| p-value: | 1e-22 |
| log p-value: | -5.231e+01 |
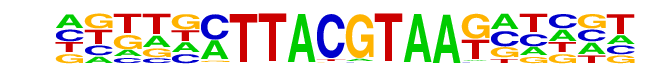
| Information Content per bp: | 1.760 |
| Number of Target Sequences with motif | 836.0 |
| Percentage of Target Sequences with motif | 22.35% |
| Number of Background Sequences with motif | 3189.1 |
| Percentage of Background Sequences with motif | 16.10% |
| Average Position of motif in Targets | 404.1 +/- 325.9bp |
| Average Position of motif in Background | 291.2 +/- 215.0bp |
| Strand Bias (log2 ratio + to - strand density) | -0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.18 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
GMEB2/MA0862.1/Jaspar
| Match Rank: | 1 |
| Score: | 0.87
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TACGTAMT
TTACGTAA- |
|


|
|
YAP3/MA0416.1/Jaspar
| Match Rank: | 2 |
| Score: | 0.84
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TACGTAMT
TACGTAAT |
|


|
|
Gmeb1/MA0615.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.82
| | Offset: | -5
| | Orientation: | forward strand |
| Alignment: | -----TACGTAMT----
GAGTGTACGTAAGATGG |
|


|
|
PB0027.1_Gmeb1_1/Jaspar
| Match Rank: | 4 |
| Score: | 0.82
| | Offset: | -5
| | Orientation: | forward strand |
| Alignment: | -----TACGTAMT----
GAGTGTACGTAAGATGG |
|


|
|
NAC043/MA1045.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.81
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --TACGTAMT
NTTACGTAAT |
|


|
|
NAC025/MA0935.1/Jaspar
| Match Rank: | 6 |
| Score: | 0.80
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --TACGTAMT
GTTACGTA-- |
|


|
|
CIN5/CIN5_H2O2Lo/[](Harbison)/Yeast
| Match Rank: | 7 |
| Score: | 0.79
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TACGTAMT
TTACGTAA- |
|


|
|
YAP6(MacIsaac)/Yeast
| Match Rank: | 8 |
| Score: | 0.78
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TACGTAMT
TTACATAA- |
|


|
|
gt/MA0447.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.78
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --TACGTAMT
ATTACGTAAT |
|


|
|
YAP1/MA0415.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.78
| | Offset: | -7
| | Orientation: | forward strand |
| Alignment: | -------TACGTAMT-----
ATTTGCTTACGTAAGCTCGT |
|

|
|





















