| p-value: | 1e-13 |
| log p-value: | -3.129e+01 |
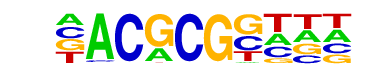
| Information Content per bp: | 1.905 |
| Number of Target Sequences with motif | 215.0 |
| Percentage of Target Sequences with motif | 4.70% |
| Number of Background Sequences with motif | 567.8 |
| Percentage of Background Sequences with motif | 2.71% |
| Average Position of motif in Targets | 392.1 +/- 285.7bp |
| Average Position of motif in Background | 281.4 +/- 206.5bp |
| Strand Bias (log2 ratio + to - strand density) | -0.8 |
| Multiplicity (# of sites on avg that occur together) | 1.06 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
MET31(MacIsaac)/Yeast
| Match Rank: | 1 |
| Score: | 0.76
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --CACACGGM
GCCACACC-- |
|


|
|
PB0044.1_Mtf1_1/Jaspar
| Match Rank: | 2 |
| Score: | 0.73
| | Offset: | -6
| | Orientation: | reverse strand |
| Alignment: | ------CACACGGM--
NNTTTGCACACGGCCC |
|


|
|
MTF1/MA0863.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.71
| | Offset: | -4
| | Orientation: | forward strand |
| Alignment: | ----CACACGGM--
TTTGCACACGGCAC |
|


|
|
CMTA3/MA0970.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.70
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --CACACGGM
NNNACGCGG- |
|


|
|
Unknown1(NR/Ini-like)/Drosophila-Promoters/Homer
| Match Rank: | 5 |
| Score: | 0.69
| | Offset: | -5
| | Orientation: | forward strand |
| Alignment: | -----CACACGGM
MYGGTCACACTG- |
|


|
|
CMTA2/MA0969.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.68
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | CACACGGM--
-ACGCGGNNN |
|

|
|
SOX10/MA0442.1/Jaspar
| Match Rank: | 7 |
| Score: | 0.67
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | CACACGGM
-ACAAAG- |
|


|
|
YDR026C(MacIsaac)/Yeast
| Match Rank: | 8 |
| Score: | 0.66
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | CACACGGM
-ACCCGG- |
|


|
|
NAC055/MA0937.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.65
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | CACACGGM-
-ACACGTAA |
|


|
|
PB0130.1_Gm397_2/Jaspar
| Match Rank: | 10 |
| Score: | 0.65
| | Offset: | -7
| | Orientation: | forward strand |
| Alignment: | -------CACACGGM-
AGCGGCACACACGCAA |
|


|
|





















