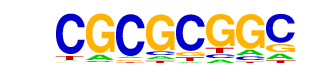
| p-value: | 1e-115 |
| log p-value: | -2.658e+02 |
| Information Content per bp: | 1.599 |
| Number of Target Sequences with motif | 1504.0 |
| Percentage of Target Sequences with motif | 36.44% |
| Number of Background Sequences with motif | 4377.7 |
| Percentage of Background Sequences with motif | 20.91% |
| Average Position of motif in Targets | 428.8 +/- 323.9bp |
| Average Position of motif in Background | 276.6 +/- 244.5bp |
| Strand Bias (log2 ratio + to - strand density) | 0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.69 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
RSC3/MA0374.1/Jaspar
| Match Rank: | 1 |
| Score: | 0.87
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCGCGGC
CGCGCGG- |
|

|
|
RSC30/MA0375.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.87
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CGCGCGGC
CGCGCGCG |
|

|
|
SUT1(MacIsaac)/Yeast
| Match Rank: | 3 |
| Score: | 0.78
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCGCGGC
CGCGGGG- |
|

|
|
SUT1/MA0399.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.78
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGCGCGGC
CGCGGGG- |
|

|
|
DPL-1(E2F)/cElegans-Adult-ChIP-Seq(modEncode)/Homer
| Match Rank: | 5 |
| Score: | 0.77
| | Offset: | -3
| | Orientation: | forward strand |
| Alignment: | ---CGCGCGGC
TAGCGCGC--- |
|


|
|
Tcfl5/MA0632.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.76
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --CGCGCGGC
GGCACGTGCC |
|


|
|
MBP1/MA0329.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.73
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --CGCGCGGC
NNCGCGT--- |
|


|
|
PB0095.1_Zfp161_1/Jaspar
| Match Rank: | 8 |
| Score: | 0.73
| | Offset: | -7
| | Orientation: | reverse strand |
| Alignment: | -------CGCGCGGC-
NCANGCGCGCGCGCCA |
|


|
|
SUT1?/SacCer-Promoters/Homer
| Match Rank: | 9 |
| Score: | 0.72
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | CGCGCGGC-
-GCGCGGGG |
|

|
|
FHY3(FAR1)/Arabidopsis-FHY3-ChIP-Seq(GSE30711)/Homer
| Match Rank: | 10 |
| Score: | 0.71
| | Offset: | -4
| | Orientation: | forward strand |
| Alignment: | ----CGCGCGGC
HHCACGCGCBTN |
|

|
|