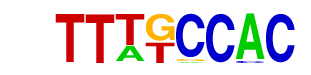
| p-value: | 1e-14 |
| log p-value: | -3.369e+01 |
| Information Content per bp: | 1.858 |
| Number of Target Sequences with motif | 218.0 |
| Percentage of Target Sequences with motif | 5.14% |
| Number of Background Sequences with motif | 592.8 |
| Percentage of Background Sequences with motif | 2.90% |
| Average Position of motif in Targets | 450.9 +/- 341.9bp |
| Average Position of motif in Background | 289.0 +/- 202.5bp |
| Strand Bias (log2 ratio + to - strand density) | 0.0 |
| Multiplicity (# of sites on avg that occur together) | 1.02 |
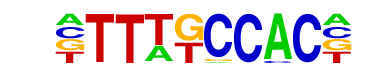
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
RPN4/RPN4_H2O2Lo/[](Harbison)/Yeast
| Match Rank: | 1 |
| Score: | 0.89
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TTTKCCAC-
TTTGCCACC |
|


|
|
RPN4(MacIsaac)/Yeast
| Match Rank: | 2 |
| Score: | 0.87
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TTTKCCAC--
TTTTGCCACCG |
|


|
|
NFATC2/MA0152.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.80
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | TTTKCCAC
TTTTCCA- |
|


|
|
NFAT5/MA0606.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.79
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TTTKCCAC-
ATTTTCCATT |
|


|
|
NFAT(RHD)/Jurkat-NFATC1-ChIP-Seq(Jolma_et_al.)/Homer
| Match Rank: | 5 |
| Score: | 0.77
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TTTKCCAC-
ATTTTCCATT |
|

|
|
MET32/MA0334.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.71
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | TTTKCCAC-
--CGCCACA |
|


|
|
NFATC3/MA0625.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.71
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TTTKCCAC-
ATTTTCCATT |
|


|
|
Su(H)/dmmpmm(Papatsenko)/fly
| Match Rank: | 8 |
| Score: | 0.71
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TTTKCCAC-
GTTCCCACG |
|


|
|
Su(H)/MA0085.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.70
| | Offset: | -6
| | Orientation: | reverse strand |
| Alignment: | ------TTTKCCAC--
ATCTCGGTTCCCACAN |
|


|
|
Rbpj1(?)/Panc1-Rbpj1-ChIP-Seq(GSE47459)/Homer
| Match Rank: | 10 |
| Score: | 0.70
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TTTKCCAC-
HTTTCCCASG |
|


|
|