| p-value: | 1e-25 |
| log p-value: | -5.910e+01 |
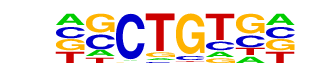
| Information Content per bp: | 1.810 |
| Number of Target Sequences with motif | 554.0 |
| Percentage of Target Sequences with motif | 16.13% |
| Number of Background Sequences with motif | 2108.8 |
| Percentage of Background Sequences with motif | 10.25% |
| Average Position of motif in Targets | 427.5 +/- 416.7bp |
| Average Position of motif in Background | 271.7 +/- 227.3bp |
| Strand Bias (log2 ratio + to - strand density) | -0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.17 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
CDC5(MYB)/Arabidopsis thaliana/AthaMap
| Match Rank: | 1 |
| Score: | 0.71
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --CGCTGTGC-
CGCGCTGAGCN |
|


|
|
CDC5/MA0579.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.71
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --CGCTGTGC-
CGCGCTGAGCN |
|


|
|
POL009.1_DCE_S_II/Jaspar
| Match Rank: | 3 |
| Score: | 0.71
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | CGCTGTGC
-GCTGTG- |
|

|
|
PB0207.1_Zic3_2/Jaspar
| Match Rank: | 4 |
| Score: | 0.67
| | Offset: | -5
| | Orientation: | reverse strand |
| Alignment: | -----CGCTGTGC--
NNTCCTGCTGTGNNN |
|


|
|
AR-halfsite(NR)/LNCaP-AR-ChIP-Seq(GSE27824)/Homer
| Match Rank: | 5 |
| Score: | 0.66
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | CGCTGTGC----
--CTGTTCCTGG |
|


|
|
POL010.1_DCE_S_III/Jaspar
| Match Rank: | 6 |
| Score: | 0.65
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CGCTGTGC
NGCTN--- |
|


|
|
MET28/MA0332.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | CGCTGTGC
--CTGTGG |
|


|
|
MET28(MacIsaac)/Yeast
| Match Rank: | 8 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | CGCTGTGC
--CTGTGG |
|


|
|
Zic(Zf)/Cerebellum-ZIC1.2-ChIP-Seq(GSE60731)/Homer
| Match Rank: | 9 |
| Score: | 0.60
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --CGCTGTGC
CCTGCTGAGH |
|


|
|
PB0205.1_Zic1_2/Jaspar
| Match Rank: | 10 |
| Score: | 0.60
| | Offset: | -5
| | Orientation: | reverse strand |
| Alignment: | -----CGCTGTGC--
TNTCCTGCTGTGNNG |
|


|
|





















