| p-value: | 1e-40 |
| log p-value: | -9.387e+01 |
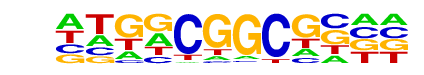
| Information Content per bp: | 1.833 |
| Number of Target Sequences with motif | 508.0 |
| Percentage of Target Sequences with motif | 11.19% |
| Number of Background Sequences with motif | 1217.3 |
| Percentage of Background Sequences with motif | 5.93% |
| Average Position of motif in Targets | 397.5 +/- 276.9bp |
| Average Position of motif in Background | 306.3 +/- 237.0bp |
| Strand Bias (log2 ratio + to - strand density) | -0.2 |
| Multiplicity (# of sites on avg that occur together) | 1.11 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
STB4/STB4_YPD/[](Harbison)/Yeast
| Match Rank: | 1 |
| Score: | 0.75
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CGGCTCGA
TCGGNNCGA |
|


|
|
STB4(MacIsaac)/Yeast
| Match Rank: | 2 |
| Score: | 0.75
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CGGCTCGA
TCGGNNCGA |
|


|
|
XBP1/MA0414.1/Jaspar
| Match Rank: | 3 |
| Score: | 0.69
| | Offset: | 3
| | Orientation: | forward strand |
| Alignment: | CGGCTCGA--
---CTCGAGA |
|


|
|
ERF069/MA0997.1/Jaspar
| Match Rank: | 4 |
| Score: | 0.66
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---CGGCTCGA
NGGCGGCGN-- |
|


|
|
STP1/MA0394.1/Jaspar
| Match Rank: | 5 |
| Score: | 0.65
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --CGGCTCGA
TGCGGCGC-- |
|


|
|
POL010.1_DCE_S_III/Jaspar
| Match Rank: | 6 |
| Score: | 0.65
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | CGGCTCGA
-NGCTN-- |
|


|
|
STP1(MacIsaac)/Yeast
| Match Rank: | 7 |
| Score: | 0.65
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --CGGCTCGA
TGCGGCGC-- |
|


|
|
XBP1/Literature(Harbison)/Yeast
| Match Rank: | 8 |
| Score: | 0.64
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | CGGCTCGA--
---CTCGAAG |
|


|
|
ERF11/MA1001.1/Jaspar
| Match Rank: | 9 |
| Score: | 0.64
| | Offset: | -4
| | Orientation: | reverse strand |
| Alignment: | ----CGGCTCGA
NTGNCGGCGN-- |
|

|
|
RSC3/MA0374.1/Jaspar
| Match Rank: | 10 |
| Score: | 0.64
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | CGGCTCGA
NNGCGCG- |
|


|
|





















