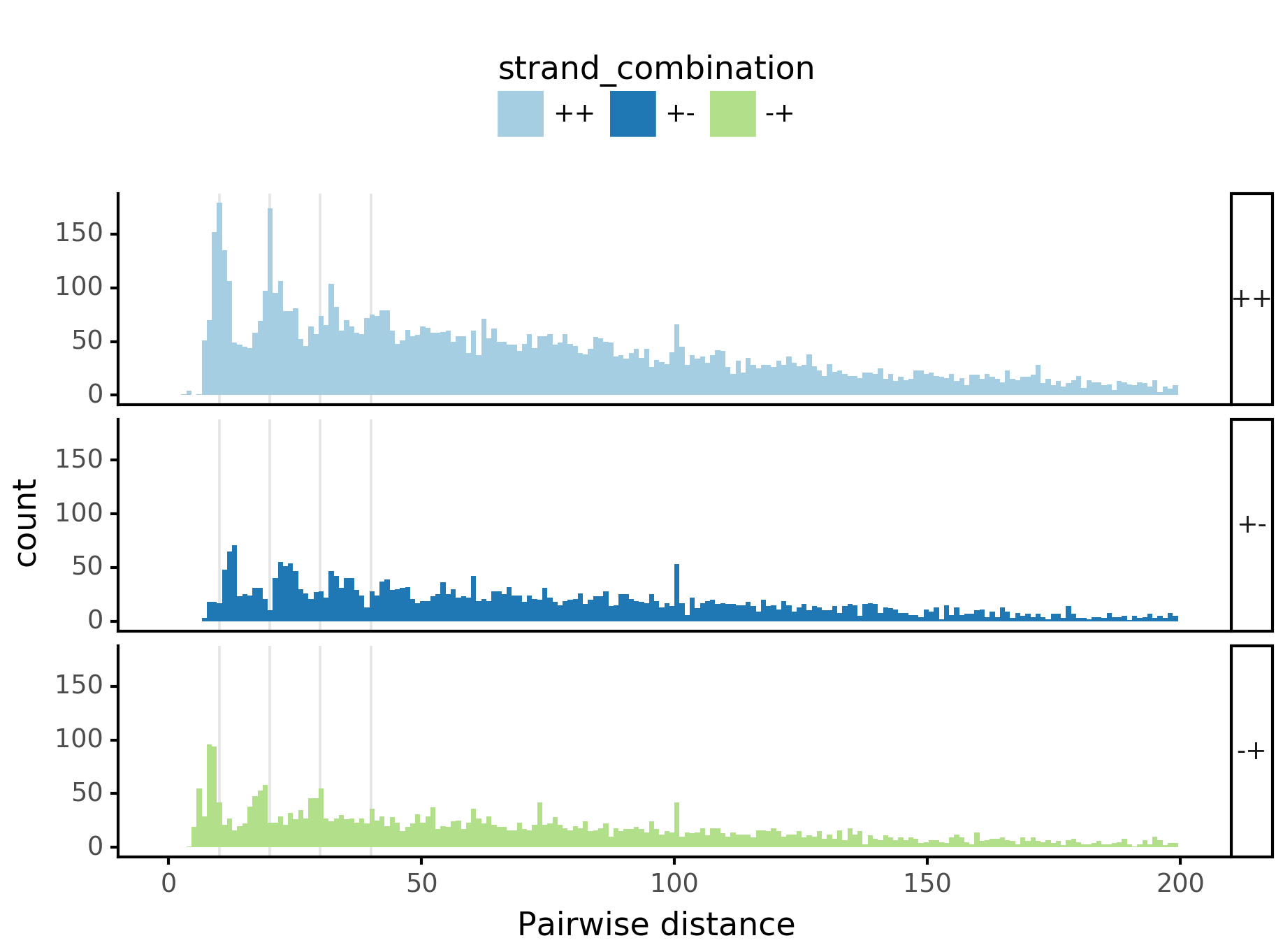
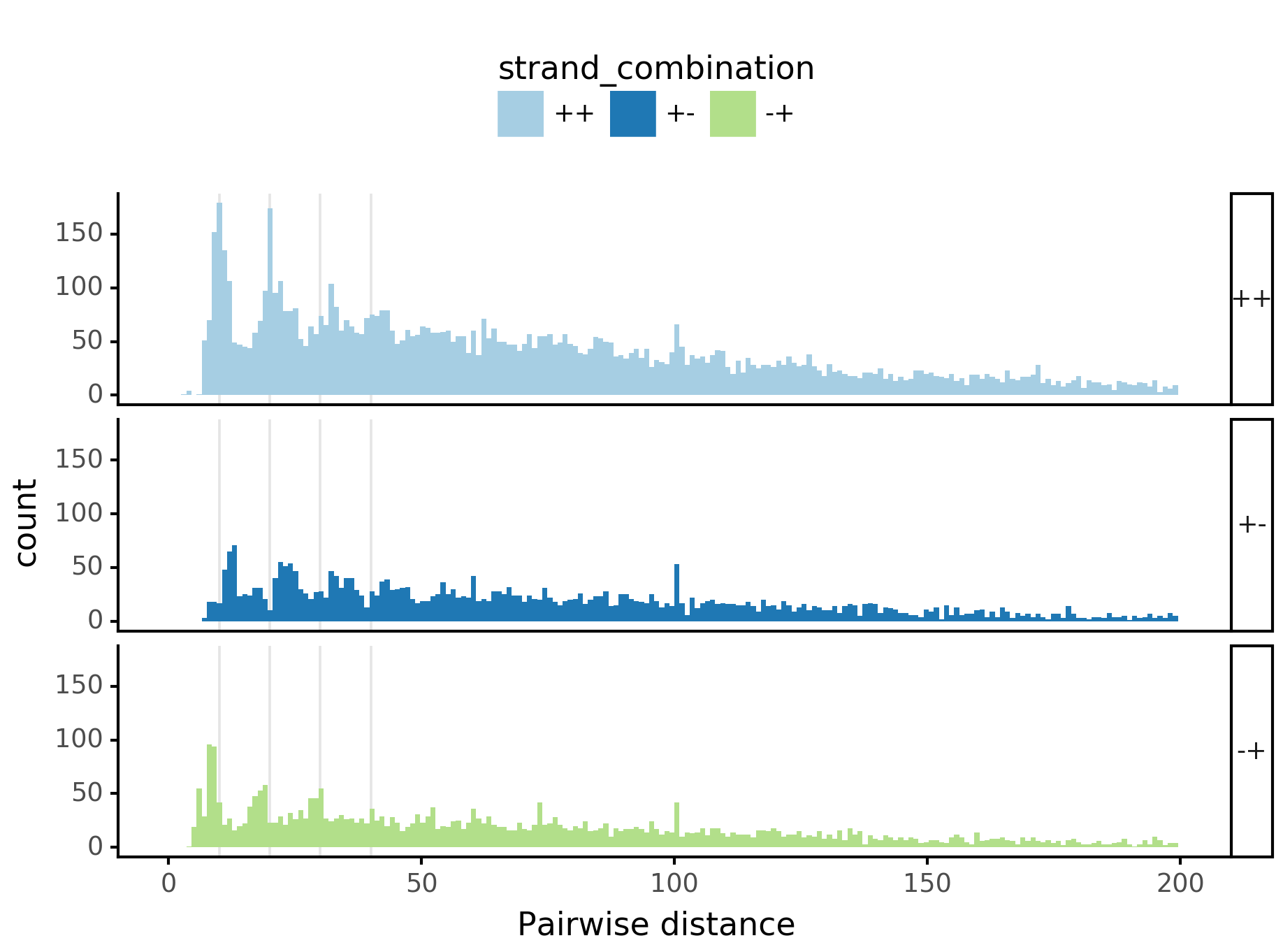
plotnine.options.figure_size = get_figsize(.5, aspect=2)# (10, 10)
max_dist = 100
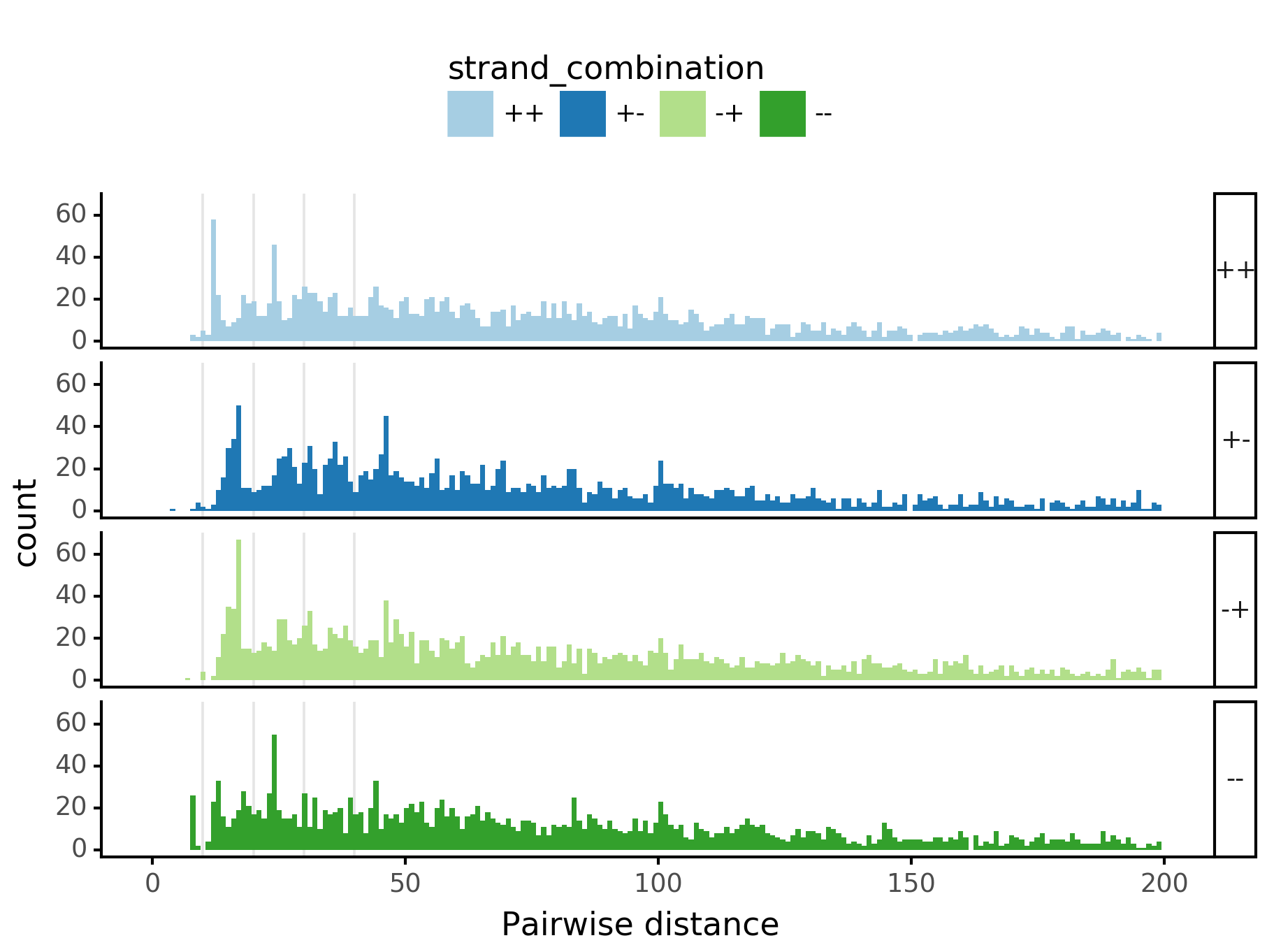
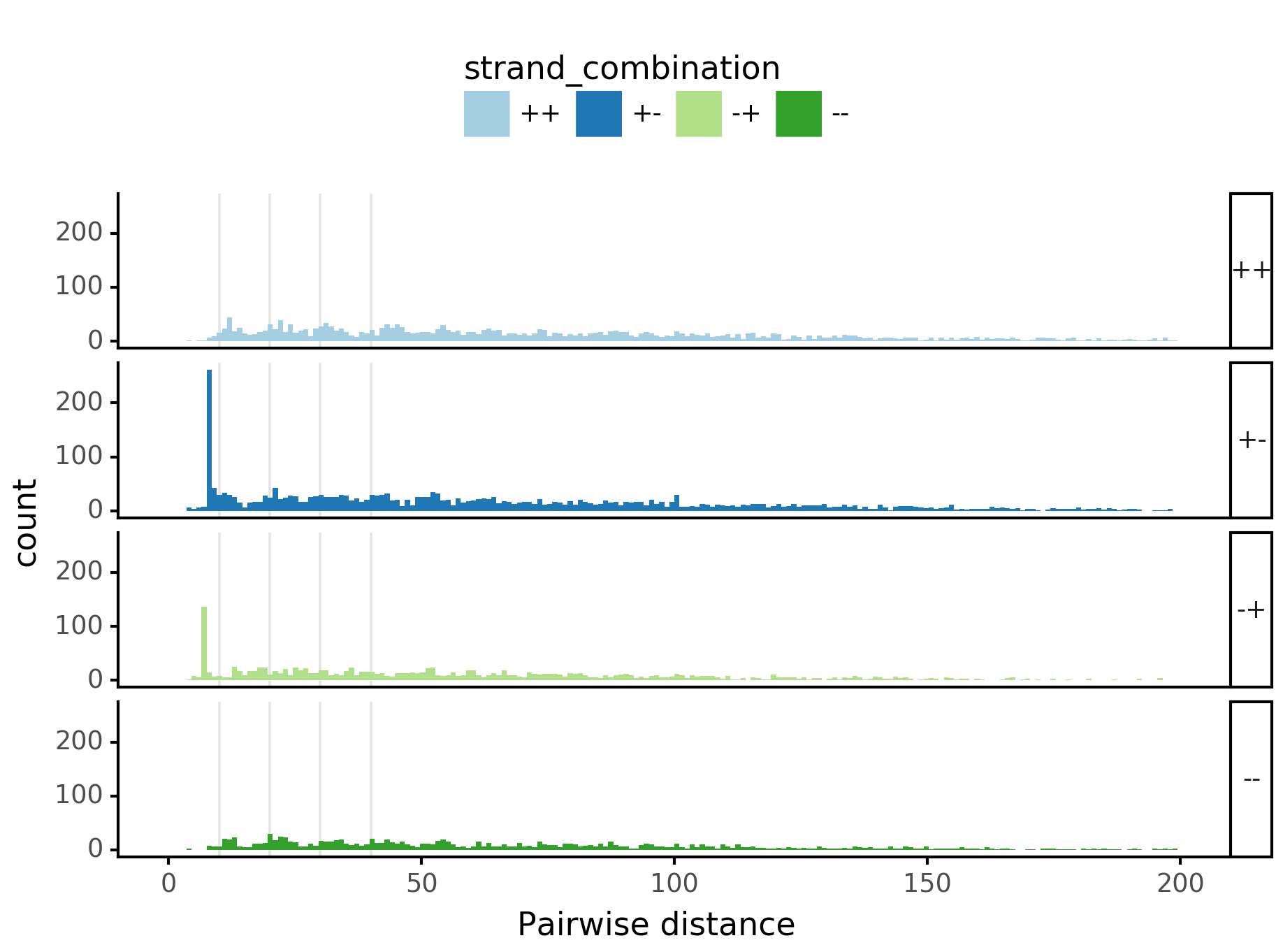
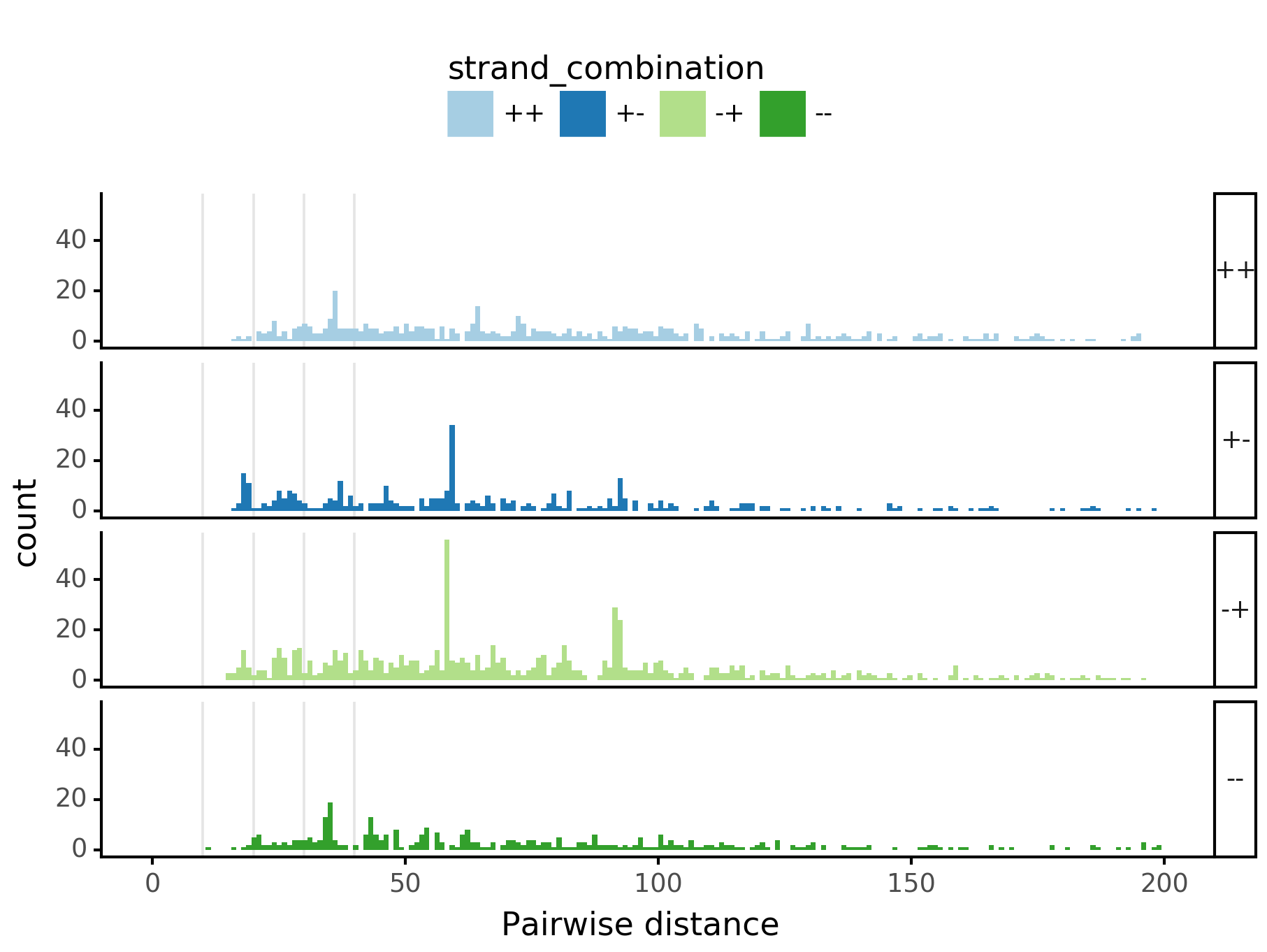
(ggplot(aes(x='center_diff', fill='strand_combination'), dfab[(dfab.center_diff <= max_dist)]) +
geom_histogram(bins=max_dist) +
facet_grid("motif_pair_cat~ .") +
theme_classic() +
theme(strip_text = element_text(rotation=0), legend_position='top') +
xlim([0, max_dist]) +
xlab("Pairwise distance") +
scale_fill_brewer(type='qual', palette=3))
Out[28]:
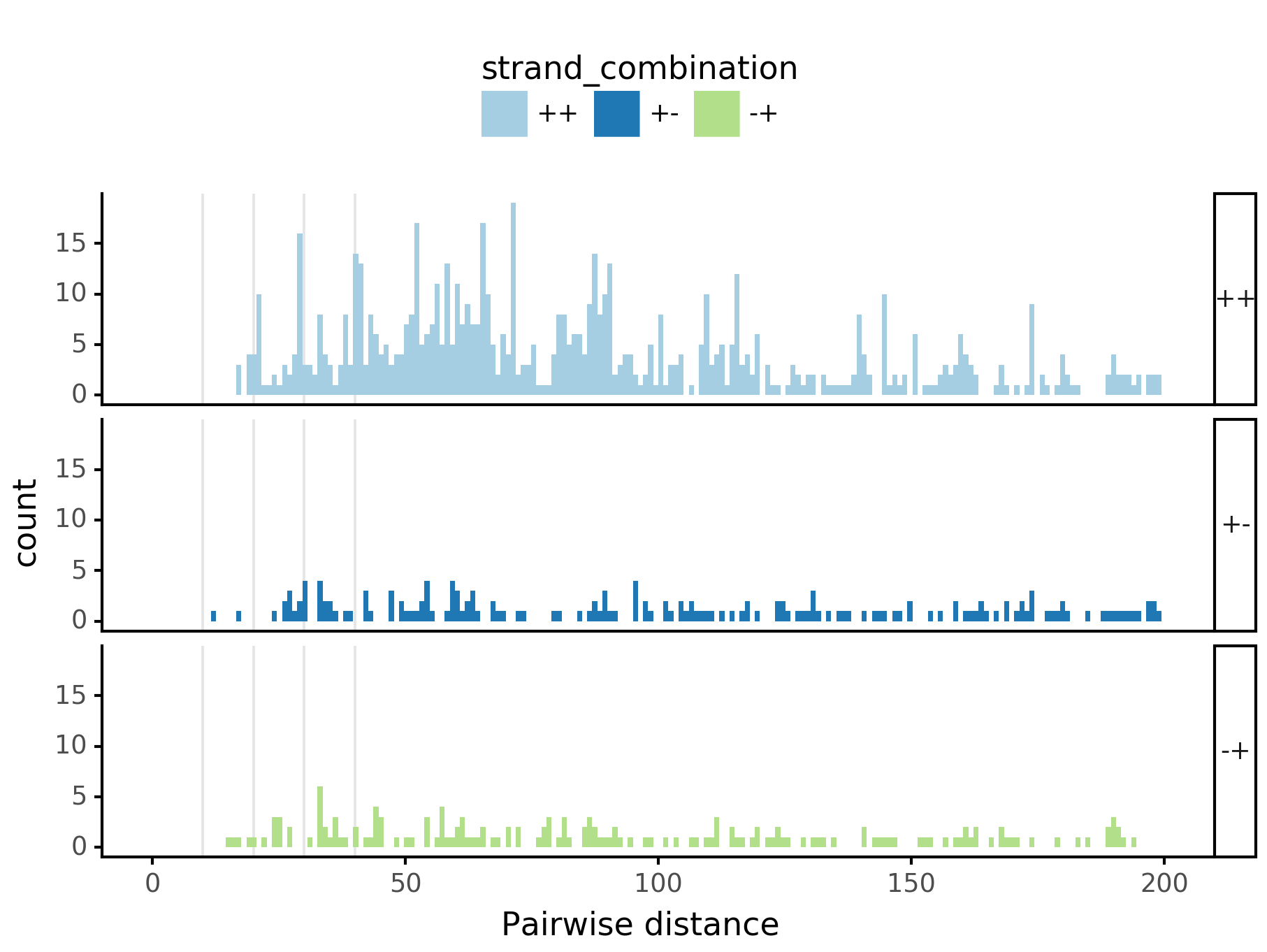
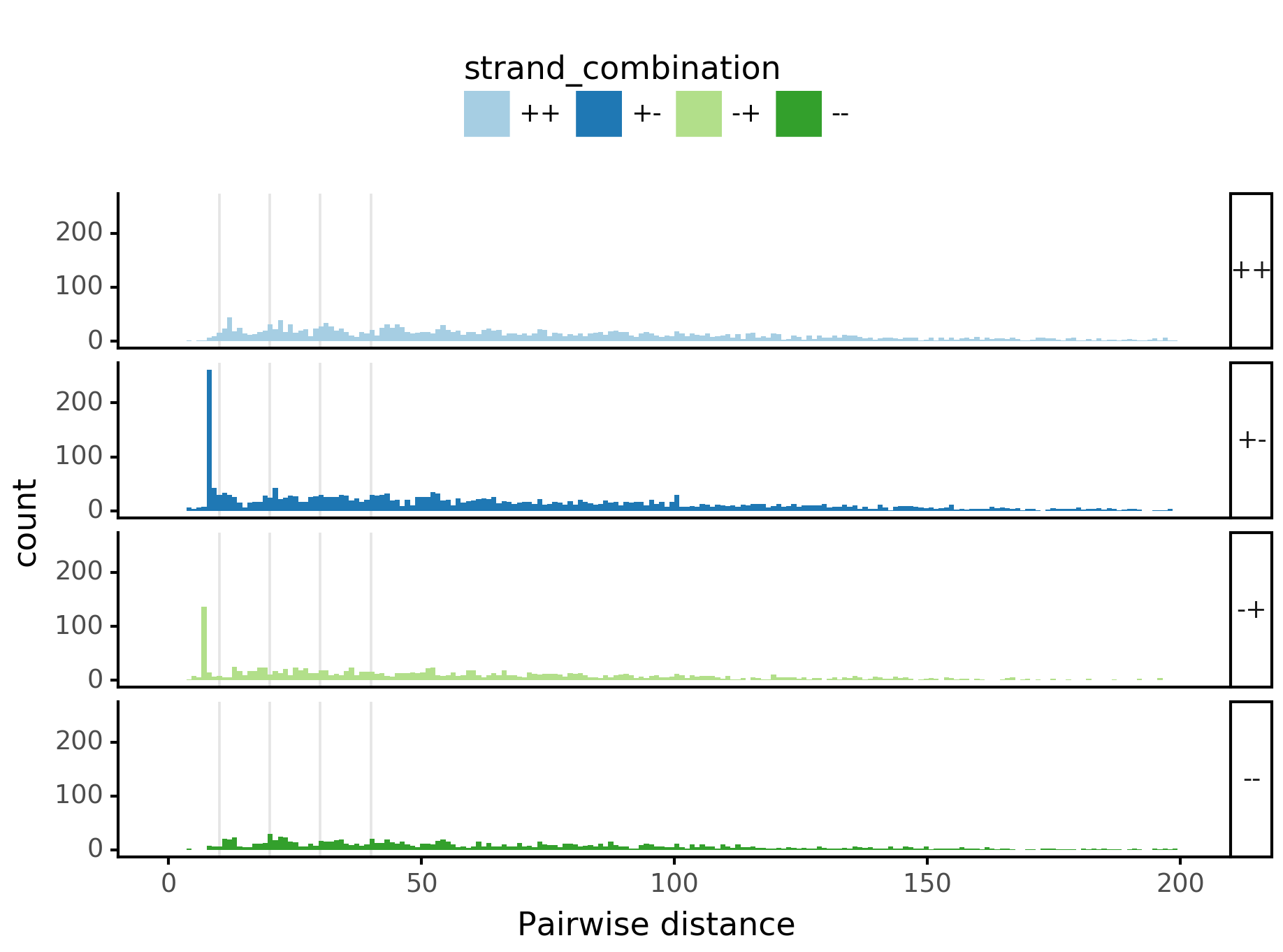
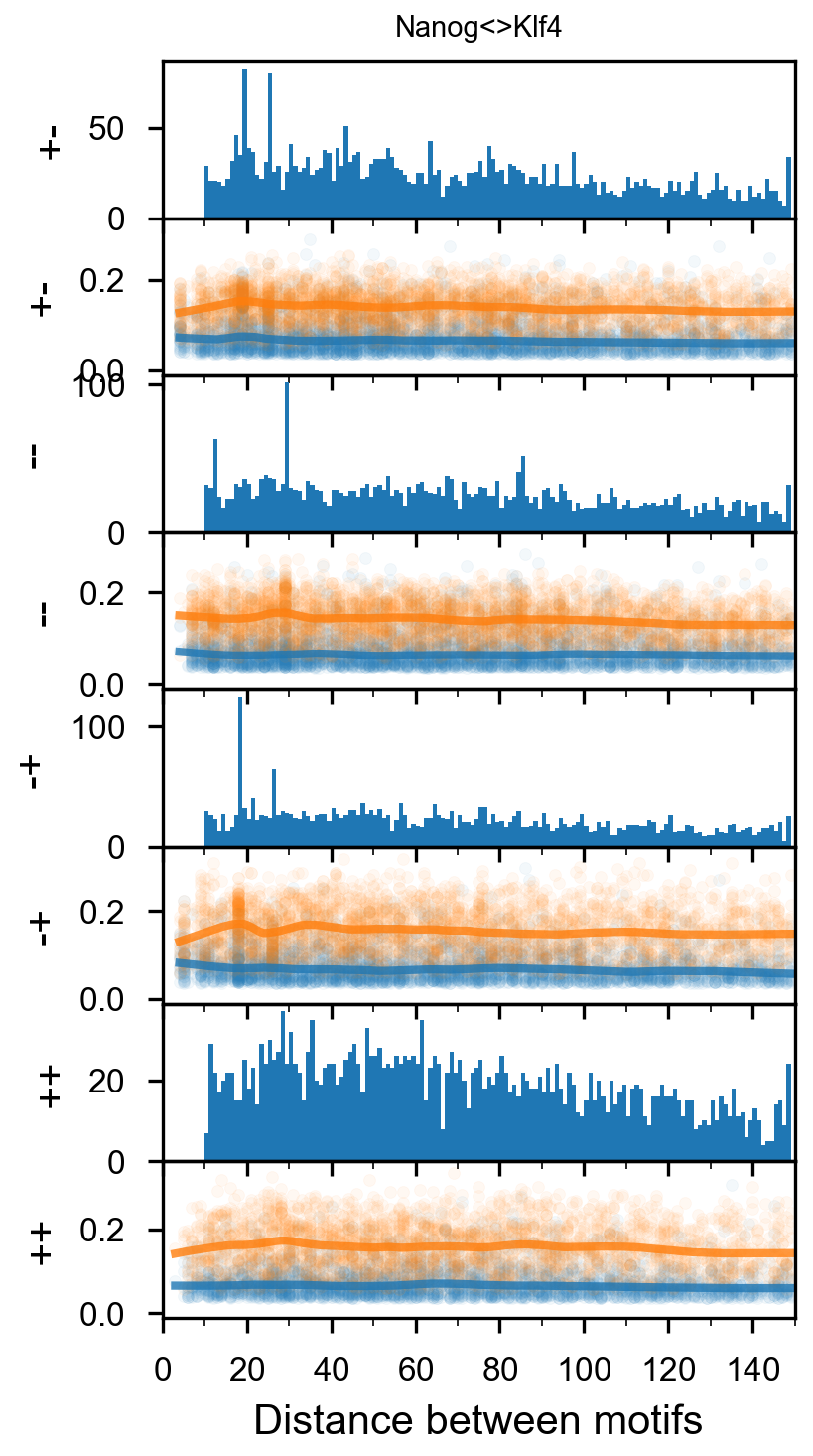
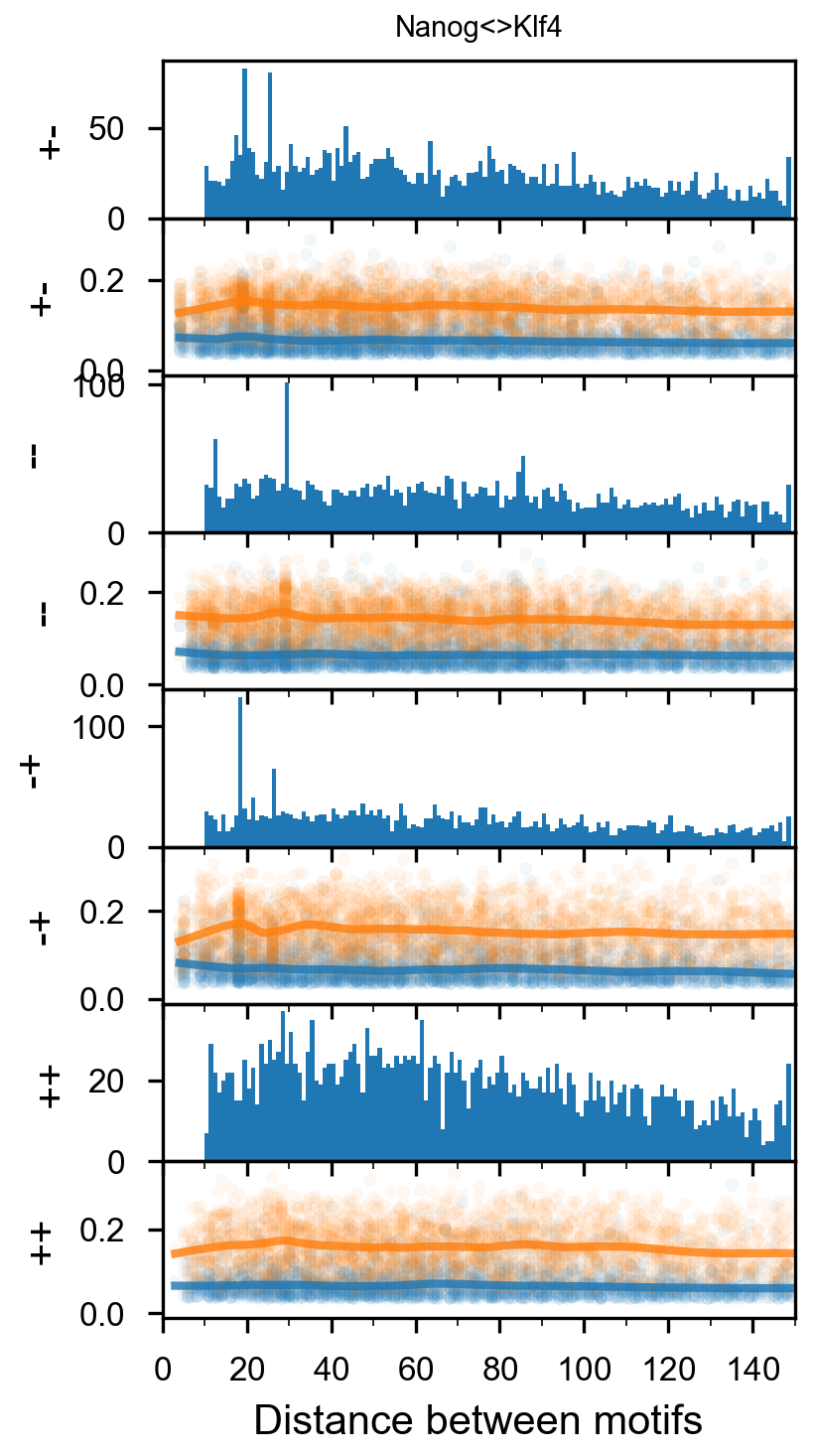
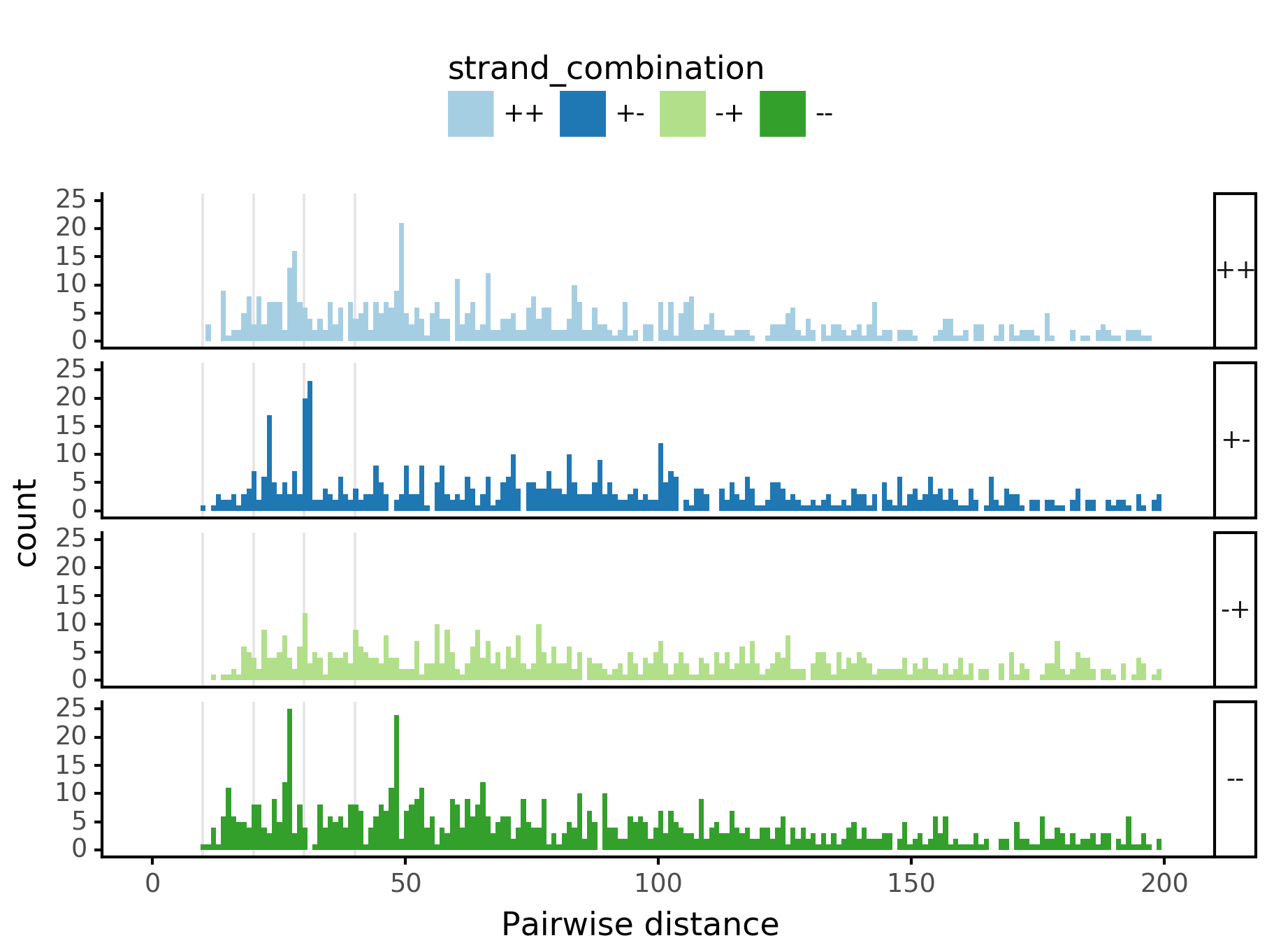
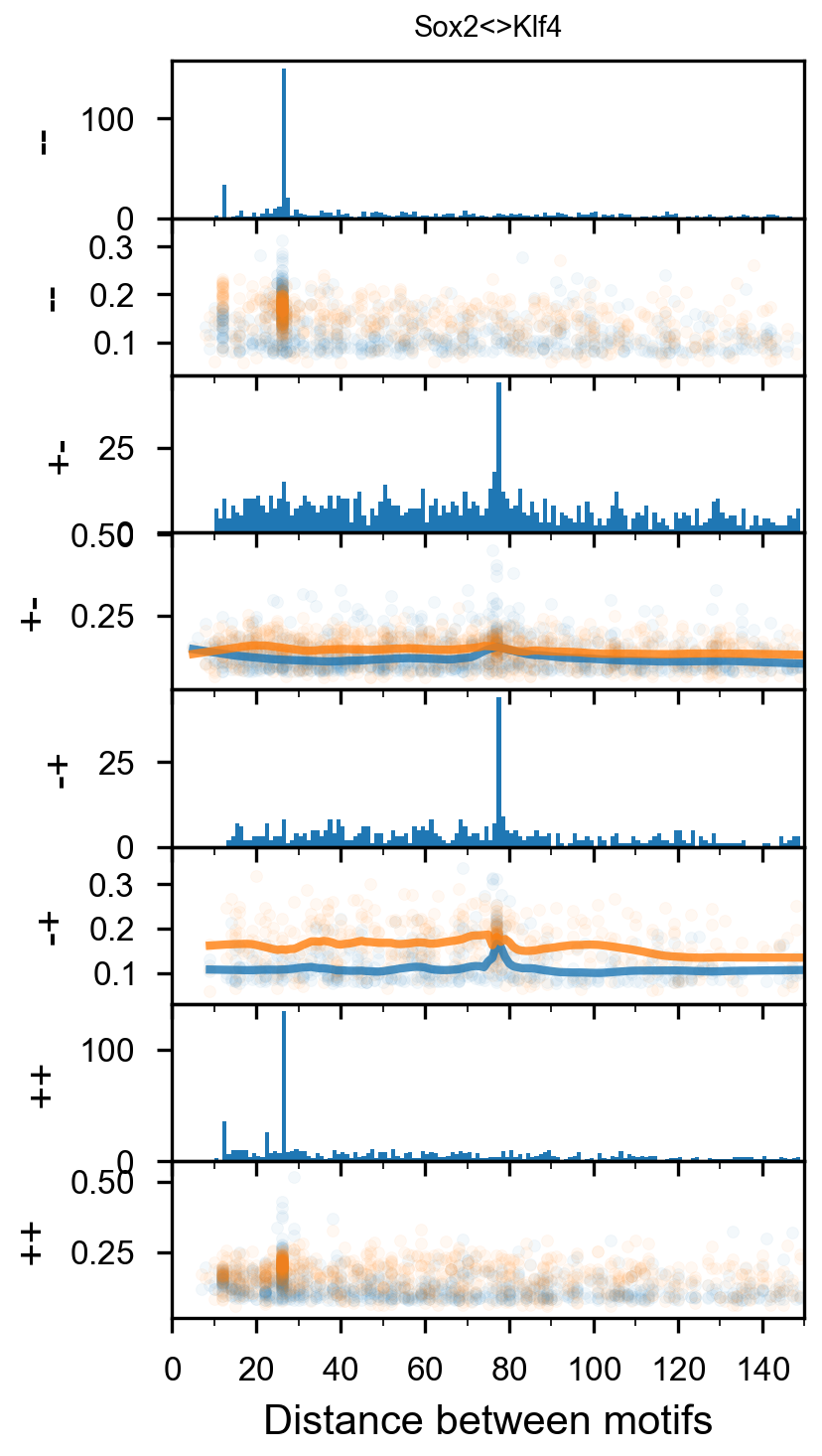
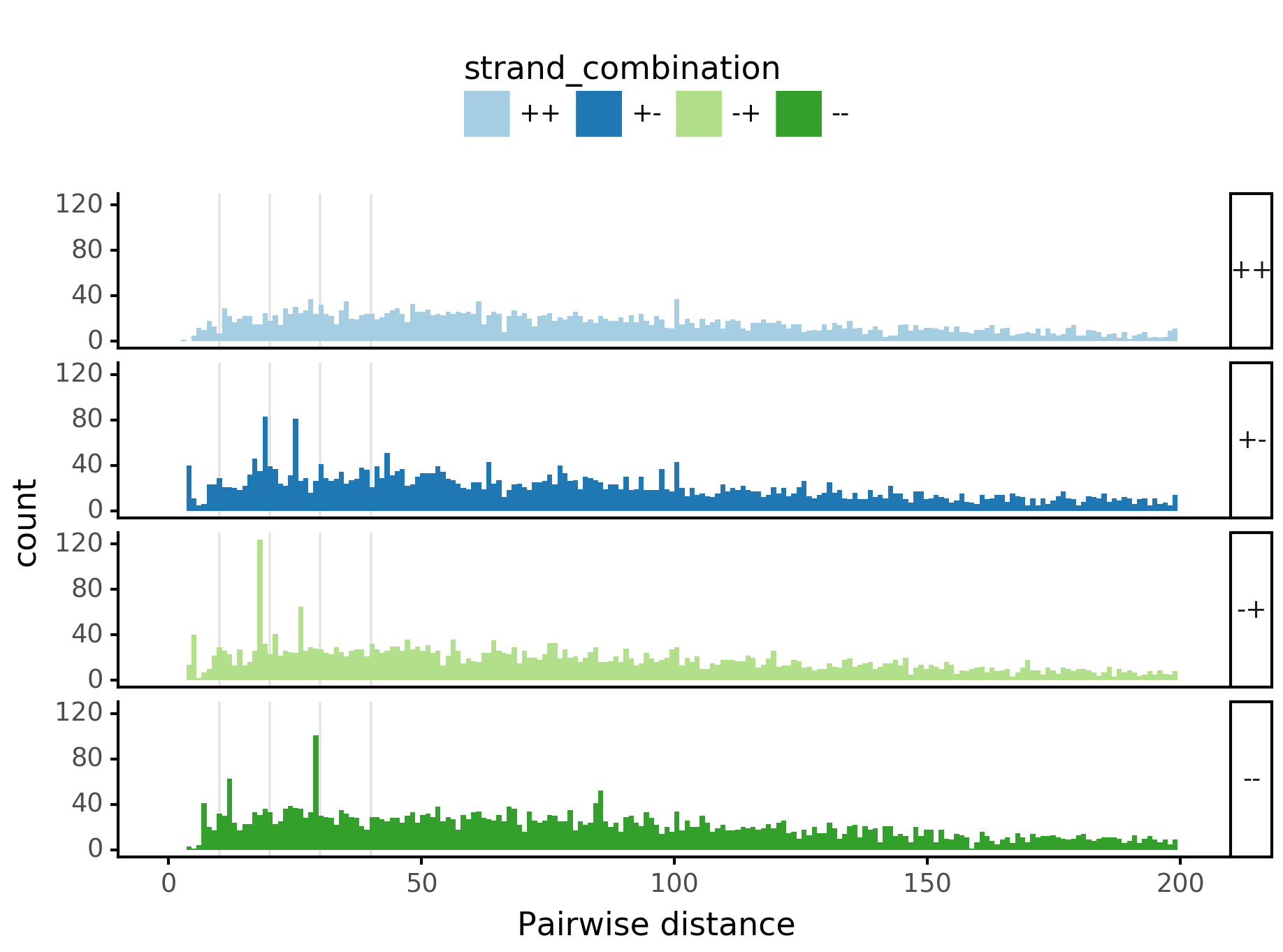
Pairwise spacing¶
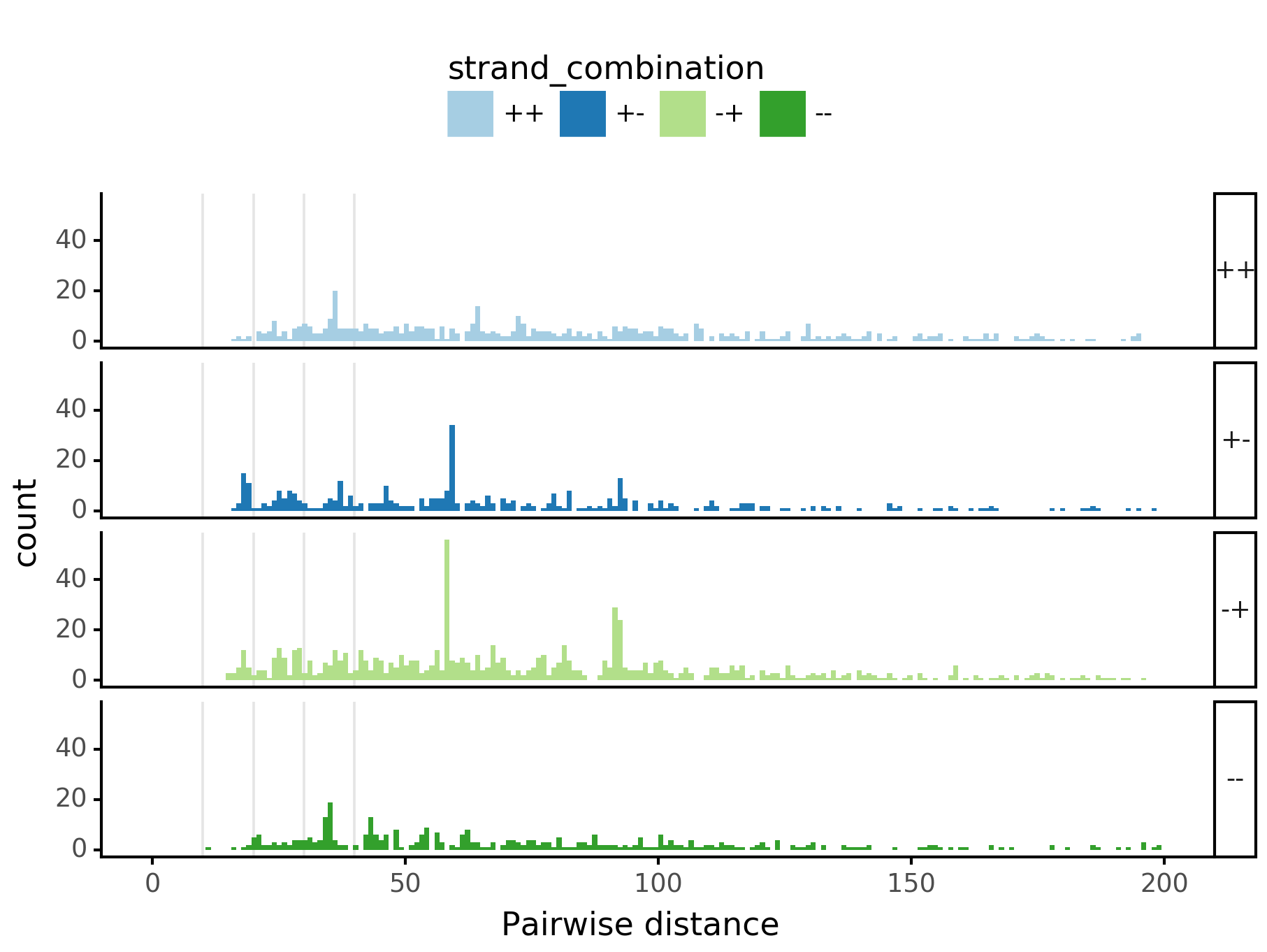
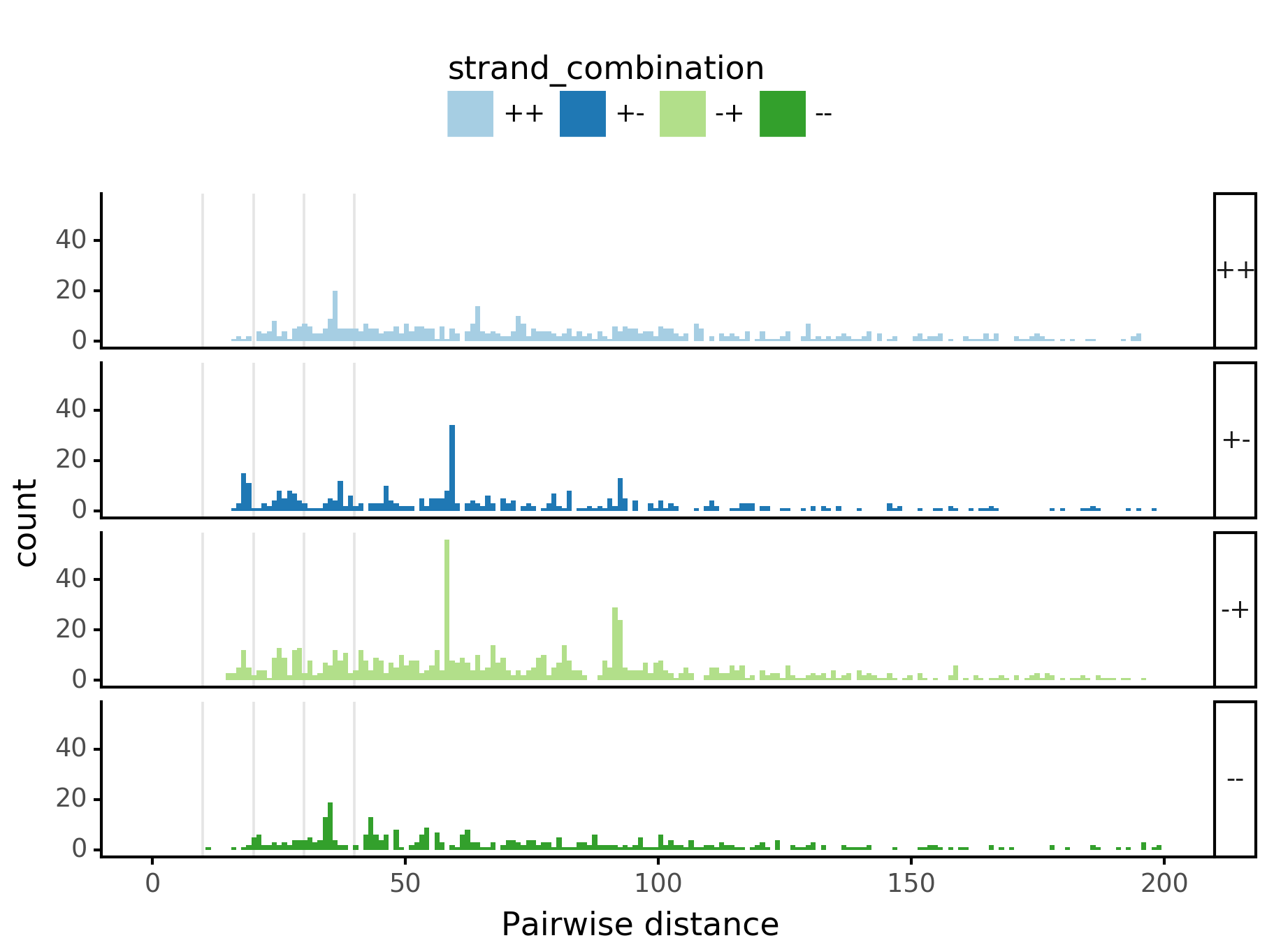
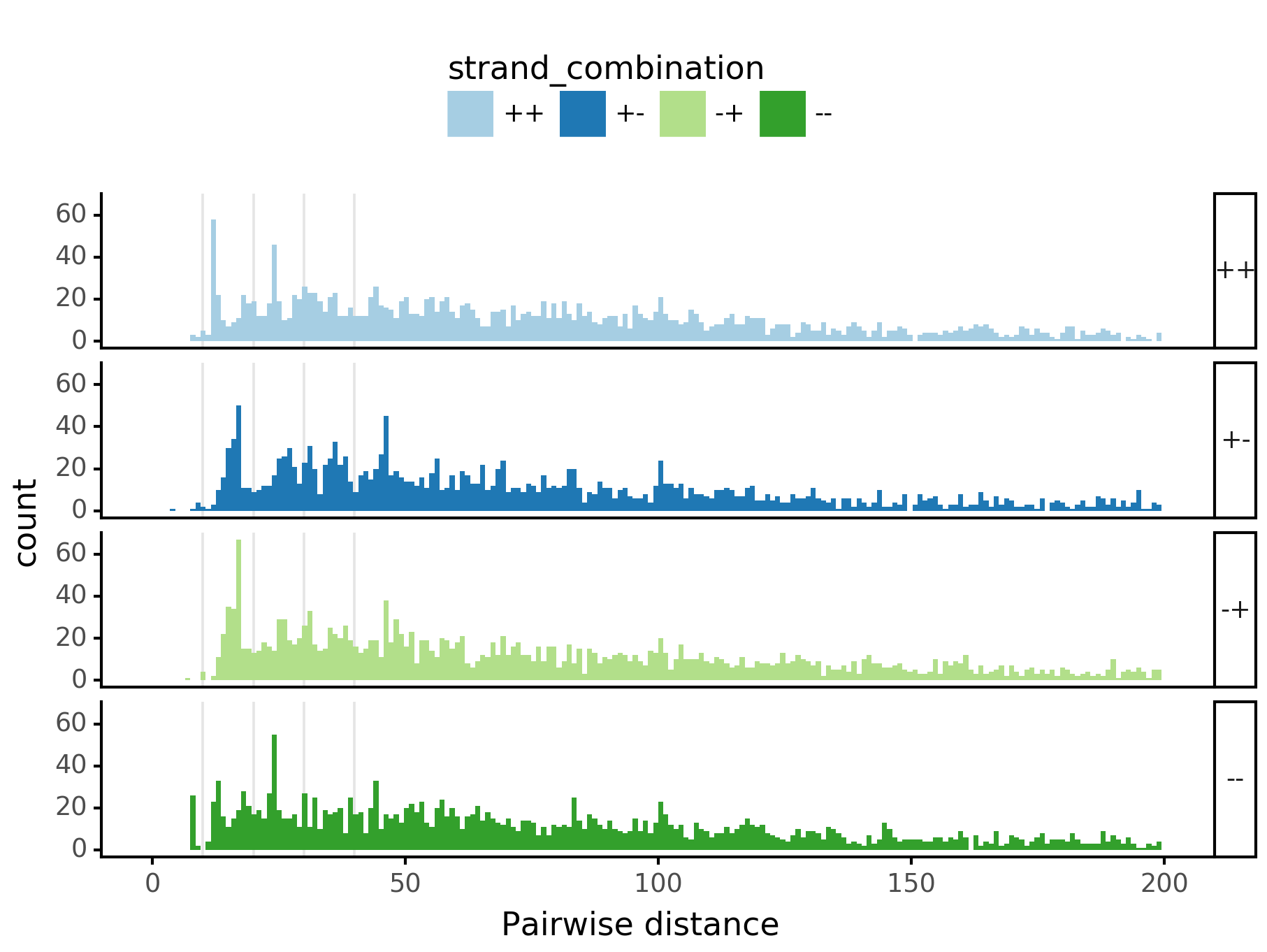
In [29]:
plotnine.options.figure_size = get_figsize(.5, aspect=2/10*3)
max_dist = 100
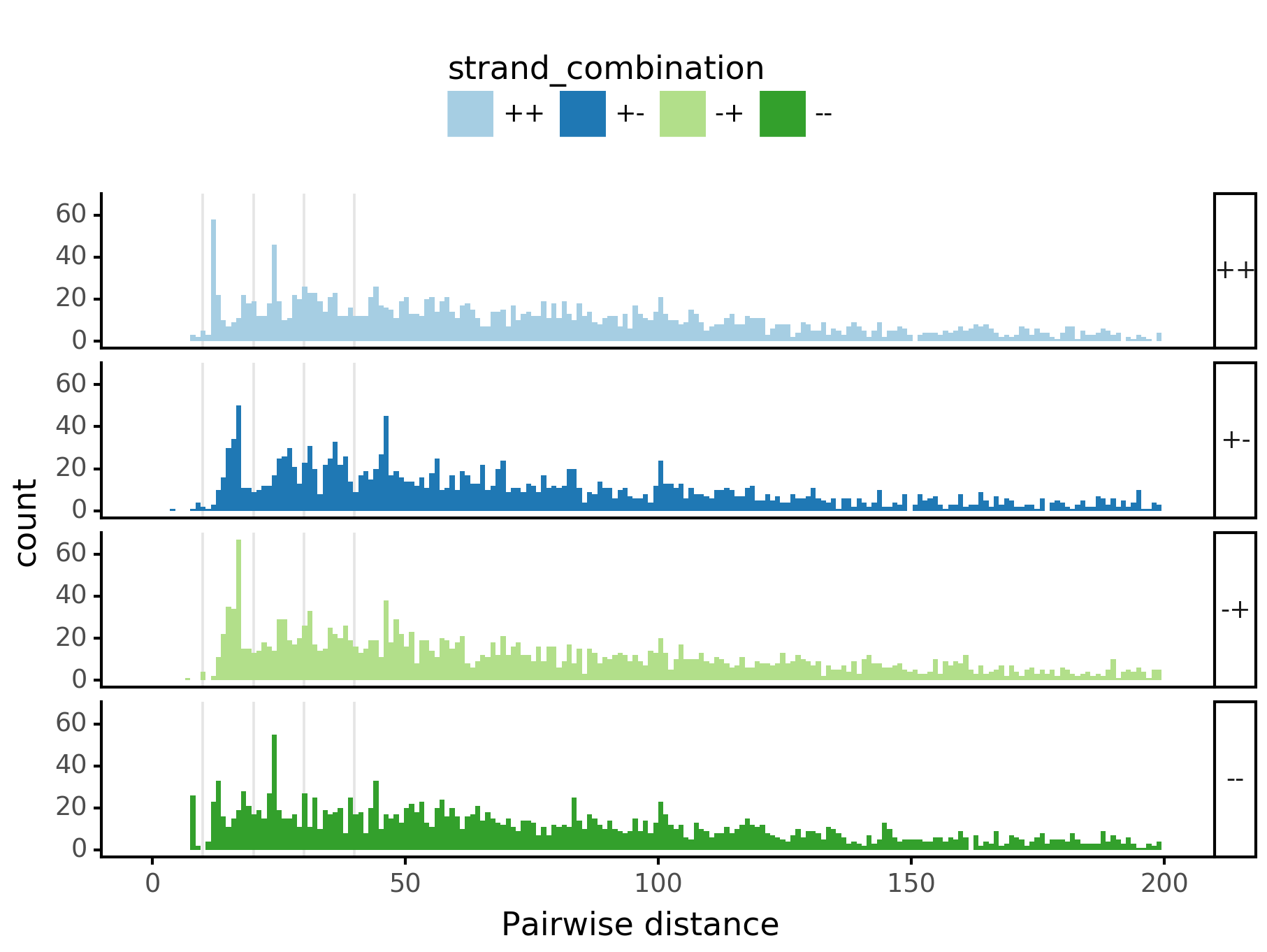
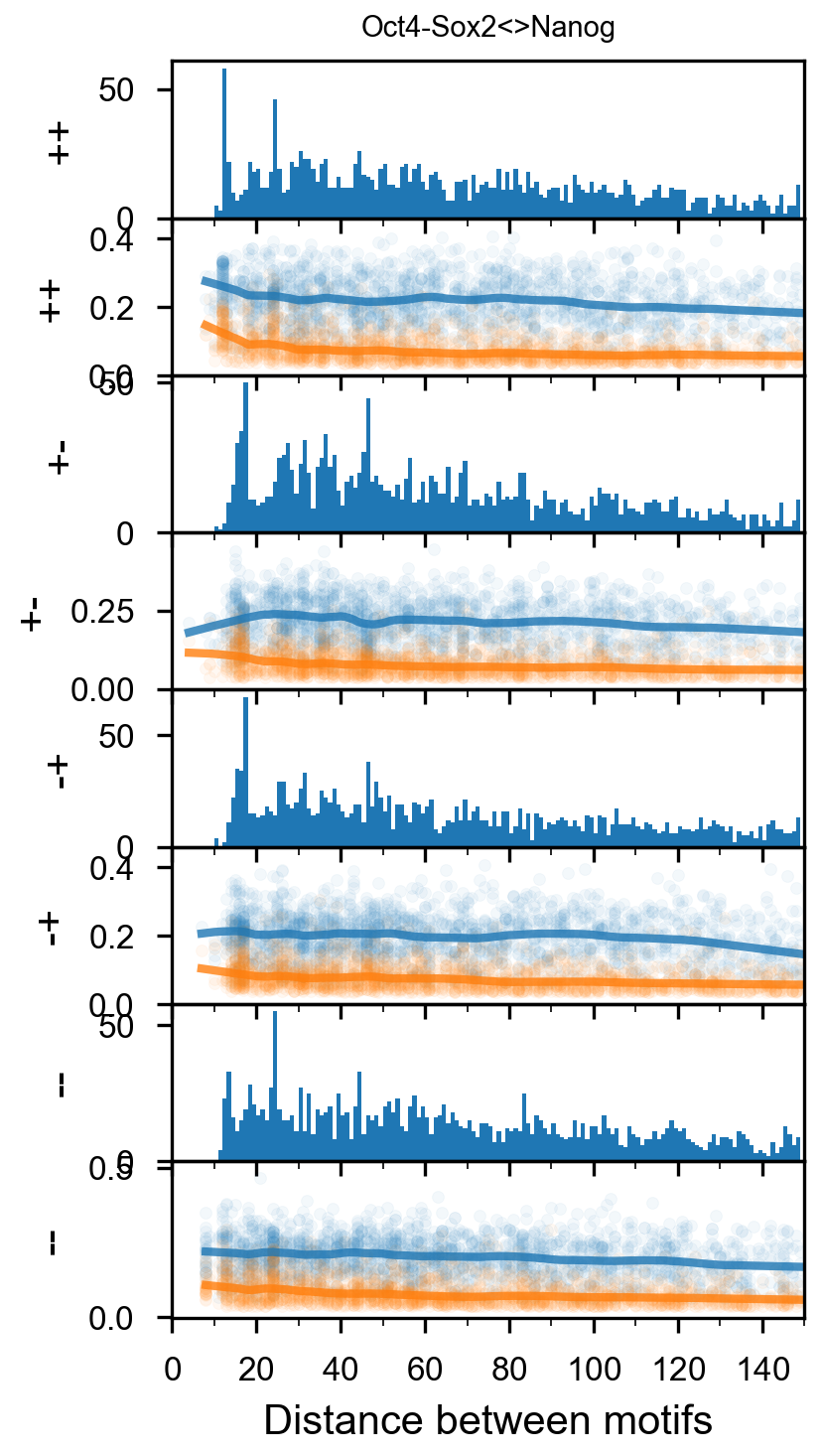
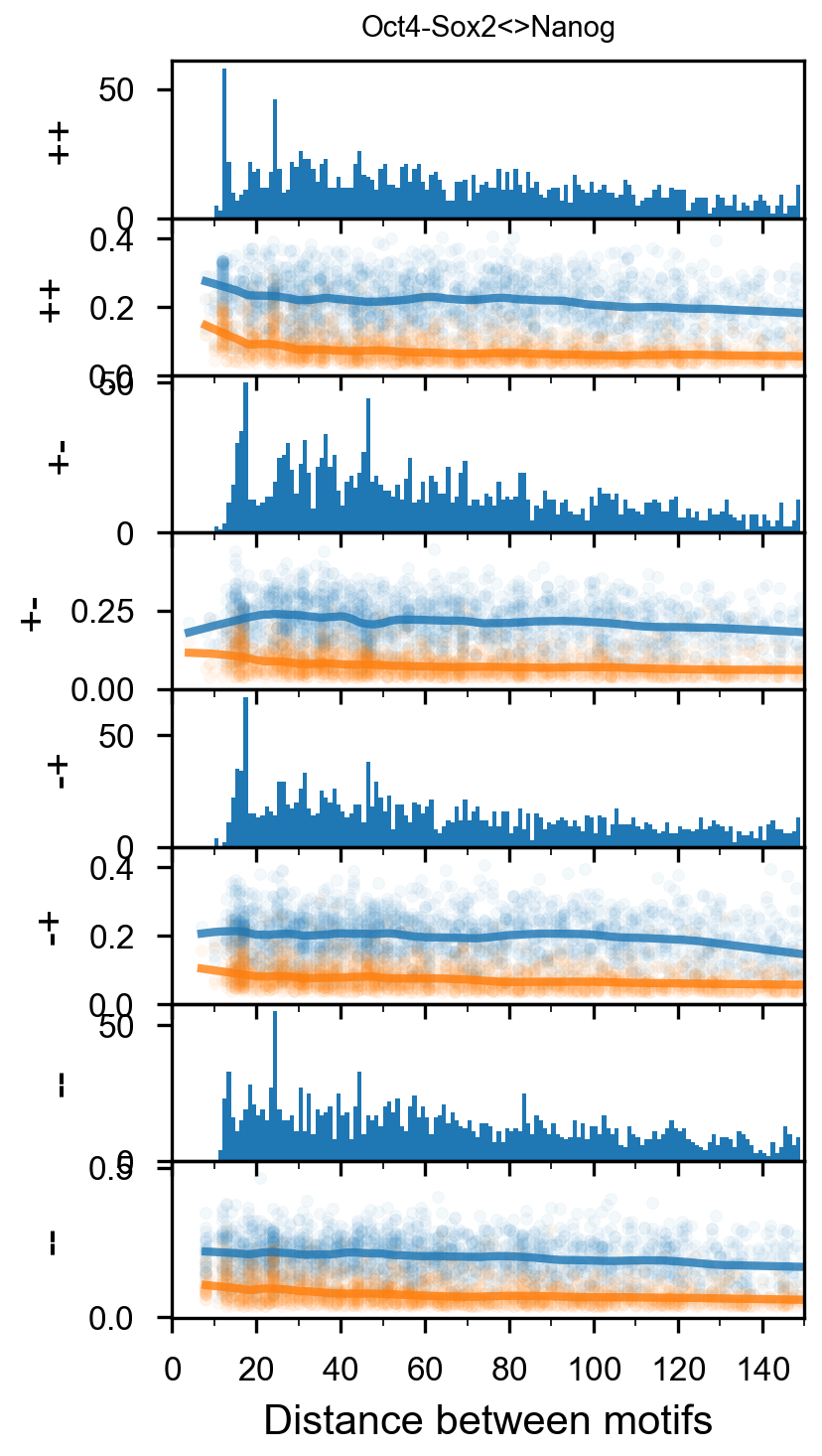
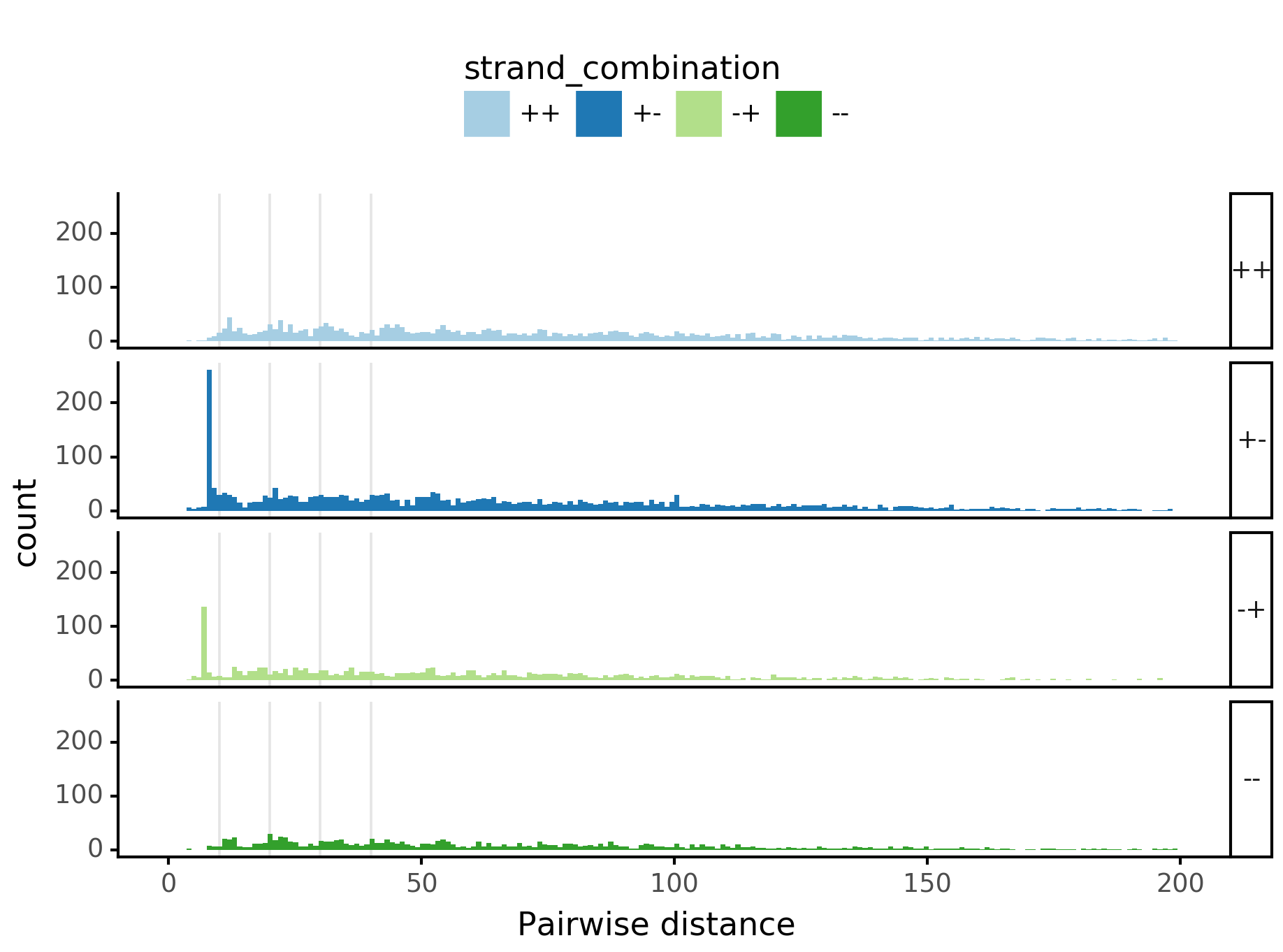
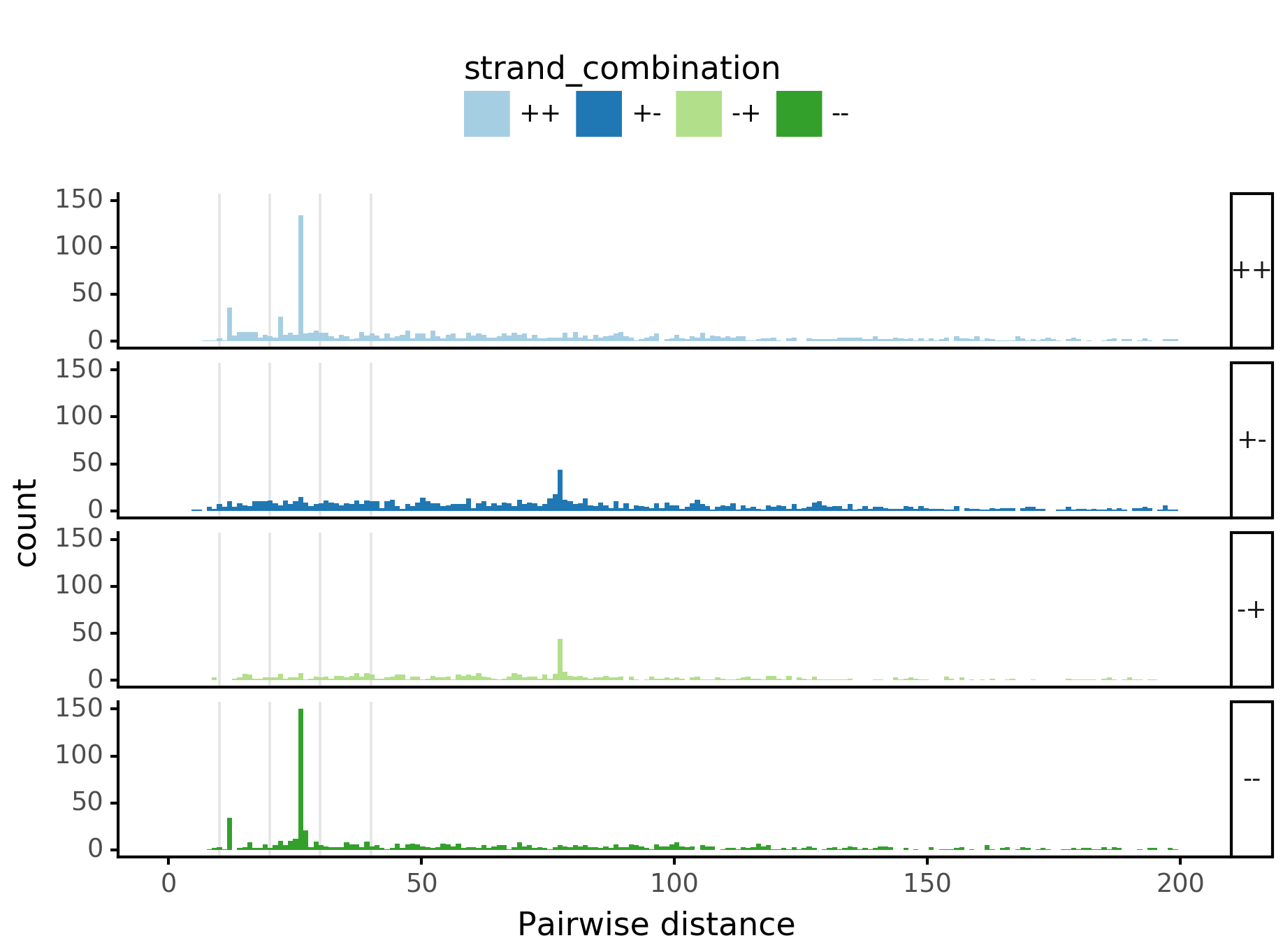
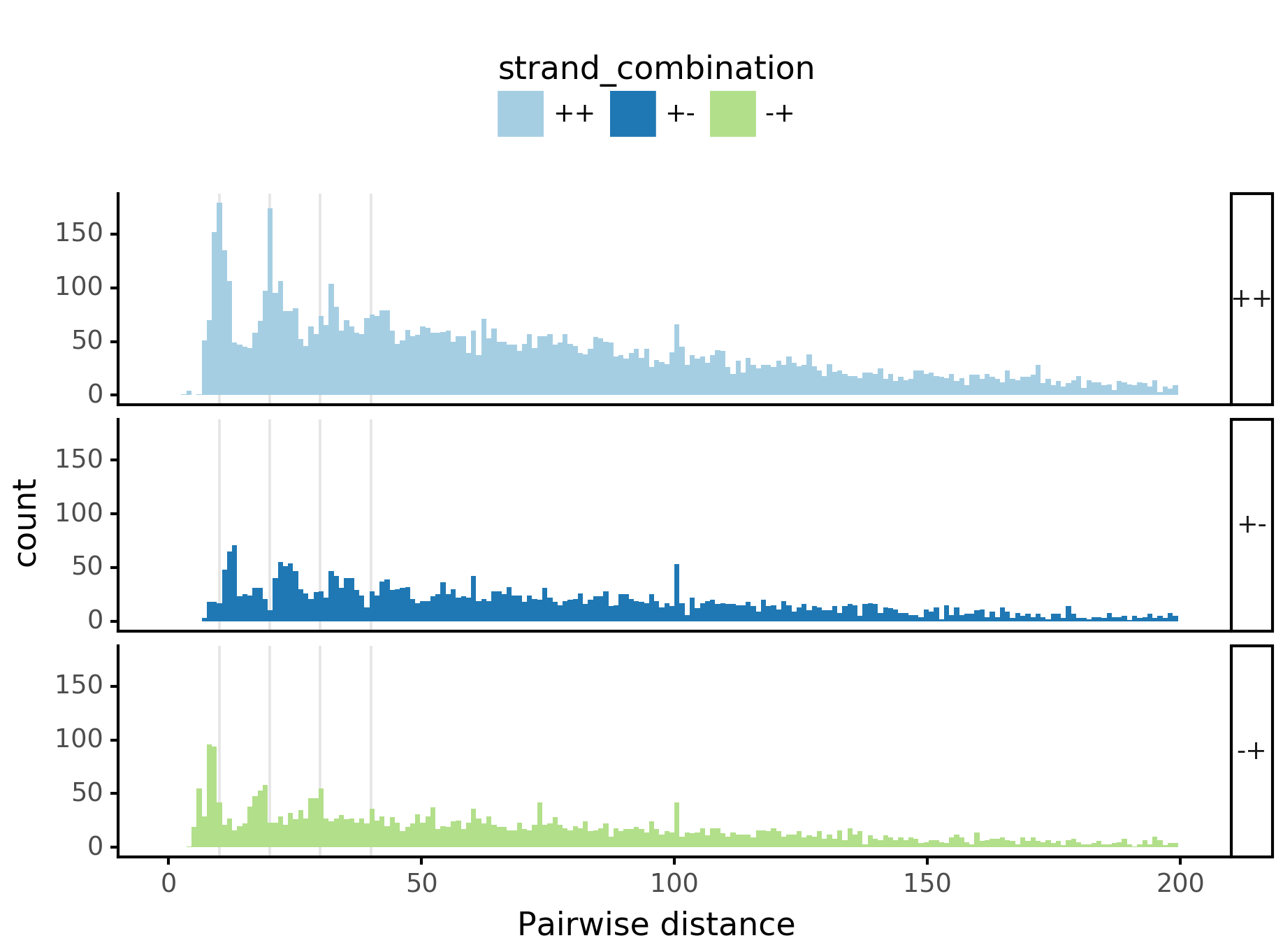
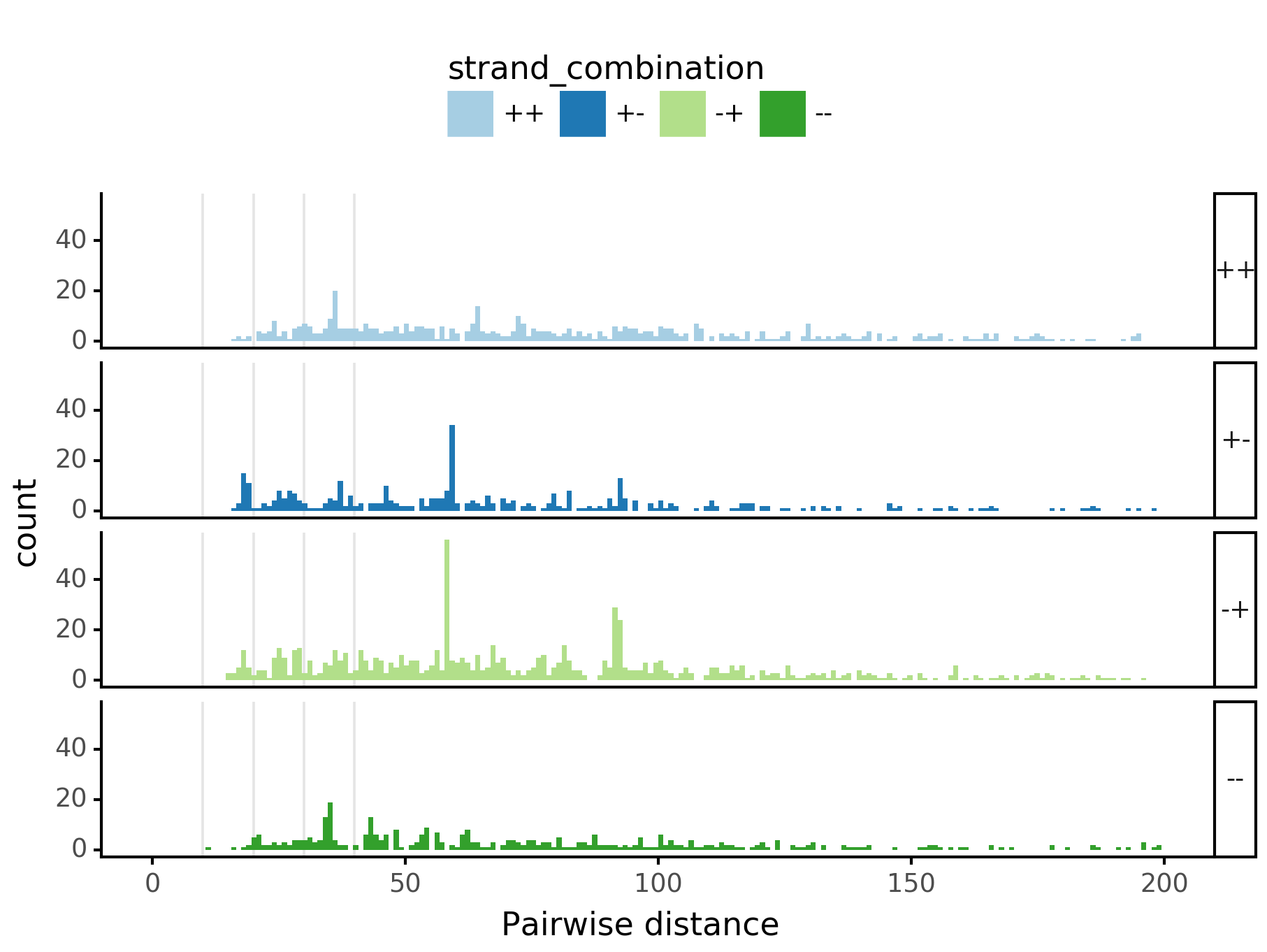
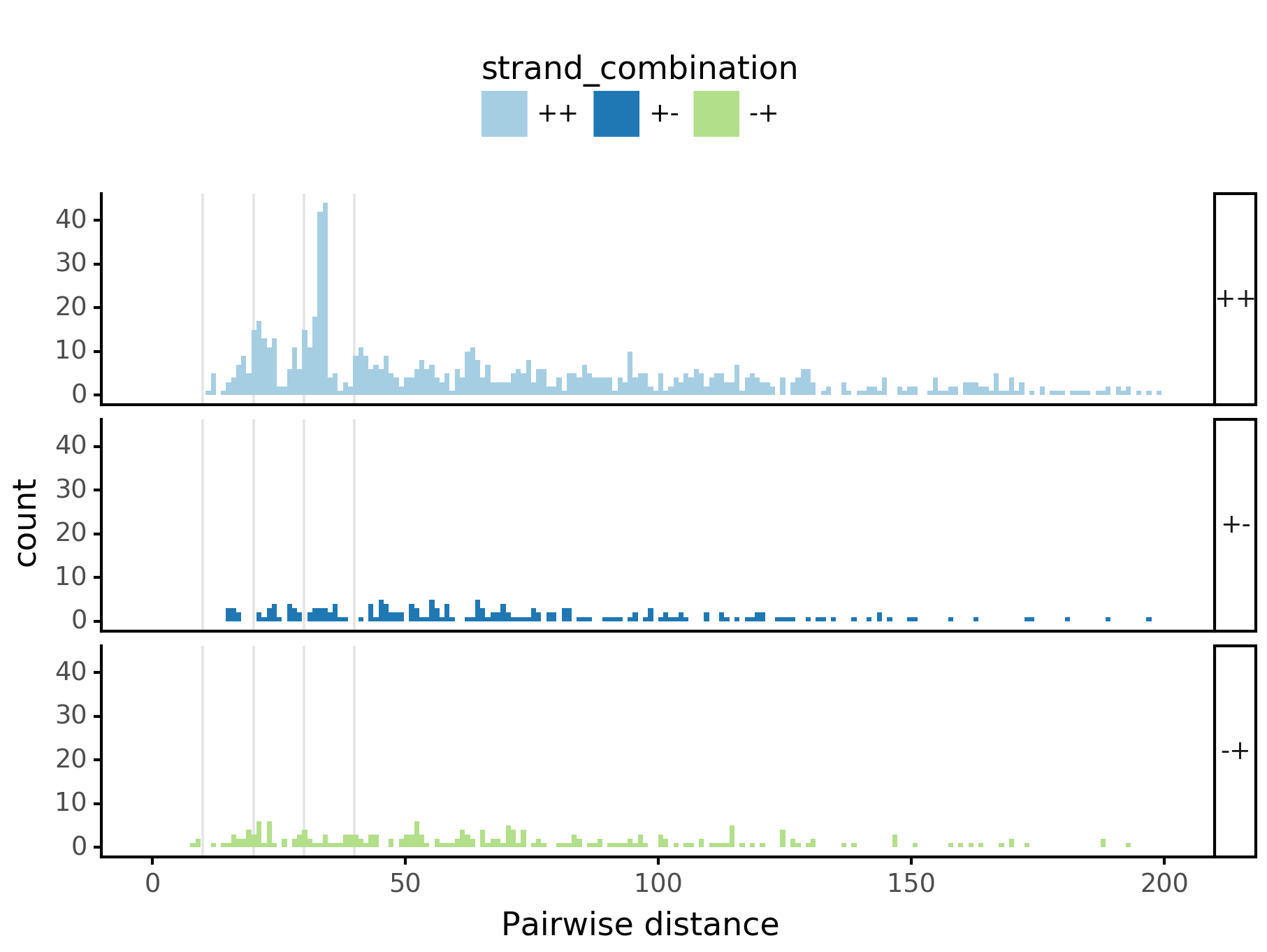
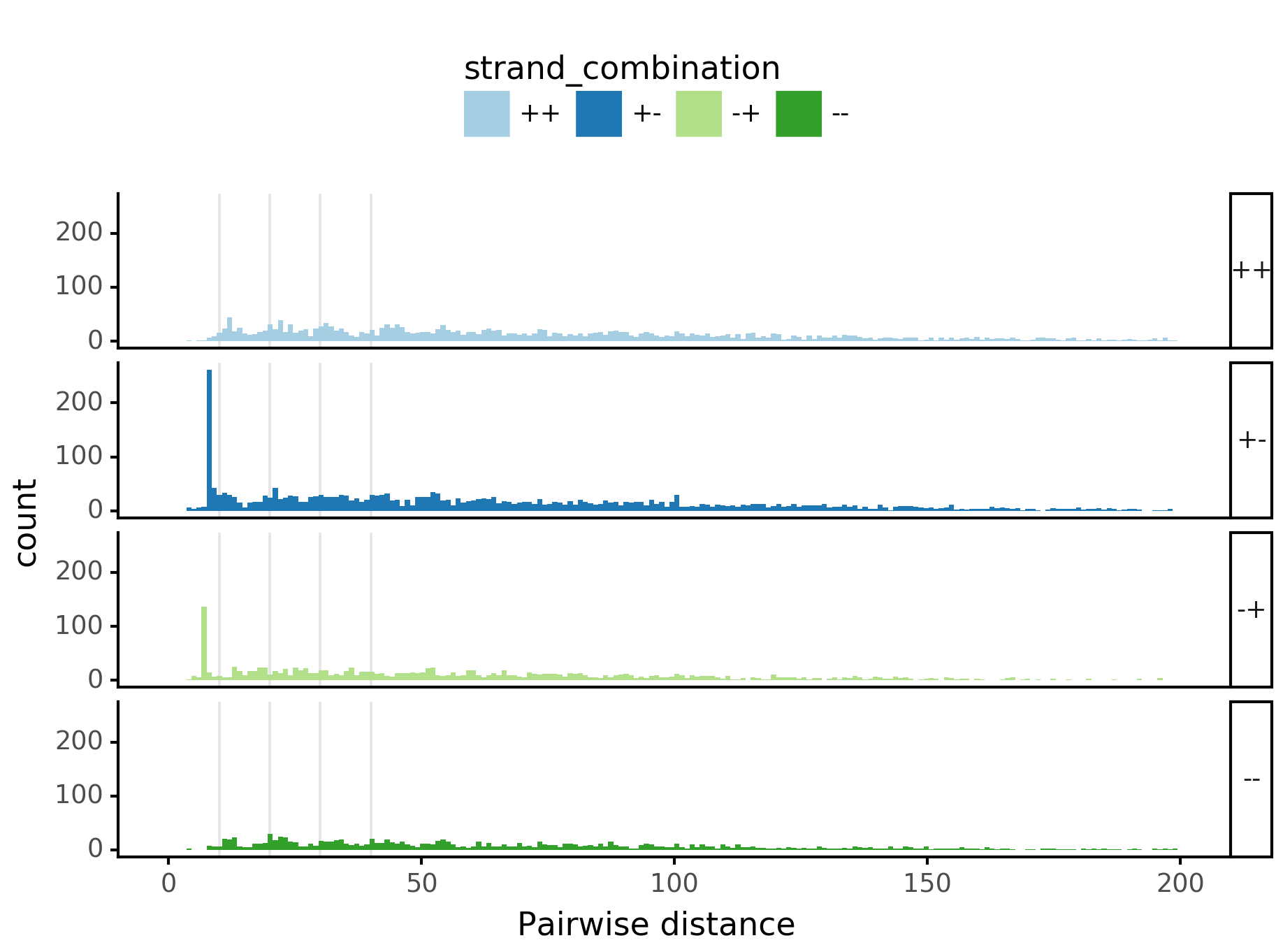
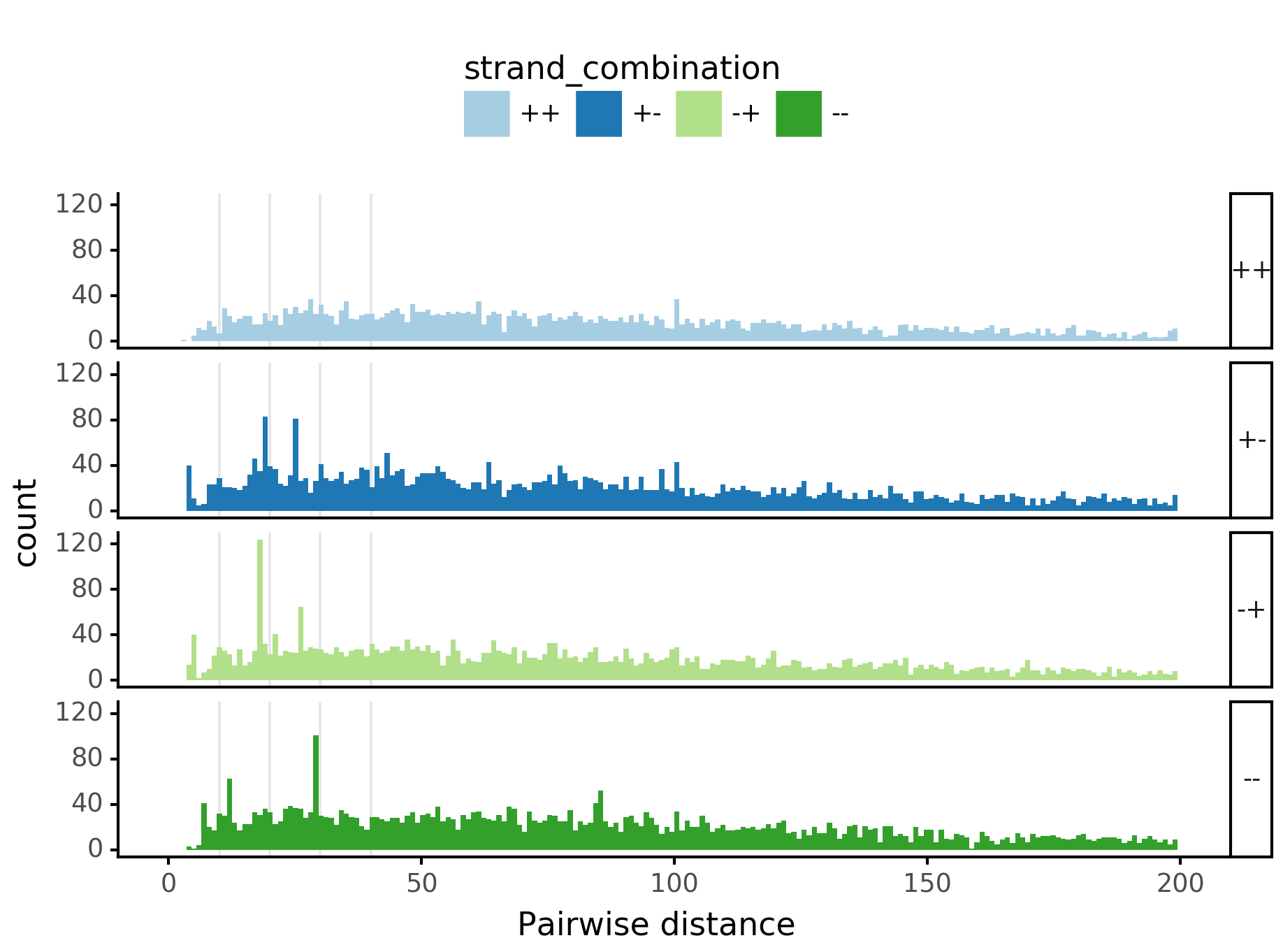
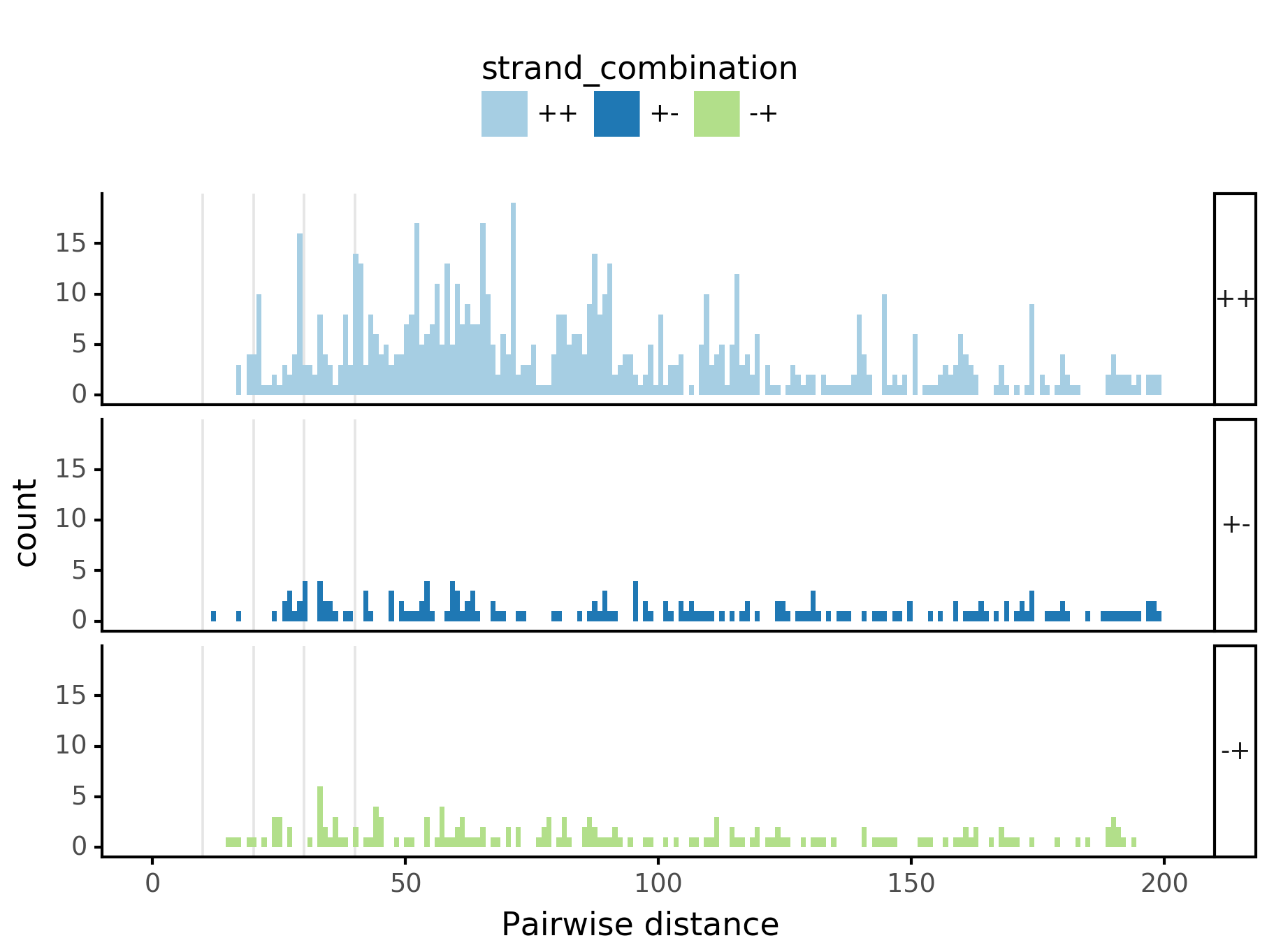
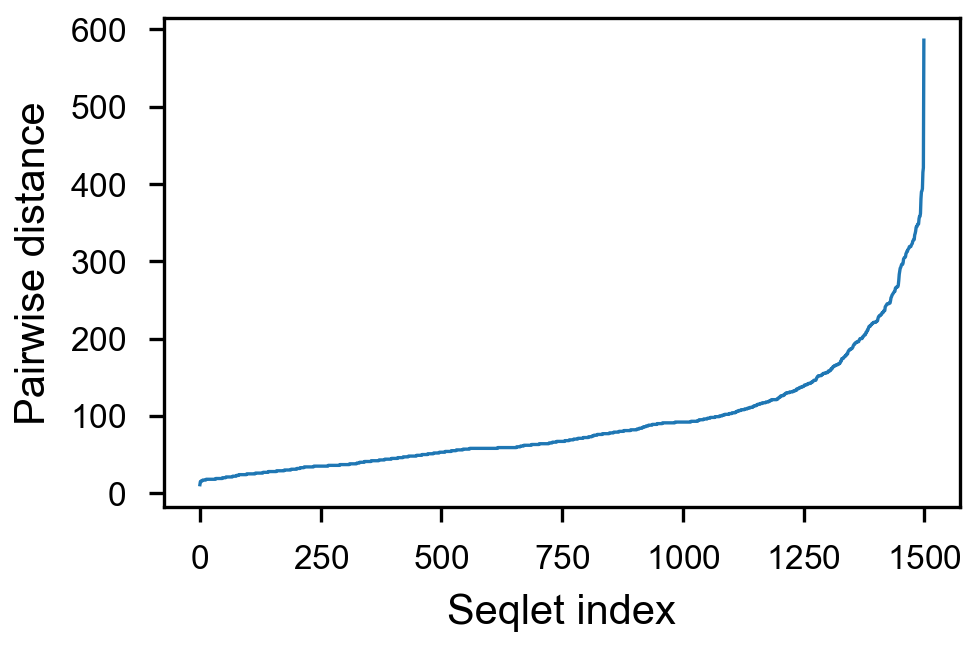
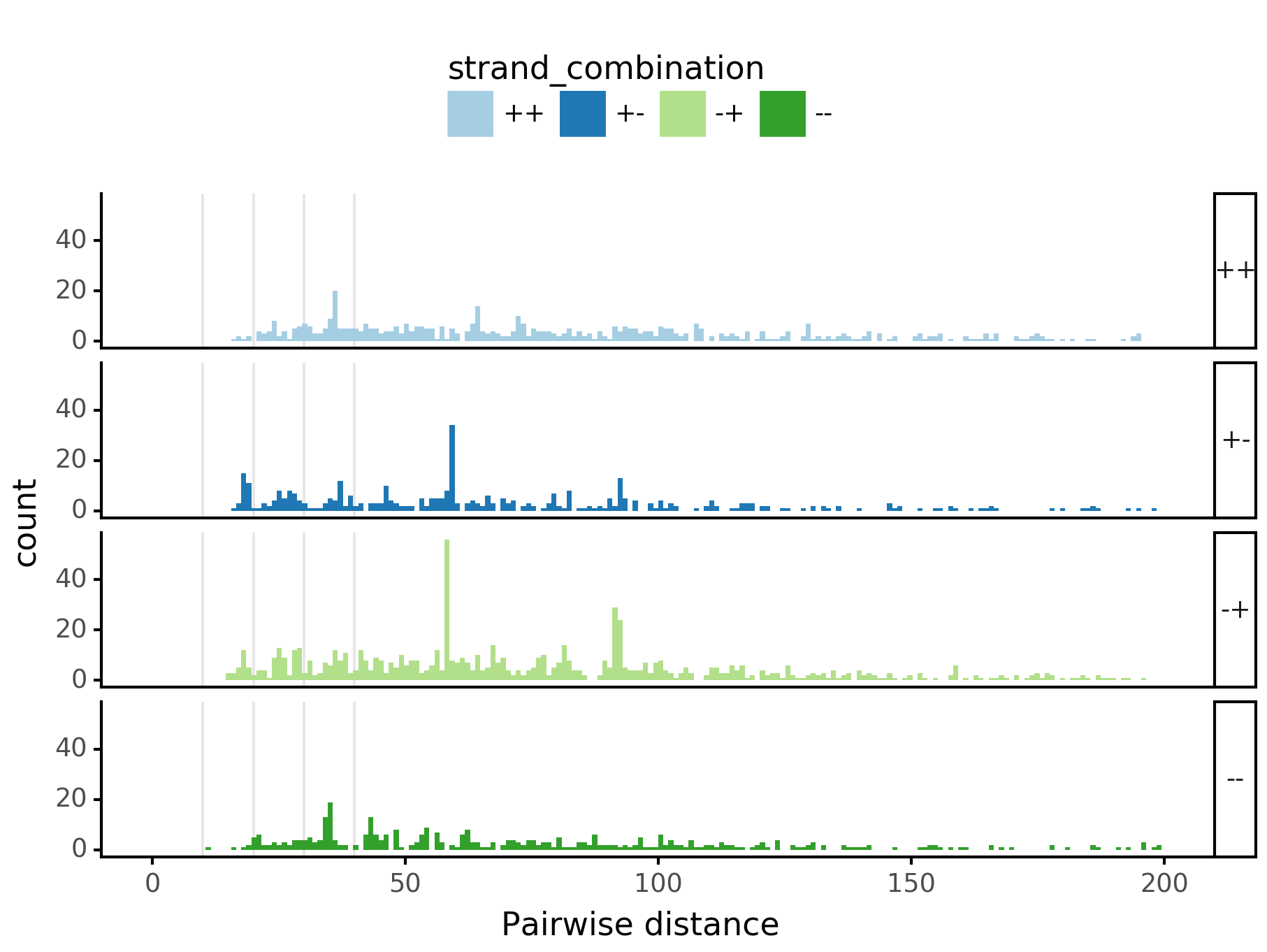
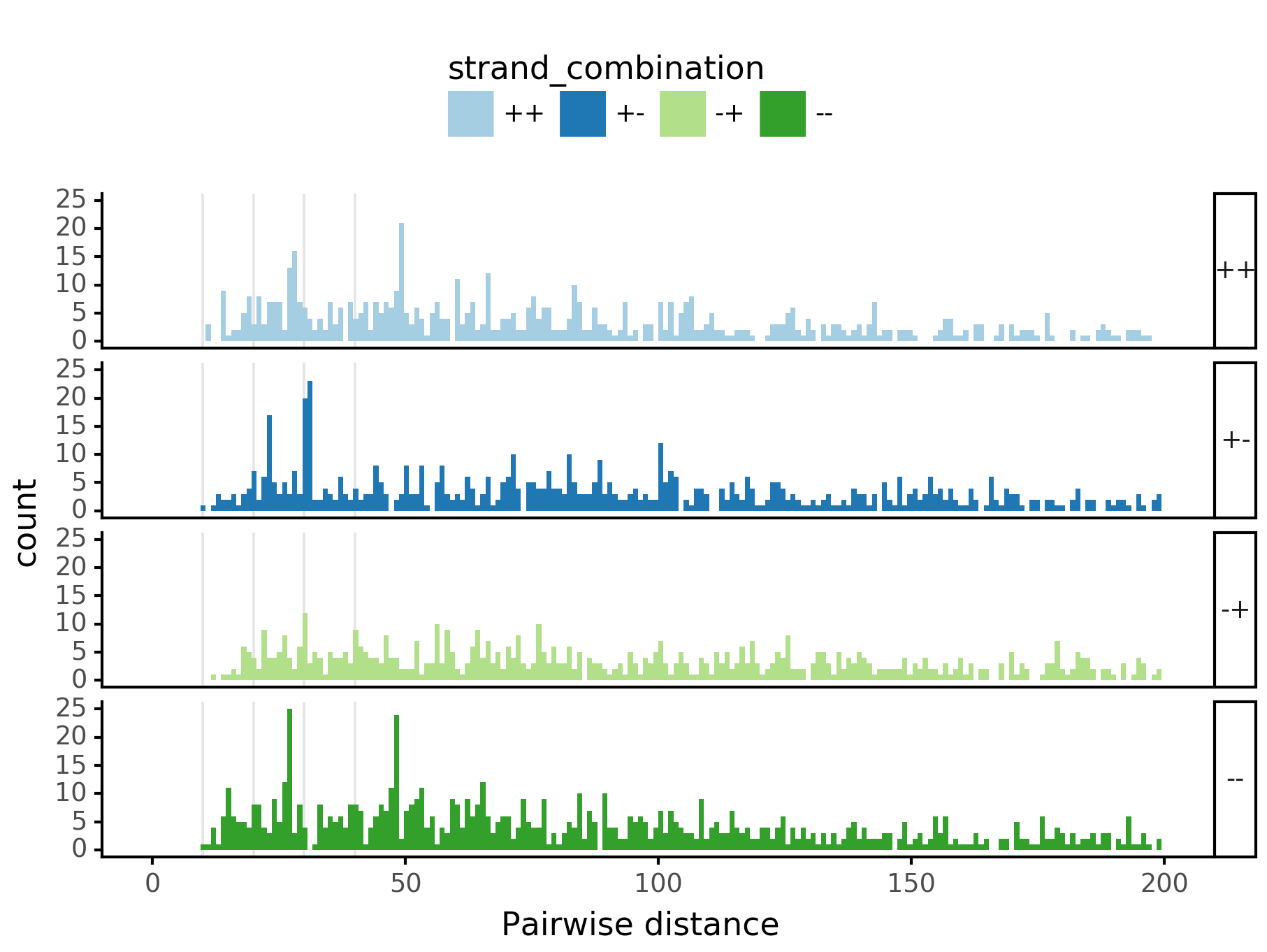
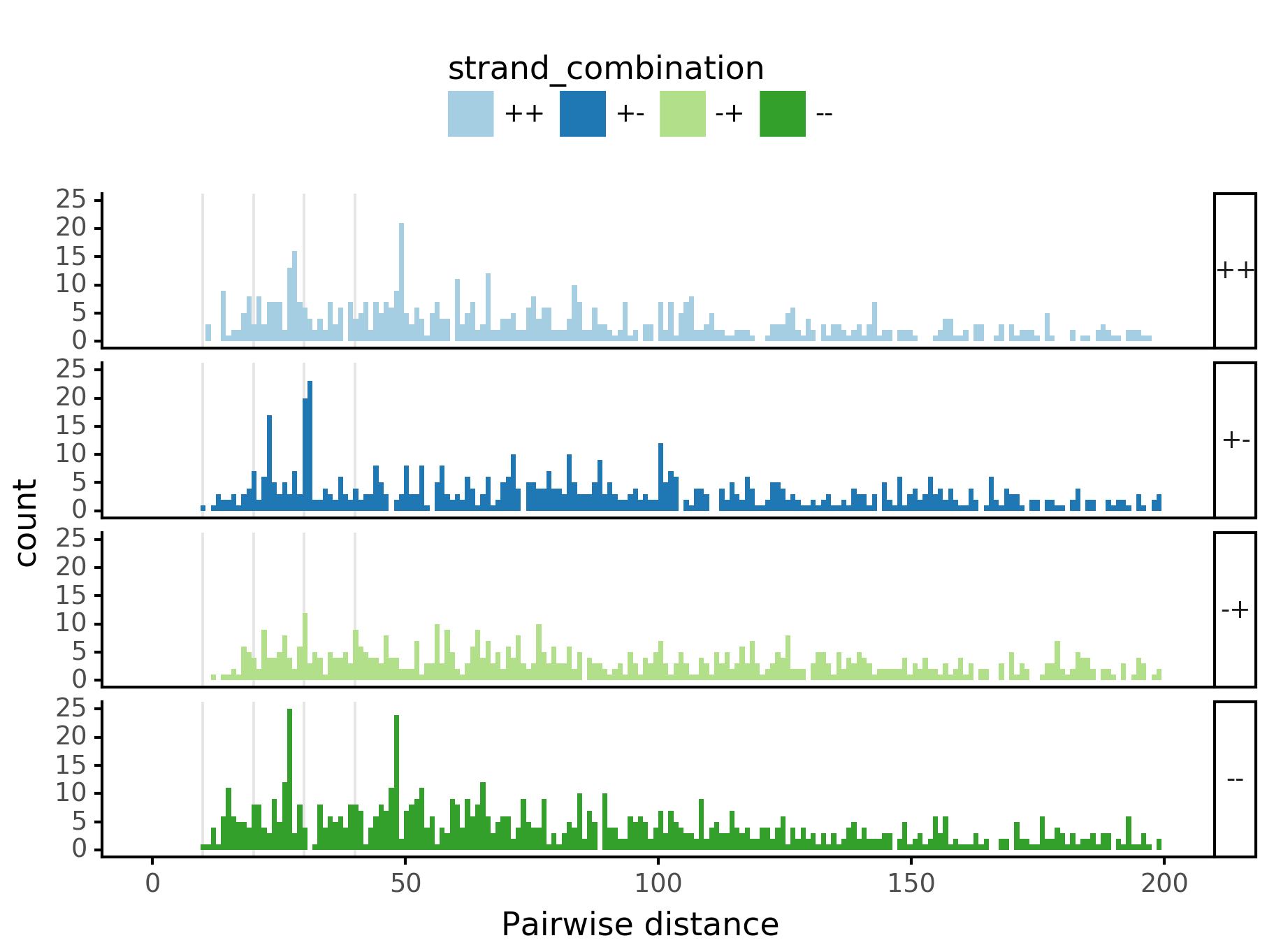
fig = (ggplot(aes(x='center_diff', fill='strand_combination'), dfab[(dfab.center_diff <= max_dist) & (dfab.motif_pair.isin(["Nanog<>Nanog"]))]) +
geom_vline(xintercept=10, alpha=0.1) +
geom_vline(xintercept=20, alpha=0.1) +
geom_vline(xintercept=30, alpha=0.1) +
geom_vline(xintercept=40, alpha=0.1) +
geom_histogram(bins=max_dist) + facet_grid("strand_combination~.") +
theme_classic() +
theme(strip_text = element_text(rotation=0), legend_position='top') +
xlim([0, max_dist]) +
xlab("Pairwise distance") +
ggtitle("Nanog<>Nanog") +
scale_fill_brewer(type='qual', palette=3))
fig
Out[29]:
TODO - export¶
Perturbation analysis¶
In [30]:
output_dir = Path('/srv/www/kundaje/avsec/chipnexus/oct-sox-nanog-klf/models/n_dil_layers=9/modisco/all/deeplift/profile/perturbation/figures.2')
figures_url = Path('/kundaje/avsec/chipnexus/oct-sox-nanog-klf/models/n_dil_layers=9/modisco/all/deeplift/profile/perturbation/figures.2')
In [31]:
!mkdir -p {output_dir}
RowQuantileNormalizer¶
In [323]:
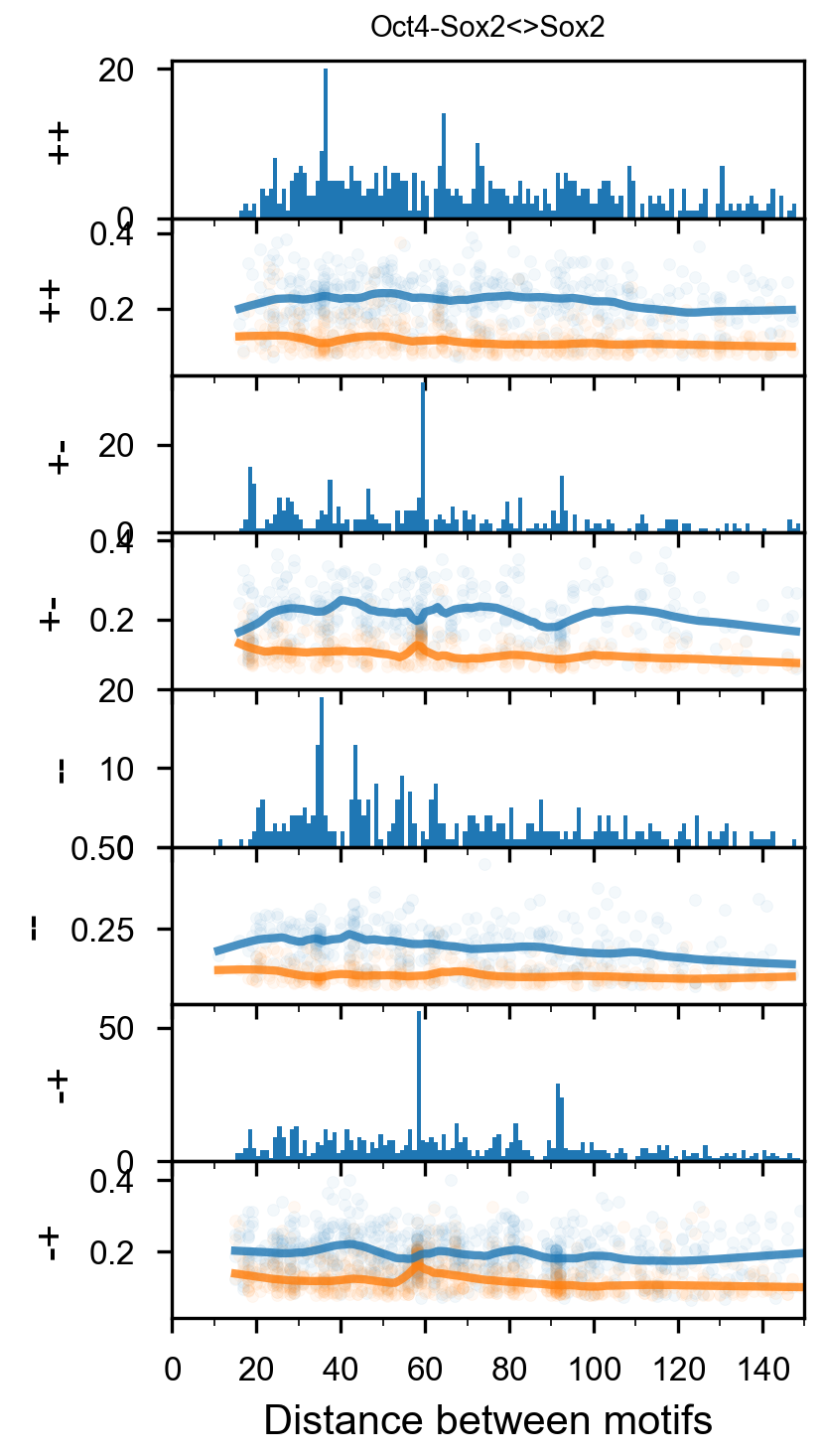
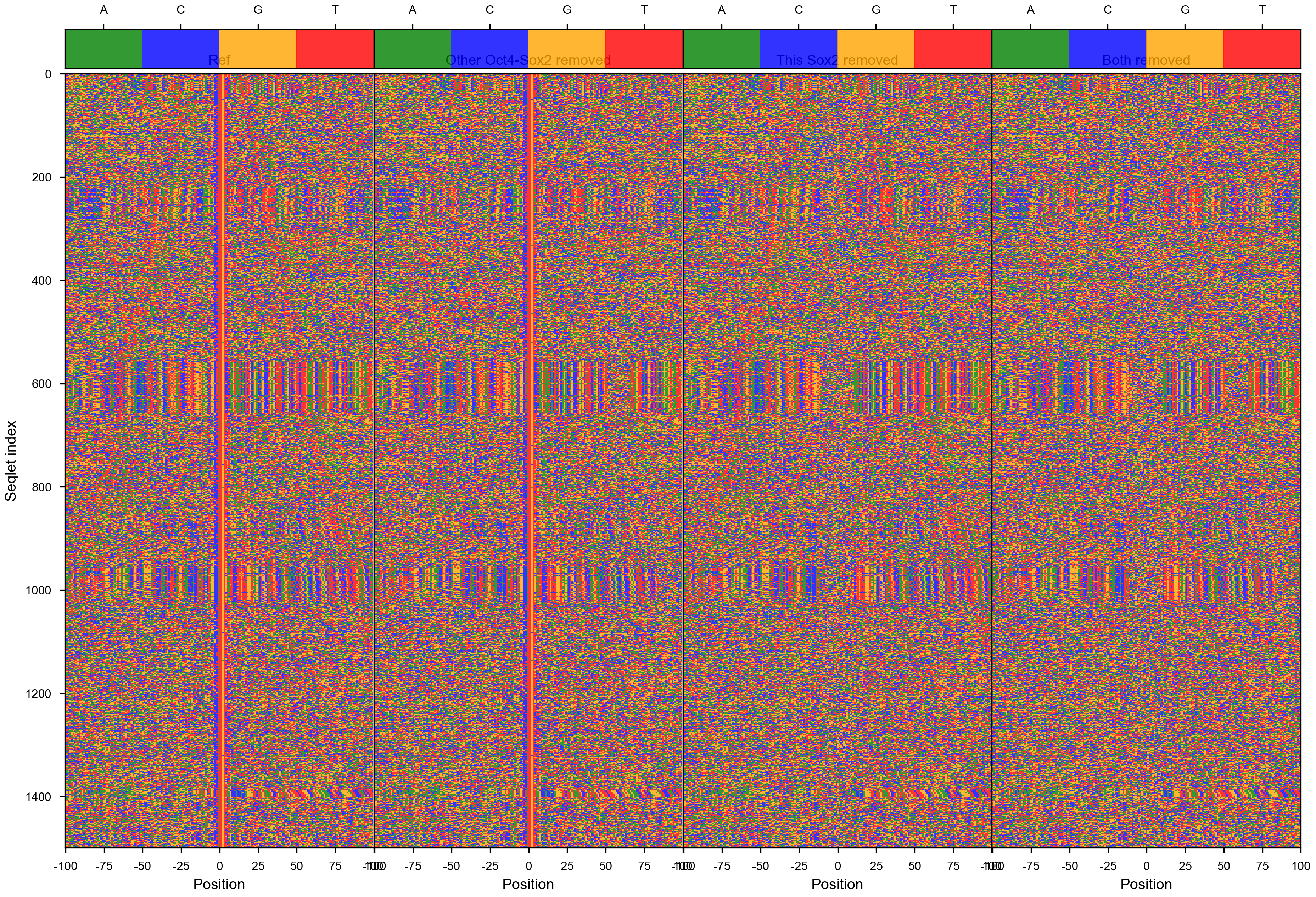
vdom_motif_pair(motif_pair_lpdata, dfab, profile_mapping, figures_dir=str(output_dir / 'RowQuantileNormalizer'), figures_url=str(figures_url / 'RowQuantileNormalizer'), profile_width=200, cache=False, normalizer=RowQuantileNormalizer())
Out[323]:
QuantileTruncateNormalizer¶
In [324]:
vdom_motif_pair(motif_pair_lpdata, dfab, profile_mapping, figures_dir=str(output_dir / 'QuantileTruncateNormalizer'), figures_url=str(figures_url / 'QuantileTruncateNormalizer'), profile_width=200, cache=False, normalizer=QuantileTruncateNormalizer())
Out[324]:
log(1+x) normalization¶
In [325]:
vdom_motif_pair(motif_pair_lpdata, dfab, profile_mapping, figures_dir=str(output_dir / 'lognorm'), figures_url=str(figures_url / 'lognorm'), profile_width=200, cache=False, normalizer=lambda x: np.log(1+x))
Out[325]:
Summarize data¶
TODO¶
- [x] implement different feature functions quantifying the interactions
- [x] given a list of arrays (pred, imp, imp_count), quantify the metric
- [X] combine differnet metrics with each other
- [x] add also the max-count metric
- positions with maximal number of counts (averaged per strands)
- [x] add scatterplots quantifying the interactions
- [ ] re-compute motif_centers and motif_diff using proper rounding schemes
Synergy scores¶
- Wt = total counts in ref
- dA = total counts after removing A
- dB = total counts after removing B
- dAB = total counts after removing both
Then TF-A depends on A with
(Wt-dA)/(Wt - dAB)
and on B with
(Wt-dB)/(Wt - dAB)
In [8]:
def ism_compute_features_tidy(motif_pair_lpdata, tasks,):
out = []
for motif_pair_name, lpdata in tqdm(motif_pair_lpdata.items()):
motif_pair = list(motif_pair_name.split("<>"))
for task in tasks:
dfab_sm = lpdata['dfab'].copy()
dfab_sm['task'] = task
whole = {k: motif_pair_lpdata[motif_pair_name]['x'][k]['whole'][task]
for k in motif_pair_lpdata[motif_pair_name]['x']}
dfab_sm['Wt_obs'] = whole['ref']['obs'].sum(axis=(1,2))
dfab_sm['Wt'] = whole['ref']['pred'].sum(axis=(1,2))
dfab_sm['dA'] = whole['dthis']['pred'].sum(axis=(1,2))
dfab_sm['dB'] = whole['dother']['pred'].sum(axis=(1,2))
dfab_sm['dAB'] = whole['dboth']['pred'].sum(axis=(1,2))
out.append(dfab_sm)
return pd.concat(out, axis=0)
In [9]:
dfabf_ism = ism_compute_features_tidy(motif_pair_lpdata, tasks)
In [ ]:
# dfs.to_csv(dataset_dir / 'dfs.csv.gz', compression='gzip', index=False)
In [10]:
dfabf_ism.head()
Out[10]:
In [134]:
dfc.head()
Out[134]:
In [10]:
dfs = dfabf_ism[['Wt_obs', 'Wt', 'dA', 'dB', 'dAB', 'motif_pair', 'task', 'center_diff', 'strand_combination', 'example_idx']]
dfs.head()
Out[10]:
In [421]:
plt.scatter(np.log10(1+dfs.Wt), np.log10(1+dfs.Wt_obs), s=1, alpha=0.1)
plt.xlabel("Predicted Wt (log10)")
plt.ylabel("Observed Wt (log10)");
In [422]:
np.log10(1+dfs.Wt_obs).plot.hist(300);
plt.xlabel("Observed Wt (log10)");
In [540]:
dfs.head()
Out[540]:
In [11]:
# load activity data
import pybedtools
from pybedtools import BedTool
from basepair.extractors import MultiAssayExtractor
from basepair.data import NumpyDataset
df = pd.read_csv(f"{ddir}/processed/chipnexus/external-data.tsv", sep='\t')
dfs = df[df.assay.isin(['PolII', 'H3K27ac'])]
df_regions = dfi_subset[['example_chrom', 'example_start', 'example_end', 'example_idx']].drop_duplicates()
bt = BedTool.from_dataframe(df_regions)
extractor = MultiAssayExtractor(dfs, None, use_strand=False, n_jobs=10)
regions = extractor.extract(list(bt))
r = NumpyDataset(regions)
dfc = pd.DataFrame(r.aggregate(np.sum, axis=1))
dfc['example_idx'] = df_regions['example_idx'].values
In [ ]:
ls {dataset_dir}
In [14]:
dfs = dfabf_ism[['Wt_obs', 'Wt', 'dA', 'dB', 'dAB', 'motif_pair', 'task', 'center_diff', 'strand_combination', 'example_idx']]
dfs.head()
Out[14]:
In [16]:
dfs = pd.merge(dfs, dfc, on='example_idx', how='left')
In [ ]:
# TODO - merge the two and export
In [20]:
dataset_dir
Out[20]:
In [17]:
dfs.to_csv(dataset_dir / 'pair.total_counts.csv.gz', index=False, compression='gzip') # store the table for Julia
In [637]:
dfs = dfs[dfs.center_diff < 35]
Fraction of positive values¶
In [638]:
terms = ['Wt-dA>0', 'Wt-dB>0', 'Wt-dAB>0', 'dA-dAB>0', 'dB-dAB>0']
c = pd.concat([dfs.groupby(['motif_pair', 'task']).apply(lambda x: x.eval(term).mean()).reset_index().rename(columns={0:'fraction'}).assign(term=term)
for term in terms], axis=0)
c['term'] = pd.Categorical(c['term'], categories=terms)
In [639]:
plotnine.options.figure_size = (10, 4)
(ggplot(aes(y='motif_pair', x='task', fill='fraction'), c) + geom_tile() + theme_classic() + facet_grid(". ~ term") +
theme(legend_position='top') +
scale_fill_gradient2(low='blue', mid='white', high='red', midpoint=.5, limits=[0, 1])
)
Out[639]:
TODO - export¶
Masked scores¶
In [640]:
mask = 'Wt-dAB>0'
pc = 100 # pseudo-counts
terms = ['(2*Wt-dA-dB)/(Wt-dAB)', f'({pc}+2*Wt-dA-dB)/({pc}+Wt-dAB)', f'(2*Wt-dA-dB)/(Wt-dAB)*({mask})']
c = pd.concat([dfs.groupby(['motif_pair', 'task']).apply(lambda x: x.eval(term).mean() if x.eval(mask).mean() > .9 else np.nan).reset_index().rename(columns={0:'average'}).assign(term=term)
for term in terms], axis=0)
c['term'] = pd.Categorical(c['term'], categories=terms)
In [551]:
plotnine.options.figure_size = (9, 4)
(ggplot(aes(y='motif_pair', x='task', fill='average')) +
geom_tile(data=c.dropna()) +
geom_text(label="/", data=c[c.average.isnull()]) +
theme_classic() +
facet_grid(". ~ term") +
theme(legend_position='top') +
scale_fill_gradient2(low='blue', mid='white', high='red', midpoint=1, limits=[0, 2])
)
Out[551]:
Alternative solutions¶
Median. Mask > .9¶
In [664]:
mask = 'Wt-dAB>0'
pc = 100 # pseudo-counts
terms = ['(2*Wt-dA-dB)/(Wt-dAB)']
c = pd.concat([dfs.groupby(['motif_pair', 'task']).apply(lambda x: x.eval(term).median() if x.eval(mask).mean() > .9 else np.nan).reset_index().rename(columns={0:'average'}).assign(term=term)
for term in terms], axis=0)
c['term'] = pd.Categorical(c['term'], categories=terms)
In [665]:
plotnine.options.figure_size = (3, 4)
(ggplot(aes(y='motif_pair', x='task', fill='average')) +
geom_tile(data=c.dropna()) +
geom_text(label="/", data=c[c.average.isnull()]) +
theme_classic() +
facet_grid(". ~ term") +
theme(legend_position='top') +
scale_fill_gradient2(low='blue', mid='white', high='red', midpoint=1, limits=[0, 2])
)
Out[665]:
TODO - export to PDF¶
mask > 0.2¶
In [666]:
mask = 'Wt-dAB>0'
pc = 100 # pseudo-counts
terms = ['(2*Wt-dA-dB)/(Wt-dAB)', f'({pc}+2*Wt-dA-dB)/({pc}+Wt-dAB)', f'(2*Wt-dA-dB)/(Wt-dAB)*({mask})']
c = pd.concat([dfs.groupby(['motif_pair', 'task']).apply(lambda x: x.eval(term).median()).reset_index().rename(columns={0:'average'}).assign(term=term)
for term in terms], axis=0)
c['term'] = pd.Categorical(c['term'], categories=terms)
In [667]:
plotnine.options.figure_size = (9, 4)
(ggplot(aes(y='motif_pair', x='task', fill='average')) +
geom_tile(data=c.dropna()) +
geom_text(label="/", data=c[c.average.isnull()]) +
theme_classic() +
facet_grid(". ~ term") +
theme(legend_position='top') +
scale_fill_gradient2(low='blue', mid='white', high='red', midpoint=1, limits=[0, 2])
)
Out[667]:
Sum up the reads in all regions¶
In [645]:
terms = ['(2*Wt-dA-dB)/(Wt-dAB)', '0.75 + 0.5 *(Wt-dAB > 0)', '0.75 + 0.5 *(Wt-dA > 0)', '0.75 + 0.5 *(Wt-dB > 0)']
features = ['Wt', 'dA', 'dB', 'dAB']
c = pd.concat([dfs.groupby(['motif_pair', 'task'])[features].sum().eval(term).reset_index().rename(columns={0:'average'}).assign(term=term)
for term in terms], axis=0)
c['term'] = pd.Categorical(c['term'], categories=terms)
In [646]:
plotnine.options.figure_size = (9, 4)
(ggplot(aes(y='motif_pair', x='task', fill='average')) +
geom_tile(data=c) +
theme_classic() +
facet_grid(". ~ term") +
theme(legend_position='top') +
scale_fill_gradient2(low='blue', mid='white', high='red', midpoint=1, limits=[0, 2])
)
Out[646]:
In [636]:
plotnine.options.figure_size = (9, 4)
(ggplot(aes(y='motif_pair', x='task', fill='average')) +
geom_tile(data=c) +
theme_classic() +
facet_grid(". ~ term") +
theme(legend_position='top') +
scale_fill_gradient2(low='blue', mid='white', high='red', midpoint=1, limits=[0, 2])
)
Out[636]:
dB -> A effect¶
In [12]:
def profile_count(narrow, wide, profile_slice=None, **kwargs):
if profile_slice is None:
profile_slice = np.arange(wide['ref']['pred'].shape[1])
return (wide['dother']['pred'][:,profile_slice].mean(axis=(1,2)),
wide['ref']['pred'][:,profile_slice].mean(axis=(1,2)))
def log_profile_count(narrow, wide, profile_slice=None, **kwargs):
if profile_slice is None:
profile_slice = np.arange(wide['ref']['pred'].shape[1])
return (np.log(1+wide['dother']['pred'][:,profile_slice].mean(axis=(1,2))),
np.log(1+wide['ref']['pred'][:,profile_slice].mean(axis=(1,2))))
def profile_count_norm(narrow, wide, profile_slice=None, **kwargs):
if profile_slice is None:
profile_slice = np.arange(wide['ref']['pred'].shape[1])
return (wide['dother']['pred'][:,profile_slice].mean(axis=(1,2)) - wide['dboth']['pred'][:,profile_slice].mean(axis=(1,2)),
wide['ref']['pred'][:,profile_slice].mean(axis=(1,2)) - wide['dthis']['pred'][:,profile_slice].mean(axis=(1,2)))
def log_profile_count_norm(narrow, wide, profile_slice=None, **kwargs):
if profile_slice is None:
profile_slice = np.arange(wide['ref']['pred'].shape[1])
return (np.log(1+wide['dother']['pred'][:,profile_slice].mean(axis=(1,2))) - np.log(1+wide['dboth']['pred'][:,profile_slice].mean(axis=(1,2))),
np.log(1+wide['ref']['pred'][:,profile_slice].mean(axis=(1,2))) - np.log(1+wide['dthis']['pred'][:,profile_slice].mean(axis=(1,2))))
def max_profile_count(narrow, wide, max_position=None, **kwargs):
if max_position is None:
max_position = np.argmax(wide['ref']['pred'].mean(axis=0), axis=0)
return (wide['dother']['pred'][:,max_position, [0,1]].mean(axis=-1),
wide['ref']['pred'][:,max_position, [0,1]].mean(axis=-1))
def log_max_profile_count(narrow, wide, max_position=None, **kwargs):
if max_position is None:
max_position = np.argmax(wide['ref']['pred'].mean(axis=0), axis=0)
return (np.log(1+wide['dother']['pred'][:,max_position, [0,1]].mean(axis=-1)),
np.log(1+wide['ref']['pred'][:,max_position, [0,1]].mean(axis=-1)))
def max_profile_count_norm(narrow, wide, max_position=None, **kwargs):
if max_position is None:
max_position = np.argmax(wide['ref']['pred'].mean(axis=0), axis=0)
return (wide['dother']['pred'][:,max_position, [0,1]].mean(axis=-1) - wide['dboth']['pred'][:,max_position, [0,1]].mean(axis=-1),
wide['ref']['pred'][:,max_position, [0,1]].mean(axis=-1) - wide['dthis']['pred'][:,max_position, [0,1]].mean(axis=-1))
def log_max_profile_count_norm(narrow, wide, max_position=None, **kwargs):
if max_position is None:
max_position = np.argmax(wide['ref']['pred'].mean(axis=0), axis=0)
return (np.log(1+wide['dother']['pred'][:,max_position, [0,1]].mean(axis=-1)) - np.log(1+wide['dboth']['pred'][:,max_position, [0,1]].mean(axis=-1)),
np.log(1+wide['ref']['pred'][:,max_position, [0,1]].mean(axis=-1)) - np.log(1+wide['dthis']['pred'][:,max_position, [0,1]].mean(axis=-1)))
def imp_profile(narrow, wide, **kwargs):
return (narrow['dother']['imp']['profile'].max(axis=-1).mean(axis=-1),
narrow['ref']['imp']['profile'].max(axis=-1).mean(axis=-1))
def imp_count(narrow, wide, **kwargs):
return (narrow['dother']['imp']['count'].max(axis=-1).mean(axis=-1),
narrow['ref']['imp']['count'].max(axis=-1).mean(axis=-1))
SCORES = [profile_count,
profile_count_norm,
max_profile_count,
max_profile_count_norm,
imp_profile,
imp_count]
LOGSCORES = [log_profile_count,
log_profile_count_norm,
log_max_profile_count,
log_max_profile_count_norm,
imp_profile,
imp_count]
def compute_features(narrow, wide, SCORES, **kwargs):
for score in SCORES:
return {score.__name__: score(narrow, wide, **kwargs)}
def compute_features_tidy(motif_pair_lpdata, tasks, plot_features=SCORES, pseudo_count_quantile=0, profile_slice=slice(82, 118), variable=None, pval=False):
from basepair.plot.config import get_figsize
nf = len(plot_features)
out = []
for motif_pair_name, lpdata in tqdm(motif_pair_lpdata.items()):
motif_pair = list(motif_pair_name.split("<>"))
dfab_sma = lpdata['dfab'].copy()
# TODO - loop through all possible combinations
xvals = list(motif_pair_lpdata[motif_pair_name]['x'])
for task in tasks:
# compute features
for score in plot_features:
dfab_sm = dfab_sma.copy()
dfab_sm['task'] = task
dfab_sm['score'] = score.__name__
for xy in ['x', 'y']:
wide = {k: motif_pair_lpdata[motif_pair_name][xy][k]['wide'][task]
for k in motif_pair_lpdata[motif_pair_name][xy]}
narrow = {k: motif_pair_lpdata[motif_pair_name][xy][k]['narrow'][task]
for k in motif_pair_lpdata[motif_pair_name][xy]}
dfab_sm[xy + '_alt'], dfab_sm[xy + '_ref'] = score(narrow, wide, profile_slice=profile_slice)
dfab_sm[xy + '_alt_ref'] = dfab_sm[xy + '_alt'] / dfab_sm[xy + '_ref']
pc = np.quantile(dfab_sm[xy + '_ref'], pseudo_count_quantile)
dfab_sm[xy + '_alt_pc'], dfab_sm[xy + '_ref_pc'] = dfab_sm[xy + '_alt'] + pc, dfab_sm[xy + '_ref'] + pc
dfab_sm[xy + '_alt_ref_pc'] = dfab_sm[xy + '_alt_pc'] / dfab_sm[xy + '_ref_pc']
out.append(dfab_sm)
return pd.concat(out, axis=0)
In [18]:
dfabf = compute_features_tidy(motif_pair_lpdata, tasks, SCORES, pseudo_count_quantile=.2, profile_slice=slice(82, 118))
In [58]:
dfabf_log = compute_features_tidy(motif_pair_lpdata, tasks, LOGSCORES, pseudo_count_quantile=.2, profile_slice=slice(82, 118))
Pairwise scatterplots¶
In [60]:
dfabf['center_diff_cat'] = pd.Categorical(pd.cut(dfabf['center_diff'], [0, 35, 70, 150, 1000]))
dfabf_log['center_diff_cat'] = pd.Categorical(pd.cut(dfabf_log['center_diff'], [0, 35, 70, 150, 1000]))
In [44]:
pairs = list(motif_pair_lpdata)
Log scale¶
In [61]:
plotnine.options.figure_size = (15, 10)
# motif_pair_name = 'Sox2<>Nanog'
for motif_pair_name in pairs:
motif_pair = list(motif_pair_name.split("<>"))
fig = (ggplot(aes(x='x_alt_ref_pc', y='y_alt_ref_pc', color='center_diff_cat'), dfabf_log[dfabf_log.motif_pair == motif_pair_name]) +
geom_point(alpha=0.2, size=.05, shape='.', show_legend=False) +
geom_point(alpha=1, size=5, data=dfabf_log[:0], shape='.') + # for the legend
facet_grid("task~score") +
xlim([0, 2]) +
ylim([0, 2]) +
theme_bw() +
xlab(motif_pair[0] + f" (d{motif_pair[1]})") +
ylab(motif_pair[1] + f" (d{motif_pair[0]})") +
ggtitle(motif_pair_name) +
scale_color_brewer(type='qual', palette=2)
)
display(fig)
Normal scale¶
In [46]:
plotnine.options.figure_size = (15, 10)
# motif_pair_name = 'Sox2<>Nanog'
for motif_pair_name in pairs:
motif_pair = list(motif_pair_name.split("<>"))
fig = (ggplot(aes(x='x_alt_ref_pc', y='y_alt_ref_pc', color='center_diff_cat'), dfabf[dfabf.motif_pair == motif_pair_name]) +
geom_point(alpha=0.2, size=.05, shape='.', show_legend=False) +
geom_point(alpha=1, size=5, data=dfabf[:0], shape='.') + # for the legend
facet_grid("task~score") +
xlim([0, 2]) +
ylim([0, 2]) +
theme_bw() +
xlab(motif_pair[0] + f" (d{motif_pair[1]})") +
ylab(motif_pair[1] + f" (d{motif_pair[0]})") +
ggtitle(motif_pair_name) +
scale_color_brewer(type='qual', palette=2)
)
display(fig)
Scratch space¶
In [32]:
from basepair.exp.chipnexus.perturb import plot_scatter
In [33]:
plot_scatter
Out[33]:
In [46]:
# compute the max
In [75]:
from basepair.plot.tracks import plot_tracks
In [453]:
s = slice(82, 118)
In [454]:
plot_tracks( dict(a=motif_pair_lpdata['Oct4-Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, s].mean(axis=0)));
In [73]:
def plot_scatter(x_ref, x_alt, y_ref, y_alt, ax, alpha=.2, s=1, label=None, xl=[0, 2], pval=True):
ax.scatter(x_alt / x_ref,
y_alt / y_ref, alpha=alpha, s=s, label=label)
if pval:
xpval = wilcoxon(x_ref, x_alt).pvalue
ypval = wilcoxon(y_ref, y_alt).pvalue
kwargs = dict(size="small", horizontalalignment='center')
ax.text(1.8, 1, f"{xpval:.2g}", **kwargs)
ax.text(1, 1.8, f"{ypval:.2g}", **kwargs)
alpha = .5
ax.plot(xl, xl, c='grey', alpha=alpha)
ax.axvline(1, c='grey', alpha=alpha)
ax.axhline(1, c='grey', alpha=alpha)
ax.set_xlim(xl)
ax.set_ylim(xl)
In [76]:
from basepair.exp.chipnexus.perturb import plot_scatter
In [92]:
sox2_max_positions = np.argmax(motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'].mean(axis=0), axis=0)
nanog_max_positions = np.argmax(motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'].mean(axis=0), axis=0)
In [ ]:
x = ((np.log(1+motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2))) -
np.log(1+motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'].sum(axis=(1,2)))))
# (motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['wide']['Nanog']['pred'].sum(axis=(1,2))))
y = motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['narrow']['Nanog']['imp']['count'].sum(axis=(1,2))
regression_eval(y, x)
plt.xlabel("ISM (Wr - dNanog)")
plt.ylabel("DeepLIFT count importance");
In [138]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=2+motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Sox2<>Nanog']['x']['dthis']['wide']['Sox2']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1),
x_alt=2+motif_pair_lpdata['Sox2<>Nanog']['x']['dother']['wide']['Sox2']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Sox2<>Nanog']['x']['dboth']['wide']['Sox2']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1),
y_ref=2+motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Sox2<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_alt=2+motif_pair_lpdata['Sox2<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Sox2<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
ax=ax, xl=[0, 2])
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[138]:
In [118]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=2+motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1),
x_alt=2+motif_pair_lpdata['Sox2<>Nanog']['x']['dother']['wide']['Sox2']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1),
y_ref=2+motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_alt=2+motif_pair_lpdata['Sox2<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
ax=ax, xl=[0, 2])
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[118]:
In [117]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=1+motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
x_alt=1+motif_pair_lpdata['Sox2<>Nanog']['x']['dother']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
y_ref=1+motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
y_alt=1+motif_pair_lpdata['Sox2<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
ax=ax, xl=[0, 2]
)
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[117]:
In [134]:
s = slice(85, 118)
In [119]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=1+motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['x']['dthis']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
x_alt=1+motif_pair_lpdata['Sox2<>Nanog']['x']['dother']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['x']['dboth']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
y_ref=1+motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
y_alt=1+motif_pair_lpdata['Sox2<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
ax=ax, xl=[0, 2]
)
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[119]:
In [129]:
s = slice(65, 135)
In [130]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['x']['dthis']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
x_alt=motif_pair_lpdata['Sox2<>Nanog']['x']['dother']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['x']['dboth']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
y_ref=motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
y_alt=motif_pair_lpdata['Sox2<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
ax=ax, xl=[-1, 3]
)
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[130]:
In [122]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=1+motif_pair_lpdata['Sox2<>Nanog']['x']['ref']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['x']['dthis']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
x_alt=1+motif_pair_lpdata['Sox2<>Nanog']['x']['dother']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['x']['dboth']['wide']['Sox2']['pred'][:, s].mean(axis=(1,2)),
y_ref=1+motif_pair_lpdata['Sox2<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
y_alt=1+motif_pair_lpdata['Sox2<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Sox2<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
ax=ax, xl=[0, 2]
)
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[122]:
In [148]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=2+motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
x_alt=2+motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
y_ref=2+motif_pair_lpdata['Nanog<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
y_alt=2+motif_pair_lpdata['Nanog<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, s].mean(axis=(1,2)),
ax=ax, xl=[0, 2]
)
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[148]:
In [137]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=2+motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1),
x_alt=2+motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'][:, sox2_max_positions,[0, 1]].mean(axis=-1),
y_ref=2+motif_pair_lpdata['Nanog<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_alt=2+motif_pair_lpdata['Nanog<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
ax=ax, xl=[0, 2])
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[137]:
In [145]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=2+motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
x_alt=2+motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_ref=2+motif_pair_lpdata['Nanog<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_alt=2+motif_pair_lpdata['Nanog<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
ax=ax, xl=[0, 2])
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[145]:
In [145]:
fig,ax = plt.subplots(figsize=get_figsize(.5, 1))
plot_scatter(x_ref=2+motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
x_alt=2+motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_ref=2+motif_pair_lpdata['Nanog<>Nanog']['y']['ref']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['y']['dthis']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
y_alt=2+motif_pair_lpdata['Nanog<>Nanog']['y']['dother']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1) - motif_pair_lpdata['Nanog<>Nanog']['y']['dboth']['wide']['Nanog']['pred'][:, nanog_max_positions,[0, 1]].mean(axis=-1),
ax=ax, xl=[0, 2])
plt.xlabel("Sox2 (dNanog)")
plt.ylabel("Nanog (dSox2)")
Out[145]:
TODO¶
- correlate ISM scores and deeplift scores
In [193]:
np.mean(x > 1)
Out[193]:
In [194]:
np.mean(x)
Out[194]:
In [ ]:
plt.scatter(motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'].sum(axis=(1,2)),
motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'].sum(axis=(1,2)))
In [192]:
x = ((2*motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2)) -
motif_pair_lpdata['Nanog<>Nanog']['x']['dother']['wide']['Nanog']['pred'].sum(axis=(1,2)) -
motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'].sum(axis=(1,2))) /
(motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['wide']['Nanog']['pred'].sum(axis=(1,2))))
plt.hist(x[(x>-1) & (x<10)], 50);
plt.xlabel("Synergy score");
In [195]:
y = motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['narrow']['Nanog']['imp']['profile']
In [240]:
motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['narrow']['Nanog']['imp']['count'][0]
Out[240]:
In [225]:
plt.hist(motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['narrow']['Nanog']['imp']['count'].sum(axis=(1,2)))
Out[225]:
In [313]:
from basepair.plot.evaluate import regression_eval
In [179]:
x = ((motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2)) -
motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'].sum(axis=(1,2))))
y = (motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2)) - motif_pair_lpdata['Nanog<>Nanog']['x']['dboth']['wide']['Nanog']['pred'].sum(axis=(1,2)))
plt.scatter(x,y, alpha=0.1);
# plt.xlabel("Single TF affinity");
In [178]:
x = ((motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Nanog']['pred'].sum(axis=(1,2))) - motif_pair_lpdata['Nanog<>Nanog']['x']['dthis']['wide']['Nanog']['pred'].sum(axis=(1,2)))
plt.hist(x[x< 100], 50);
plt.xlabel("Single TF affinity");
In [163]:
motif_pair_lpdata['Nanog<>Nanog']['x']['ref']['wide']['Klf4']['obs'].shape
Out[163]:
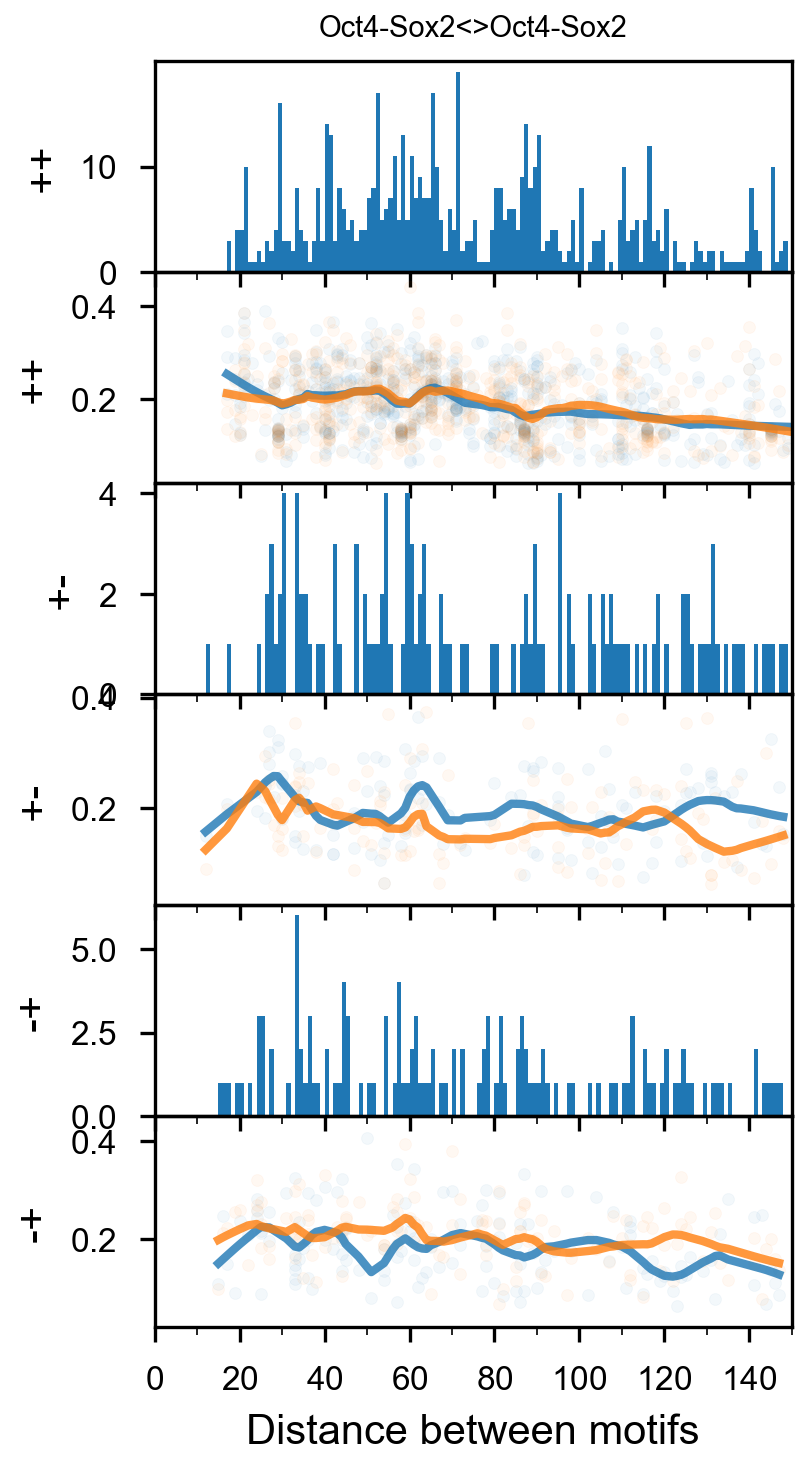

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

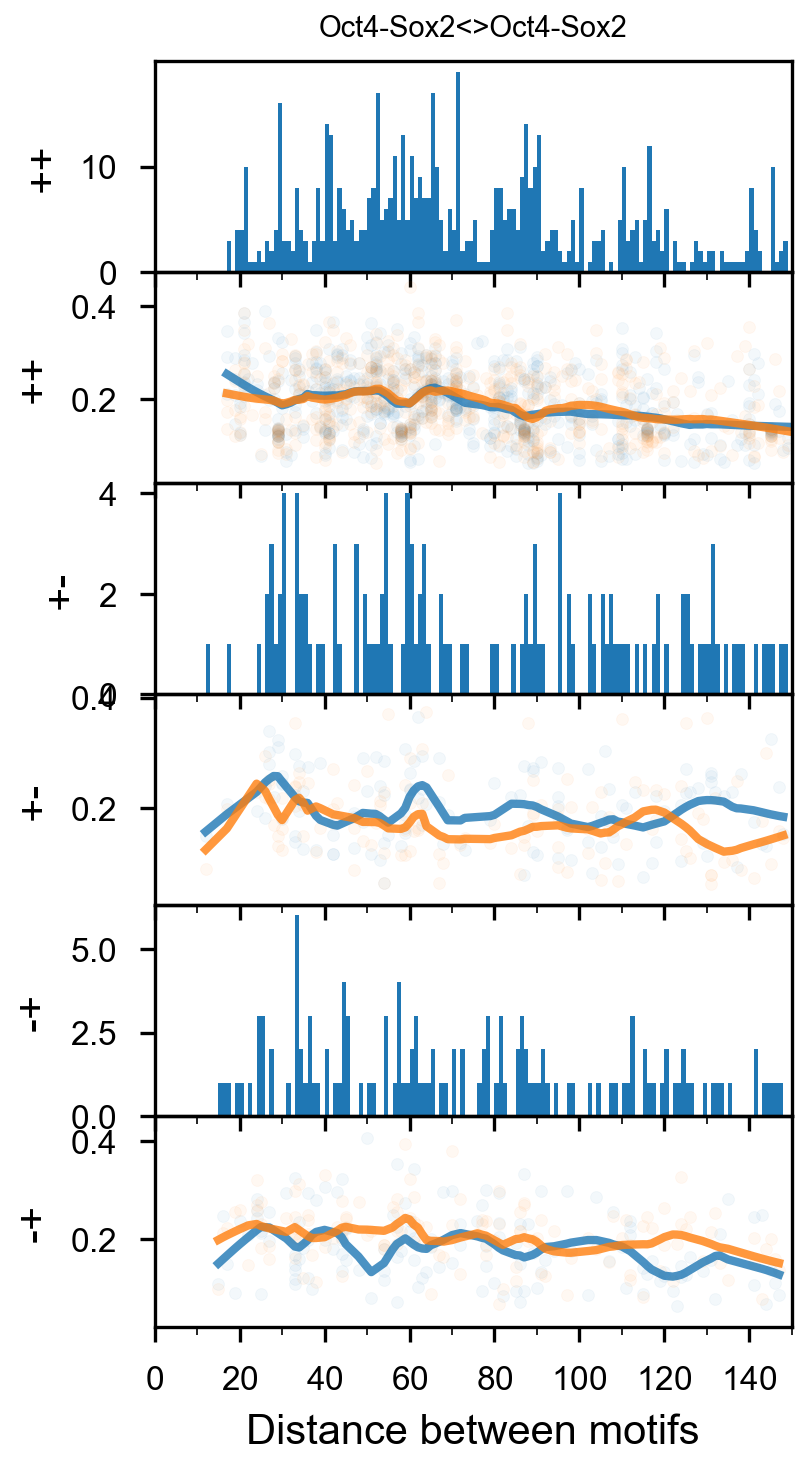
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
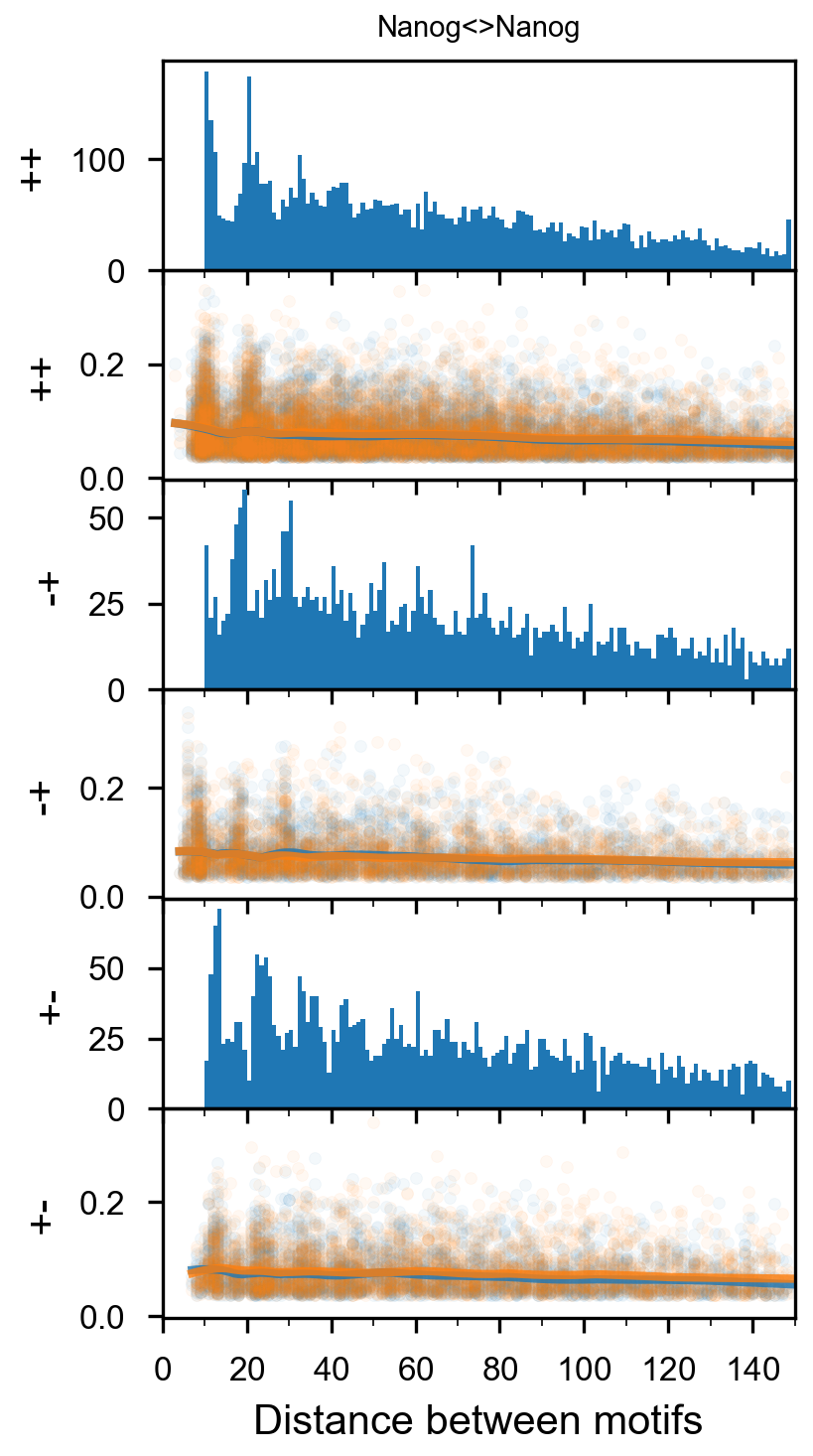
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)




.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)




.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
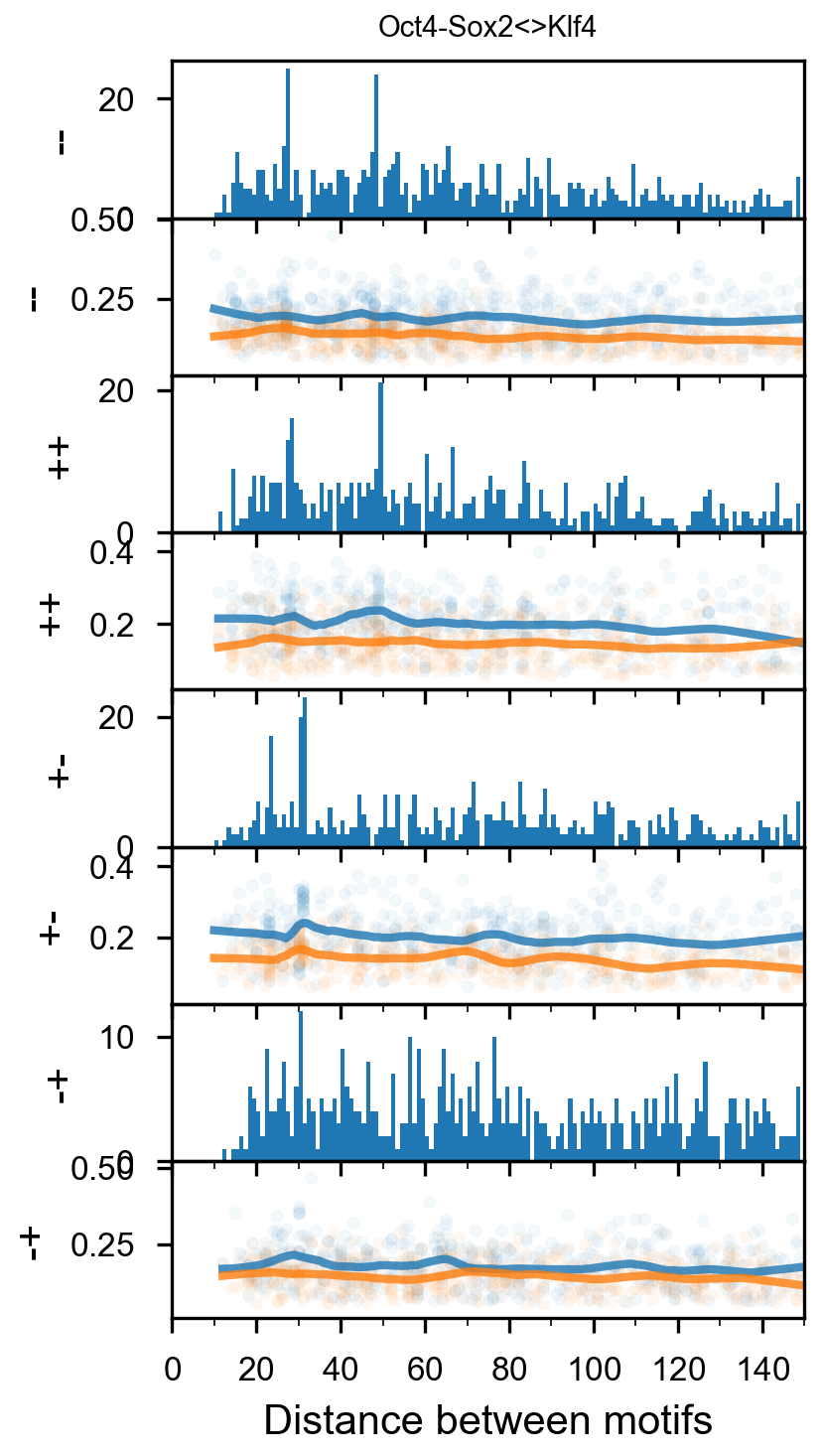

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)


.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
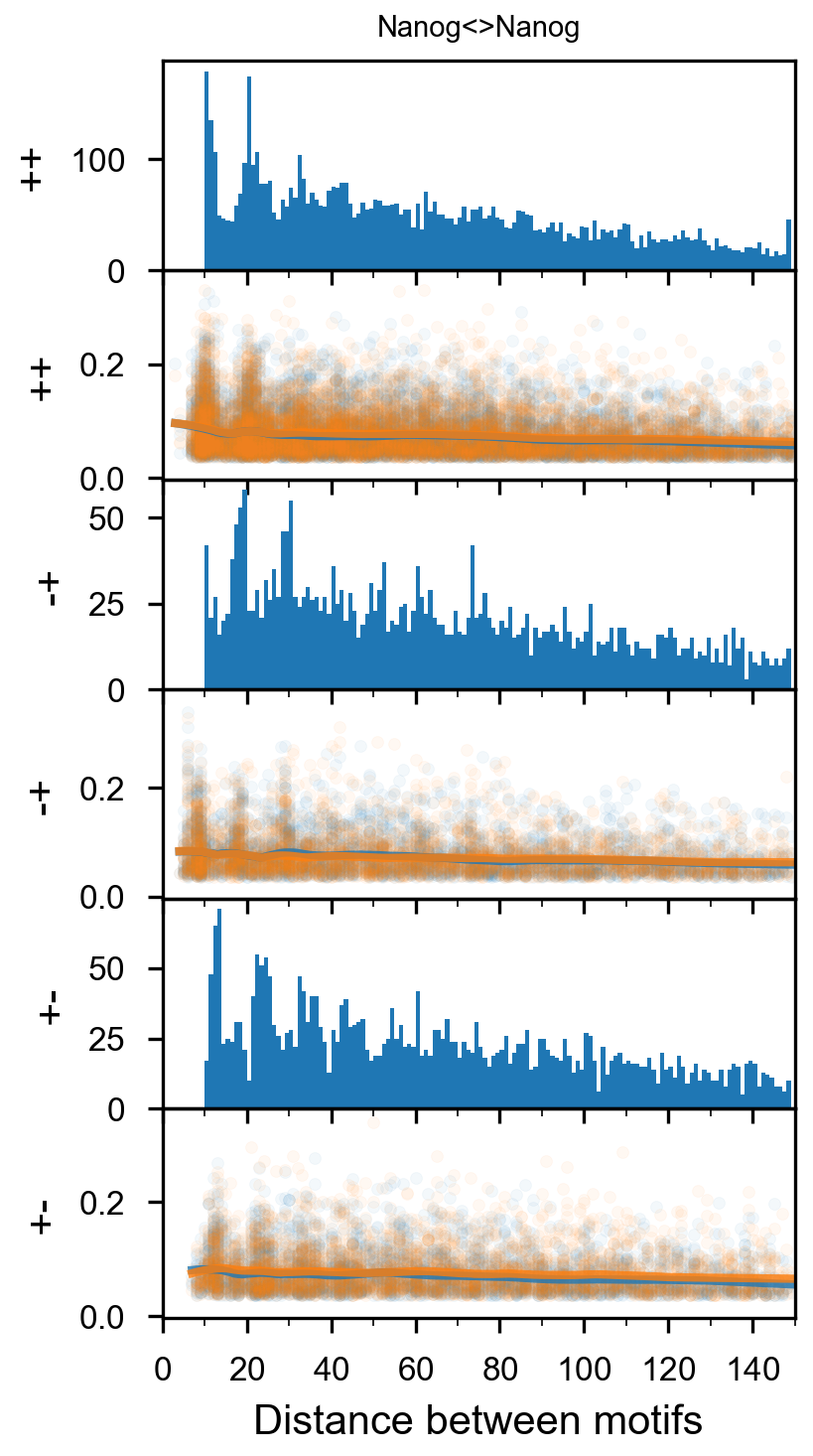
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)



.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)
.png)