| Rank | Motif | Name | P-value | log P-pvalue | q-value (Benjamini) | # Target Sequences with Motif | % of Targets Sequences with Motif | # Background Sequences with Motif | % of Background Sequences with Motif | Motif File |
PDF |
| 1 |  | ABF1/SacCer-Promoters/Homer | 1e-219 | -5.065e+02 | 0.0000 | 1568.0 | 35.13% | 3133.5 | 15.73% | motif file (matrix) |
pdf |
| 2 |  | REB1/SacCer-Promoters/Homer | 1e-189 | -4.359e+02 | 0.0000 | 985.0 | 22.07% | 1571.1 | 7.89% | motif file (matrix) |
pdf |
| 3 |  | RLR1?/SacCer-Promoters/Homer | 1e-65 | -1.511e+02 | 0.0000 | 2100.0 | 47.04% | 6883.7 | 34.55% | motif file (matrix) |
pdf |
| 4 |  | c-Myc(bHLH)/LNCAP-cMyc-ChIP-Seq(Unpublished)/Homer | 1e-58 | -1.353e+02 | 0.0000 | 626.0 | 14.02% | 1402.3 | 7.04% | motif file (matrix) |
pdf |
| 5 |  | SeqBias: CG-repeat | 1e-46 | -1.062e+02 | 0.0000 | 812.0 | 18.19% | 2179.9 | 10.94% | motif file (matrix) |
pdf |
| 6 |  | n-Myc(bHLH)/mES-nMyc-ChIP-Seq(GSE11431)/Homer | 1e-45 | -1.041e+02 | 0.0000 | 708.0 | 15.86% | 1826.1 | 9.16% | motif file (matrix) |
pdf |
| 7 |  | Cbf1(bHLH)/Yeast-Cbf1-ChIP-Seq(GSE29506)/Homer | 1e-44 | -1.027e+02 | 0.0000 | 442.0 | 9.90% | 953.1 | 4.78% | motif file (matrix) |
pdf |
| 8 |  | bHLHE40(bHLH)/HepG2-BHLHE40-ChIP-Seq(GSE31477)/Homer | 1e-41 | -9.601e+01 | 0.0000 | 459.0 | 10.28% | 1034.3 | 5.19% | motif file (matrix) |
pdf |
| 9 |  | Usf2(bHLH)/C2C12-Usf2-ChIP-Seq(GSE36030)/Homer | 1e-41 | -9.462e+01 | 0.0000 | 430.0 | 9.63% | 948.5 | 4.76% | motif file (matrix) |
pdf |
| 10 |  | CLOCK(bHLH)/Liver-Clock-ChIP-Seq(GSE39860)/Homer | 1e-39 | -9.138e+01 | 0.0000 | 604.0 | 13.53% | 1537.2 | 7.72% | motif file (matrix) |
pdf |
| 11 |  | USF1(bHLH)/GM12878-Usf1-ChIP-Seq(GSE32465)/Homer | 1e-36 | -8.436e+01 | 0.0000 | 517.0 | 11.58% | 1279.0 | 6.42% | motif file (matrix) |
pdf |
| 12 |  | DPL-1(E2F)/cElegans-Adult-ChIP-Seq(modEncode)/Homer | 1e-35 | -8.257e+01 | 0.0000 | 1114.0 | 24.96% | 3477.0 | 17.45% | motif file (matrix) |
pdf |
| 13 |  | SFP1/SacCer-Promoters/Homer | 1e-35 | -8.135e+01 | 0.0000 | 660.0 | 14.78% | 1791.6 | 8.99% | motif file (matrix) |
pdf |
| 14 |  | E-box/Arabidopsis-Promoters/Homer | 1e-31 | -7.302e+01 | 0.0000 | 566.0 | 12.68% | 1511.4 | 7.59% | motif file (matrix) |
pdf |
| 15 |  | Max(bHLH)/K562-Max-ChIP-Seq(GSE31477)/Homer | 1e-28 | -6.672e+01 | 0.0000 | 625.0 | 14.00% | 1762.5 | 8.85% | motif file (matrix) |
pdf |
| 16 |  | TATA-box/SacCer-Promoters/Homer | 1e-28 | -6.523e+01 | 0.0000 | 2079.0 | 46.57% | 7646.4 | 38.38% | motif file (matrix) |
pdf |
| 17 |  | E-box(bHLH)/Promoter/Homer | 1e-28 | -6.478e+01 | 0.0000 | 143.0 | 3.20% | 215.7 | 1.08% | motif file (matrix) |
pdf |
| 18 |  | IBL1(bHLH)/Seedling-IBL1-ChIP-Seq(GSE51120)/Homer | 1e-24 | -5.675e+01 | 0.0000 | 1419.0 | 31.79% | 4961.4 | 24.90% | motif file (matrix) |
pdf |
| 19 |  | PIF4(bHLH)/Seedling-PIF4-ChIP-Seq(GSE35315)/Homer | 1e-23 | -5.511e+01 | 0.0000 | 1021.0 | 22.87% | 3369.9 | 16.91% | motif file (matrix) |
pdf |
| 20 |  | PIF5ox(bHLH)/Arabidopsis-PIF5ox-ChIP-Seq(GSE35062)/Homer | 1e-23 | -5.361e+01 | 0.0000 | 916.0 | 20.52% | 2971.8 | 14.92% | motif file (matrix) |
pdf |
| 21 |  | Pho2(bHLH)/Yeast-Pho2-ChIP-Seq(GSE29506)/Homer | 1e-23 | -5.313e+01 | 0.0000 | 452.0 | 10.13% | 1236.5 | 6.21% | motif file (matrix) |
pdf |
| 22 |  | Egr2(Zf)/Thymocytes-Egr2-ChIP-Seq(GSE34254)/Homer | 1e-20 | -4.763e+01 | 0.0000 | 149.0 | 3.34% | 276.8 | 1.39% | motif file (matrix) |
pdf |
| 23 |  | SPCH(bHLH)/Seedling-SPCH-ChIP-Seq(GSE57497)/Homer | 1e-19 | -4.581e+01 | 0.0000 | 1154.0 | 25.85% | 4009.9 | 20.13% | motif file (matrix) |
pdf |
| 24 |  | E2F4(E2F)/K562-E2F4-ChIP-Seq(GSE31477)/Homer | 1e-18 | -4.268e+01 | 0.0000 | 812.0 | 18.19% | 2677.7 | 13.44% | motif file (matrix) |
pdf |
| 25 |  | FHY3(FAR1)/Arabidopsis-FHY3-ChIP-Seq(GSE30711)/Homer | 1e-17 | -4.077e+01 | 0.0000 | 429.0 | 9.61% | 1241.6 | 6.23% | motif file (matrix) |
pdf |
| 26 |  | TOD6?/SacCer-Promoters/Homer | 1e-17 | -4.004e+01 | 0.0000 | 345.0 | 7.73% | 946.2 | 4.75% | motif file (matrix) |
pdf |
| 27 |  | NPAS2(bHLH)/Liver-NPAS2-ChIP-Seq(GSE39860)/Homer | 1e-15 | -3.491e+01 | 0.0000 | 842.0 | 18.86% | 2887.0 | 14.49% | motif file (matrix) |
pdf |
| 28 |  | KLF14(Zf)/HEK293-KLF14.GFP-ChIP-Seq(GSE58341)/Homer | 1e-15 | -3.481e+01 | 0.0000 | 578.0 | 12.95% | 1851.9 | 9.29% | motif file (matrix) |
pdf |
| 29 |  | c-Myc(bHLH)/mES-cMyc-ChIP-Seq(GSE11431)/Homer | 1e-13 | -3.007e+01 | 0.0000 | 365.0 | 8.18% | 1093.5 | 5.49% | motif file (matrix) |
pdf |
| 30 |  | Arnt:Ahr(bHLH)/MCF7-Arnt-ChIP-Seq(Lo_et_al.)/Homer | 1e-12 | -2.886e+01 | 0.0000 | 1083.0 | 24.26% | 3953.3 | 19.84% | motif file (matrix) |
pdf |
| 31 |  | BMAL1(bHLH)/Liver-Bmal1-ChIP-Seq(GSE39860)/Homer | 1e-12 | -2.805e+01 | 0.0000 | 1473.0 | 33.00% | 5606.6 | 28.14% | motif file (matrix) |
pdf |
| 32 |  | ZNF467(Zf)/HEK293-ZNF467.GFP-ChIP-Seq(GSE58341)/Homer | 1e-11 | -2.755e+01 | 0.0000 | 332.0 | 7.44% | 993.4 | 4.99% | motif file (matrix) |
pdf |
| 33 |  | Egr1(Zf)/K562-Egr1-ChIP-Seq(GSE32465)/Homer | 1e-10 | -2.358e+01 | 0.0000 | 326.0 | 7.30% | 1006.4 | 5.05% | motif file (matrix) |
pdf |
| 34 |  | MITF(bHLH)/MastCells-MITF-ChIP-Seq(GSE48085)/Homer | 1e-10 | -2.328e+01 | 0.0000 | 798.0 | 17.88% | 2869.8 | 14.40% | motif file (matrix) |
pdf |
| 35 |  | GAGA-repeat/Arabidopsis-Promoters/Homer | 1e-9 | -2.133e+01 | 0.0000 | 548.0 | 12.28% | 1890.2 | 9.49% | motif file (matrix) |
pdf |
| 36 |  | Pho4(bHLH)/Yeast-Pho4-ChIP-Seq(GSE29506)/Homer | 1e-9 | -2.102e+01 | 0.0000 | 167.0 | 3.74% | 451.1 | 2.26% | motif file (matrix) |
pdf |
| 37 |  | KLF10(Zf)/HEK293-KLF10.GFP-ChIP-Seq(GSE58341)/Homer | 1e-9 | -2.089e+01 | 0.0000 | 249.0 | 5.58% | 745.9 | 3.74% | motif file (matrix) |
pdf |
| 38 |  | Maz(Zf)/HepG2-Maz-ChIP-Seq(GSE31477)/Homer | 1e-9 | -2.087e+01 | 0.0000 | 286.0 | 6.41% | 882.4 | 4.43% | motif file (matrix) |
pdf |
| 39 |  | E2F1(E2F)/Hela-E2F1-ChIP-Seq(GSE22478)/Homer | 1e-8 | -2.032e+01 | 0.0000 | 210.0 | 4.70% | 608.3 | 3.05% | motif file (matrix) |
pdf |
| 40 |  | Klf4(Zf)/mES-Klf4-ChIP-Seq(GSE11431)/Homer | 1e-8 | -1.948e+01 | 0.0000 | 158.0 | 3.54% | 430.6 | 2.16% | motif file (matrix) |
pdf |
| 41 |  | Klf9(Zf)/GBM-Klf9-ChIP-Seq(GSE62211)/Homer | 1e-8 | -1.918e+01 | 0.0000 | 133.0 | 2.98% | 346.8 | 1.74% | motif file (matrix) |
pdf |
| 42 |  | SUT1?/SacCer-Promoters/Homer | 1e-7 | -1.744e+01 | 0.0000 | 3045.0 | 68.21% | 12818.6 | 64.34% | motif file (matrix) |
pdf |
| 43 | 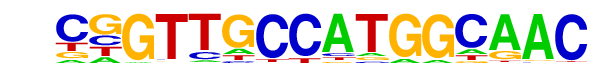 | RFX(HTH)/K562-RFX3-ChIP-Seq(SRA012198)/Homer | 1e-7 | -1.736e+01 | 0.0000 | 100.0 | 2.24% | 246.8 | 1.24% | motif file (matrix) |
pdf |
| 44 |  | ZBTB33(Zf)/GM12878-ZBTB33-ChIP-Seq(GSE32465)/Homer | 1e-7 | -1.727e+01 | 0.0000 | 61.0 | 1.37% | 124.9 | 0.63% | motif file (matrix) |
pdf |
| 45 |  | HIF-1a(bHLH)/MCF7-HIF1a-ChIP-Seq(GSE28352)/Homer | 1e-7 | -1.710e+01 | 0.0000 | 516.0 | 11.56% | 1822.6 | 9.15% | motif file (matrix) |
pdf |
| 46 |  | BATF(bZIP)/Th17-BATF-ChIP-Seq(GSE39756)/Homer | 1e-6 | -1.518e+01 | 0.0000 | 638.0 | 14.29% | 2349.7 | 11.79% | motif file (matrix) |
pdf |
| 47 |  | E2F7(E2F)/Hela-E2F7-ChIP-Seq(GSE32673)/Homer | 1e-5 | -1.372e+01 | 0.0000 | 126.0 | 2.82% | 358.0 | 1.80% | motif file (matrix) |
pdf |
| 48 |  | HIF-1b(HLH)/T47D-HIF1b-ChIP-Seq(GSE59937)/Homer | 1e-5 | -1.347e+01 | 0.0000 | 1692.0 | 37.90% | 6881.9 | 34.54% | motif file (matrix) |
pdf |
| 49 |  | SeqBias: TA-repeat | 1e-5 | -1.222e+01 | 0.0000 | 3950.0 | 88.49% | 17184.5 | 86.25% | motif file (matrix) |
pdf |
| 50 |  | BORIS(Zf)/K562-CTCFL-ChIP-Seq(GSE32465)/Homer | 1e-5 | -1.214e+01 | 0.0000 | 52.0 | 1.16% | 117.4 | 0.59% | motif file (matrix) |
pdf |
| 51 |  | Rfx2(HTH)/LoVo-RFX2-ChIP-Seq(GSE49402)/Homer | 1e-5 | -1.208e+01 | 0.0000 | 102.0 | 2.28% | 285.6 | 1.43% | motif file (matrix) |
pdf |
| 52 |  | TATA-box/Drosophila-Promoters/Homer | 1e-5 | -1.196e+01 | 0.0000 | 228.0 | 5.11% | 755.8 | 3.79% | motif file (matrix) |
pdf |
| 53 |  | E2F(E2F)/Hela-CellCycle-Expression/Homer | 1e-5 | -1.195e+01 | 0.0000 | 131.0 | 2.93% | 390.7 | 1.96% | motif file (matrix) |
pdf |
| 54 |  | EKLF(Zf)/Erythrocyte-Klf1-ChIP-Seq(GSE20478)/Homer | 1e-5 | -1.183e+01 | 0.0001 | 106.0 | 2.37% | 301.6 | 1.51% | motif file (matrix) |
pdf |
| 55 |  | p53(p53)/Saos-p53-ChIP-Seq(GSE15780)/Homer | 1e-4 | -1.142e+01 | 0.0001 | 80.0 | 1.79% | 213.7 | 1.07% | motif file (matrix) |
pdf |
| 56 |  | p53(p53)/Saos-p53-ChIP-Seq/Homer | 1e-4 | -1.142e+01 | 0.0001 | 80.0 | 1.79% | 213.7 | 1.07% | motif file (matrix) |
pdf |
| 57 |  | E2F6(E2F)/Hela-E2F6-ChIP-Seq(GSE31477)/Homer | 1e-4 | -1.054e+01 | 0.0002 | 284.0 | 6.36% | 993.9 | 4.99% | motif file (matrix) |
pdf |
| 58 |  | KLF5(Zf)/LoVo-KLF5-ChIP-Seq(GSE49402)/Homer | 1e-4 | -1.053e+01 | 0.0002 | 453.0 | 10.15% | 1676.8 | 8.42% | motif file (matrix) |
pdf |
| 59 |  | FOXP1(Forkhead)/H9-FOXP1-ChIP-Seq(GSE31006)/Homer | 1e-4 | -1.023e+01 | 0.0002 | 807.0 | 18.08% | 3161.0 | 15.87% | motif file (matrix) |
pdf |
| 60 |  | SpiB(ETS)/OCILY3-SPIB-ChIP-Seq(GSE56857)/Homer | 1e-4 | -9.807e+00 | 0.0004 | 300.0 | 6.72% | 1068.6 | 5.36% | motif file (matrix) |
pdf |
| 61 |  | PU.1-IRF(ETS:IRF)/Bcell-PU.1-ChIP-Seq(GSE21512)/Homer | 1e-4 | -9.211e+00 | 0.0006 | 1472.0 | 32.97% | 6054.4 | 30.39% | motif file (matrix) |
pdf |
| 62 |  | NRF(NRF)/Promoter/Homer | 1e-3 | -8.973e+00 | 0.0008 | 92.0 | 2.06% | 273.1 | 1.37% | motif file (matrix) |
pdf |
| 63 |  | CRE(bZIP)/Promoter/Homer | 1e-3 | -8.915e+00 | 0.0008 | 204.0 | 4.57% | 700.9 | 3.52% | motif file (matrix) |
pdf |
| 64 |  | Dorsal(RHD)/Embryo-dl-ChIP-Seq(GSE65441)/Homer | 1e-3 | -8.698e+00 | 0.0010 | 222.0 | 4.97% | 774.1 | 3.89% | motif file (matrix) |
pdf |
| 65 |  | NRF1(NRF)/MCF7-NRF1-ChIP-Seq(Unpublished)/Homer | 1e-3 | -8.370e+00 | 0.0014 | 43.0 | 0.96% | 106.4 | 0.53% | motif file (matrix) |
pdf |
| 66 |  | Elk4(ETS)/Hela-Elk4-ChIP-Seq(GSE31477)/Homer | 1e-3 | -7.875e+00 | 0.0022 | 1050.0 | 23.52% | 4268.9 | 21.43% | motif file (matrix) |
pdf |
| 67 |  | Fra1(bZIP)/BT549-Fra1-ChIP-Seq(GSE46166)/Homer | 1e-3 | -7.232e+00 | 0.0042 | 543.0 | 12.16% | 2123.4 | 10.66% | motif file (matrix) |
pdf |
| 68 |  | Rfx5(HTH)/GM12878-Rfx5-ChIP-Seq(GSE31477)/Homer | 1e-3 | -7.037e+00 | 0.0050 | 283.0 | 6.34% | 1047.8 | 5.26% | motif file (matrix) |
pdf |
| 69 |  | Pax8(Paired,Homeobox)/Thyroid-Pax8-ChIP-Seq(GSE26938)/Homer | 1e-3 | -6.960e+00 | 0.0053 | 126.0 | 2.82% | 421.3 | 2.11% | motif file (matrix) |
pdf |
| 70 |  | Elk1(ETS)/Hela-Elk1-ChIP-Seq(GSE31477)/Homer | 1e-2 | -6.828e+00 | 0.0060 | 864.0 | 19.35% | 3502.6 | 17.58% | motif file (matrix) |
pdf |
| 71 |  | HIF2a(bHLH)/785_O-HIF2a-ChIP-Seq(GSE34871)/Homer | 1e-2 | -6.818e+00 | 0.0060 | 733.0 | 16.42% | 2942.0 | 14.77% | motif file (matrix) |
pdf |
| 72 |  | RXR(NR),DR1/3T3L1-RXR-ChIP-Seq(GSE13511)/Homer | 1e-2 | -6.715e+00 | 0.0065 | 521.0 | 11.67% | 2044.7 | 10.26% | motif file (matrix) |
pdf |
| 73 |  | Tbox:Smad(T-box,MAD)/ESCd5-Smad2_3-ChIP-Seq(GSE29422)/Homer | 1e-2 | -6.610e+00 | 0.0071 | 113.0 | 2.53% | 375.4 | 1.88% | motif file (matrix) |
pdf |
| 74 |  | CTCF(Zf)/CD4+-CTCF-ChIP-Seq(Barski_et_al.)/Homer | 1e-2 | -6.457e+00 | 0.0082 | 45.0 | 1.01% | 124.7 | 0.63% | motif file (matrix) |
pdf |
| 75 |  | Trl(Zf)/S2-GAGAfactor-ChIP-Seq(GSE40646)/Homer | 1e-2 | -6.262e+00 | 0.0098 | 1737.0 | 38.91% | 7334.0 | 36.81% | motif file (matrix) |
pdf |
| 76 |  | Unknown-ESC-element(?)/mES-Nanog-ChIP-Seq(GSE11724)/Homer | 1e-2 | -6.192e+00 | 0.0104 | 193.0 | 4.32% | 697.4 | 3.50% | motif file (matrix) |
pdf |
| 77 |  | PAX5(Paired,Homeobox)/GM12878-PAX5-ChIP-Seq(GSE32465)/Homer | 1e-2 | -6.183e+00 | 0.0104 | 214.0 | 4.79% | 782.5 | 3.93% | motif file (matrix) |
pdf |
| 78 |  | X-box(HTH)/NPC-H3K4me1-ChIP-Seq(GSE16256)/Homer | 1e-2 | -5.900e+00 | 0.0136 | 135.0 | 3.02% | 470.8 | 2.36% | motif file (matrix) |
pdf |
| 79 |  | ETS(ETS)/Promoter/Homer | 1e-2 | -5.832e+00 | 0.0144 | 354.0 | 7.93% | 1367.0 | 6.86% | motif file (matrix) |
pdf |
| 80 |  | Stat3(Stat)/mES-Stat3-ChIP-Seq(GSE11431)/Homer | 1e-2 | -5.805e+00 | 0.0146 | 489.0 | 10.95% | 1934.3 | 9.71% | motif file (matrix) |
pdf |
| 81 |  | AP-2alpha(AP2)/Hela-AP2alpha-ChIP-Seq(GSE31477)/Homer | 1e-2 | -5.549e+00 | 0.0186 | 165.0 | 3.70% | 595.3 | 2.99% | motif file (matrix) |
pdf |
| 82 |  | GEI-11(Myb?)/cElegans-L4-GEI11-ChIP-Seq(modEncode)/Homer | 1e-2 | -4.803e+00 | 0.0387 | 43.0 | 0.96% | 129.6 | 0.65% | motif file (matrix) |
pdf |
| 83 |  | AP-1(bZIP)/ThioMac-PU.1-ChIP-Seq(GSE21512)/Homer | 1e-2 | -4.730e+00 | 0.0411 | 637.0 | 14.27% | 2601.0 | 13.05% | motif file (matrix) |
pdf |
| 84 |  | Fosl2(bZIP)/3T3L1-Fosl2-ChIP-Seq(GSE56872)/Homer | 1e-2 | -4.725e+00 | 0.0411 | 195.0 | 4.37% | 732.2 | 3.67% | motif file (matrix) |
pdf |







































































































































































