| p-value: | 1e-12 |
| log p-value: | -2.909e+01 |
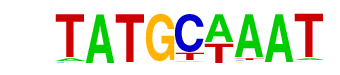
| Information Content per bp: | 1.747 |
| Number of Target Sequences with motif | 445.0 |
| Percentage of Target Sequences with motif | 10.23% |
| Number of Background Sequences with motif | 1567.6 |
| Percentage of Background Sequences with motif | 7.22% |
| Average Position of motif in Targets | 387.5 +/- 337.3bp |
| Average Position of motif in Background | 273.4 +/- 194.2bp |
| Strand Bias (log2 ratio + to - strand density) | -0.4 |
| Multiplicity (# of sites on avg that occur together) | 1.14 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
P0510F09.23/MA1030.1/Jaspar
| Match Rank: | 1 |
| Score: | 0.79
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA--
AACCCTAGAT |
|


|
|
TRB2/MA1073.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.74
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA
TGCCCTAA |
|


|
|
TBF1/MA0403.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.70
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA
AACCCTAA |
|


|
|
cad/dmmpmm(SeSiMCMC)/fly
| Match Rank: | 4 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA-
--GCATAAA |
|


|
|
MYB59/MA1042.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.63
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | TAMSCTAA-
-TACCTAAC |
|


|
|
NAC043/MA1045.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.63
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TAMSCTAA-
ATTACGTAAA |
|


|
|
Barhl1/MA0877.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.62
| | Offset: | 3
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA-----
---GCTAATTGCT |
|


|
|
CIN5/MA0284.1/Jaspar
| Match Rank: | 8 |
| Score: | 0.61
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA--
TTACGTAATC |
|


|
|
STP3/MA0396.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.61
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TAMSCTAA-
TGCGCTAGC |
|


|
|
POU3F4/MA0789.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.60
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TAMSCTAA-
TATGCAAAT |
|

|
|





















