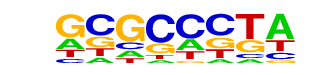
| p-value: | 1e-30 |
| log p-value: | -6.981e+01 |
| Information Content per bp: | 1.405 |
| Number of Target Sequences with motif | 881.0 |
| Percentage of Target Sequences with motif | 23.39% |
| Number of Background Sequences with motif | 3267.6 |
| Percentage of Background Sequences with motif | 16.11% |
| Average Position of motif in Targets | 434.1 +/- 362.7bp |
| Average Position of motif in Background | 292.5 +/- 218.6bp |
| Strand Bias (log2 ratio + to - strand density) | -0.2 |
| Multiplicity (# of sites on avg that occur together) | 1.21 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
DPL-1(E2F)/cElegans-Adult-ChIP-Seq(modEncode)/Homer
| Match Rank: | 1 |
| Score: | 0.72
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TAGGGCGC
TAGCGCGC |
|


|
|
PB0009.1_E2F3_1/Jaspar
| Match Rank: | 2 |
| Score: | 0.72
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --TAGGGCGC-----
ATAAGGGCGCGCGAT |
|


|
|
TRB2/MA1073.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.71
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TAGGGCGC
TTAGGGCA- |
|


|
|
PB0008.1_E2F2_1/Jaspar
| Match Rank: | 4 |
| Score: | 0.70
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --TAGGGCGC-----
ATAAAGGCGCGCGAT |
|


|
|
TBF1/MA0403.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.69
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TAGGGCGC
NTAGGGTT- |
|


|
|
POL006.1_BREu/Jaspar
| Match Rank: | 6 |
| Score: | 0.67
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | TAGGGCGC---
---GGCGCGCT |
|


|
|
STP2/MA0395.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.67
| | Offset: | -6
| | Orientation: | reverse strand |
| Alignment: | ------TAGGGCGC------
NNGNNGTGCGGCGCCGNTNN |
|


|
|
brk/dmmpmm(Bigfoot)/fly
| Match Rank: | 8 |
| Score: | 0.66
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | TAGGGCGC-
--CGGCGCC |
|


|
|
PB0143.1_Klf7_2/Jaspar
| Match Rank: | 9 |
| Score: | 0.65
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---TAGGGCGC------
NNNTNGGGCGTATNNTN |
|


|
|
PB0133.1_Hic1_2/Jaspar
| Match Rank: | 10 |
| Score: | 0.64
| | Offset: | -4
| | Orientation: | reverse strand |
| Alignment: | ----TAGGGCGC----
NNNNTTGGGCACNNCN |
|


|
|